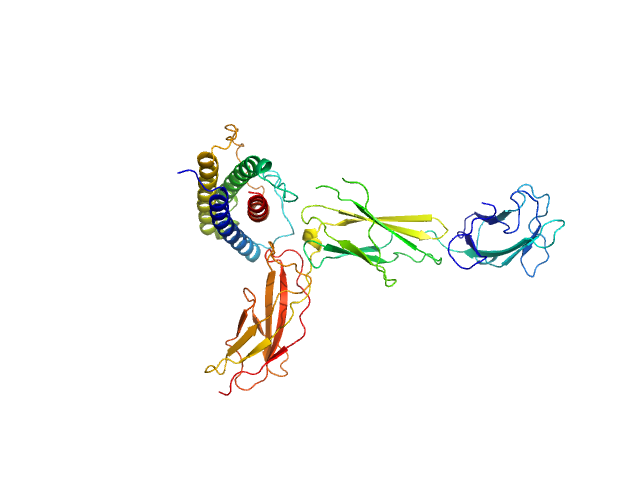
UniProt ID: Q14626 (23-319) Interleukin-11 receptor subunit alpha
UniProt ID: A8K3F7 (32-199) Interleukin 11
|
|
|
|
| Sample: |
Interleukin-11 receptor subunit alpha monomer, 32 kDa Homo sapiens protein
Interleukin 11 monomer, 18 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.2% sodium azide, pH: 8.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2019 Jun 8
|
The structure of the extracellular domains of human interleukin 11 α-receptor reveals mechanisms of cytokine engagement
Journal of Biological Chemistry :jbc.RA119.012351 (2020)
Metcalfe R, Aizel K, Zlatic C, Nguyen P, Morton C, Lio D, Cheng H, Dobson R, Parker M, Gooley P, Putoczki T, Griffin M
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
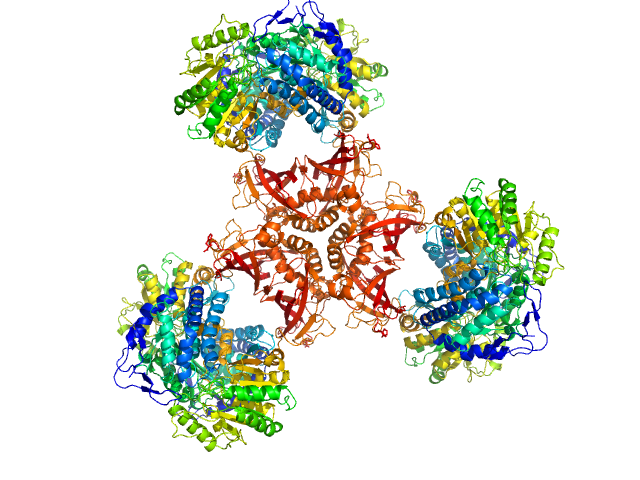
UniProt ID: P77455 (1-681) Bifunctional protein PaaZ
|
|
|
|
| Sample: |
Bifunctional protein PaaZ hexamer, 438 kDa Escherichia coli protein
|
| Buffer: |
25 mM HEPES, 50 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Feb 24
|
Molecular basis for metabolite channeling in a ring opening enzyme of the phenylacetate degradation pathway.
Nat Commun 10(1):4127 (2019)
Sathyanarayanan N, Cannone G, Gakhar L, Katagihallimath N, Sowdhamini R, Ramaswamy S, Vinothkumar KR
|
| RgGuinier |
6.2 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
636 |
nm3 |
|
|
UniProt ID: P11532 (1990-3040) Human dystrophin central domain R16-24 del44-54 fragment
|
|
|
|
| Sample: |
Human dystrophin central domain R16-24 del44-54 fragment monomer, 57 kDa Homo sapiens protein
|
| Buffer: |
20 mM Na-phosphate, 300 mM NaCl, 1 mM EDTA, 2% glycerol,, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Jul 8
|
Dystrophin SAXS data
Raphael Dos Santos Morais
|
| RgGuinier |
6.4 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
159 |
nm3 |
|
|
UniProt ID: Q9U6M1 (2-827) Thymine dioxygenase JBP1
|
|
|
|
| Sample: |
Thymine dioxygenase JBP1 monomer, 93 kDa Leishmania tarentolae protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 21
|
The domain architecture of protozoan protein J-DNA-binding protein 1 suggests synergy between base J DNA binding and thymidine hydroxylase activity.
J Biol Chem (2019)
Adamopoulos A, Heidebrecht T, Roosendaal J, Touw WG, Phan IQ, Beijnen J, Perrakis A
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
128 |
nm3 |
|
|
UniProt ID: Q9U6M1 (2-827) Delta-JDBD
|
|
|
|
| Sample: |
Delta-JDBD monomer, 72 kDa Leismania tarentolae protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 21
|
The domain architecture of protozoan protein J-DNA-binding protein 1 suggests synergy between base J DNA binding and thymidine hydroxylase activity.
J Biol Chem (2019)
Adamopoulos A, Heidebrecht T, Roosendaal J, Touw WG, Phan IQ, Beijnen J, Perrakis A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
108 |
nm3 |
|
|
UniProt ID: Q9U6M1 (382-561) J-DNA binding domain
|
|
|
|
| Sample: |
J-DNA binding domain monomer, 21 kDa Leishmania tarentolae protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 4
|
The domain architecture of protozoan protein J-DNA-binding protein 1 suggests synergy between base J DNA binding and thymidine hydroxylase activity.
J Biol Chem (2019)
Adamopoulos A, Heidebrecht T, Roosendaal J, Touw WG, Phan IQ, Beijnen J, Perrakis A
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
UniProt ID: Q9U6M1 (382-561) J-DNA binding domain
UniProt ID: None (None-None) J-DNA (23mer)
|
|
|
|
| Sample: |
J-DNA binding domain monomer, 21 kDa Leishmania tarentolae protein
J-DNA (23mer) monomer, 14 kDa DNA
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 21
|
The domain architecture of protozoan protein J-DNA-binding protein 1 suggests synergy between base J DNA binding and thymidine hydroxylase activity.
J Biol Chem (2019)
Adamopoulos A, Heidebrecht T, Roosendaal J, Touw WG, Phan IQ, Beijnen J, Perrakis A
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
43 |
nm3 |
|
|
UniProt ID: Q9U6M1 (2-827) Thymine dioxygenase JBP1
UniProt ID: None (None-None) J-DNA (23mer)
|
|
|
|
| Sample: |
Thymine dioxygenase JBP1 monomer, 93 kDa Leishmania tarentolae protein
J-DNA (23mer) monomer, 14 kDa DNA
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 21
|
The domain architecture of protozoan protein J-DNA-binding protein 1 suggests synergy between base J DNA binding and thymidine hydroxylase activity.
J Biol Chem (2019)
Adamopoulos A, Heidebrecht T, Roosendaal J, Touw WG, Phan IQ, Beijnen J, Perrakis A
|
| RgGuinier |
4.1 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
148 |
nm3 |
|
|
UniProt ID: Q8U4M2 (1-227) Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase
UniProt ID: Q8U160 (1-123) 50S ribosomal protein L7Ae
UniProt ID: Q8U4M1 (1-402) NOP5/NOP56 related protein
UniProt ID: None (None-None) pyrococcus furiosus sR26 stabilized construct
UniProt ID: None (None-None) Pyrococcus furiosus sR26 substrate D'
|
|
|
|
| Sample: |
Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase dimer, 52 kDa Pyrococcus furiosus protein
50S ribosomal protein L7Ae dimer, 27 kDa Pyrococcus furiosus protein
NOP5/NOP56 related protein dimer, 94 kDa Pyrococcus furiosus protein
Pyrococcus furiosus sR26 stabilized construct monomer, 24 kDa Pyrococcus furiosus RNA
Pyrococcus furiosus sR26 substrate D' monomer, 4 kDa Pyrococcus furiosus RNA
|
| Buffer: |
50 mM phosphate 500 mM NaCl 42%D2O, pH: 6.6 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2015 May 24
|
The guide sRNA sequence determines the activity level of box C/D RNPs.
Elife 9 (2020)
Graziadei A, Gabel F, Kirkpatrick J, Carlomagno T
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
UniProt ID: Q8U4M2 (1-227) Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase
UniProt ID: Q8U160 (1-123) 50S ribosomal protein L7Ae
UniProt ID: Q8U4M1 (1-402) NOP5/NOP56 related protein
UniProt ID: None (None-None) pyrococcus furiosus sR26 stabilized construct
UniProt ID: None (None-None) Pyrococcus furiosus sR26 substrate D'
|
|
|
|
| Sample: |
Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase dimer, 52 kDa Pyrococcus furiosus protein
50S ribosomal protein L7Ae dimer, 27 kDa Pyrococcus furiosus protein
NOP5/NOP56 related protein dimer, 94 kDa Pyrococcus furiosus protein
Pyrococcus furiosus sR26 stabilized construct monomer, 24 kDa Pyrococcus furiosus RNA
Pyrococcus furiosus sR26 substrate D' monomer, 4 kDa Pyrococcus furiosus RNA
|
| Buffer: |
50 mM phosphate 500 mM NaCl 42%D2O, pH: 6.6 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2015 Sep 21
|
The guide sRNA sequence determines the activity level of box C/D RNPs.
Elife 9 (2020)
Graziadei A, Gabel F, Kirkpatrick J, Carlomagno T
|
| RgGuinier |
4.1 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
210 |
nm3 |
|
|