UniProt ID: Q9UPT6 (22-187) C-Jun-amino-terminal kinase-interacting protein 3
|
|
|

|
| Sample: |
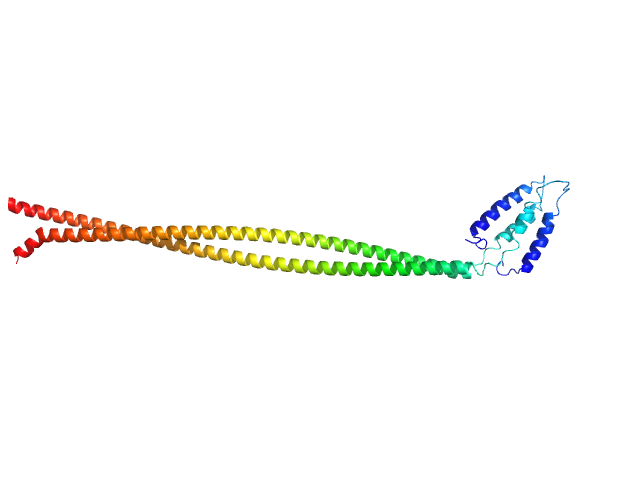
C-Jun-amino-terminal kinase-interacting protein 3 dimer, 40 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 0.5 mM TCEP, pH: 7.1 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Sep 22
|
Structural characterization of the RH1-LZI tandem of JIP3/4 highlights RH1 domains as a cytoskeletal motor-binding motif.
Sci Rep 9(1):16036 (2019)
Vilela F, Velours C, Chenon M, Aumont-Nicaise M, Campanacci V, Thureau A, Pylypenko O, Andreani J, Llinas P, Ménétrey J
|
| RgGuinier |
6.3 |
nm |
| Dmax |
23.1 |
nm |
| VolumePorod |
140 |
nm3 |
|
|
UniProt ID: Q9UPT6 (22-187) C-Jun-amino-terminal kinase-interacting protein 3
|
|
|

|
| Sample: |
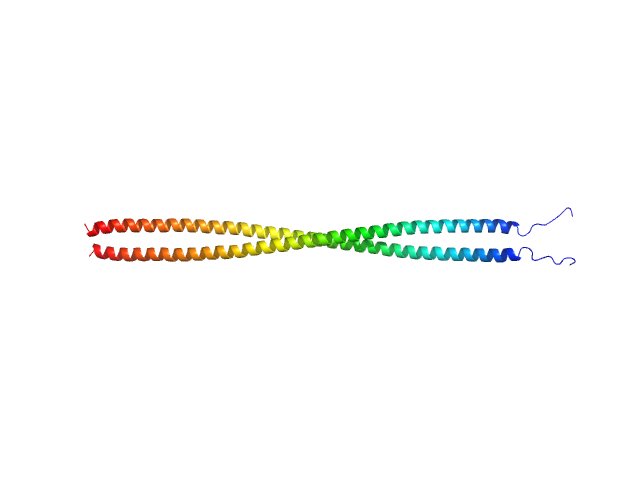
C-Jun-amino-terminal kinase-interacting protein 3 dimer, 40 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 0.5 mM TCEP, pH: 7.1 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Sep 22
|
Structural characterization of the RH1-LZI tandem of JIP3/4 highlights RH1 domains as a cytoskeletal motor-binding motif.
Sci Rep 9(1):16036 (2019)
Vilela F, Velours C, Chenon M, Aumont-Nicaise M, Campanacci V, Thureau A, Pylypenko O, Andreani J, Llinas P, Ménétrey J
|
| RgGuinier |
5.0 |
nm |
| Dmax |
19.8 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
UniProt ID: Q9NZN9 (None-None) Aryl-hydrocarbon-interacting protein-like 1(1-316)
|
|
|
|
| Sample: |
Aryl-hydrocarbon-interacting protein-like 1(1-316) monomer, 37 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 2.5 % glycerol and 6 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jul 17
|
Interaction of the tetratricopeptide repeat domain of aryl hydrocarbon receptor-interacting protein-like 1 with the regulatory Pγ subunit of phosphodiesterase 6.
J Biol Chem 294(43):15795-15807 (2019)
Yadav RP, Boyd K, Yu L, Artemyev NO
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
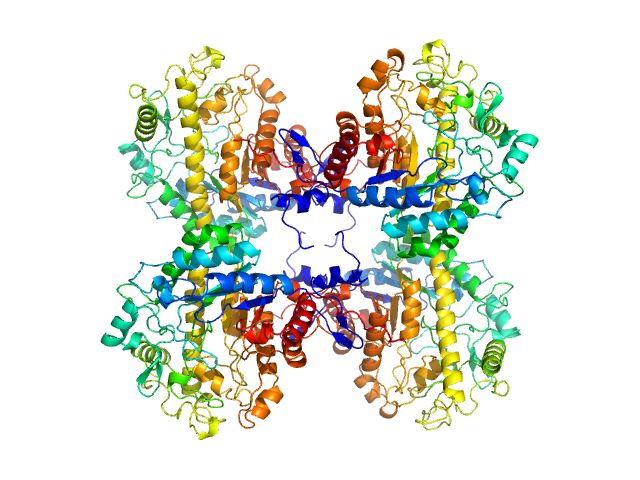
UniProt ID: O65258 (56-542) Beta-amylase 2, chloroplastic
|
|
|
|
| Sample: |
Beta-amylase 2, chloroplastic tetramer, 229 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Jun 11
|
Solution structure and assembly of β-amylase2 from Arabidopsis thaliana
(2019)
Chandrasekharan N, Ravenburg C, Roy I, Monroe J, Berndsen C
|
| RgGuinier |
4.2 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
308 |
nm3 |
|
|
UniProt ID: O65258 (86-542) Beta-amylase 2, chloroplastic
|
|
|
|
| Sample: |
Beta-amylase 2, chloroplastic tetramer, 215 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Jun 11
|
Solution structure and assembly of β-amylase2 from Arabidopsis thaliana
(2019)
Chandrasekharan N, Ravenburg C, Roy I, Monroe J, Berndsen C
|
| RgGuinier |
4.4 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
272 |
nm3 |
|
|
UniProt ID: P32849 (2-1169) DNA repair protein RAD5
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG25_fit1_model1.png)
|
| Sample: |
DNA repair protein RAD5 monomer, 138 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
40 mM Tris, 150 mM KCl, 5 mM DTT, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Mar 6
|
Conformational flexibility of fork-remodeling helicase Rad5 shown by full-ensemble hybrid methods.
PLoS One 14(10):e0223875 (2019)
Gildenberg MS, Washington MT
|
| RgGuinier |
4.7 |
nm |
| Dmax |
17.8 |
nm |
| VolumePorod |
254 |
nm3 |
|
|
UniProt ID: Q15796 (10-174) Mothers against decapentaplegic homolog 2
|
|
|
|
| Sample: |
Mothers against decapentaplegic homolog 2 monomer, 19 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jul 2
|
Structural basis for distinct roles of SMAD2 and SMAD3 in FOXH1 pioneer-directed TGF-β signaling.
Genes Dev 33(21-22):1506-1524 (2019)
Aragón E, Wang Q, Zou Y, Morgani SM, Ruiz L, Kaczmarska Z, Su J, Torner C, Tian L, Hu J, Shu W, Agrawal S, Gomes T, Márquez JA, Hadjantonakis AK, Macias MJ, Massagué J
|
| RgGuinier |
1.9 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
UniProt ID: Q15796-2 (10-143) Mothers against decapentaplegic homolog 2
|
|
|
|
| Sample: |
Mothers against decapentaplegic homolog 2 monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jul 2
|
Structural basis for distinct roles of SMAD2 and SMAD3 in FOXH1 pioneer-directed TGF-β signaling.
Genes Dev 33(21-22):1506-1524 (2019)
Aragón E, Wang Q, Zou Y, Morgani SM, Ruiz L, Kaczmarska Z, Su J, Torner C, Tian L, Hu J, Shu W, Agrawal S, Gomes T, Márquez JA, Hadjantonakis AK, Macias MJ, Massagué J
|
| RgGuinier |
1.7 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
UniProt ID: P01137 (30-277) Latency associated peptide
|
|
|
|
| Sample: |
Latency associated peptide dimer, 58 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Apr 20
|
Structural insights into conformational switching in latency-associated peptide between transforming growth factor β-1 bound and unbound states
IUCrJ 7(2) (2020)
Stachowski T, Snell M, Snell E
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
129 |
nm3 |
|
|
UniProt ID: O00482 (1-541) Liver Receptor Homolog-1
UniProt ID: None (None-None) Peroxisome Proliferator-Activated Receptor Gamma Coactivator-1 Alpha
UniProt ID: None (None-None) CYP7A1 Promoter Forward
UniProt ID: None (None-None) CYP7A1 Promoter Reverse
|
|
|
|
| Sample: |
Liver Receptor Homolog-1 monomer, 64 kDa Homo sapiens protein
Peroxisome Proliferator-Activated Receptor Gamma Coactivator-1 Alpha monomer, 2 kDa Homo sapiens protein
CYP7A1 Promoter Forward monomer, 4 kDa Homo sapiens DNA
CYP7A1 Promoter Reverse monomer, 4 kDa Homo sapiens DNA
|
| Buffer: |
20 mM TRIS, 150 mM NaCl, 2% v/v glycerol, 0.5 mM CHAPS, 5 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 26
|
Integrated Structural Modeling of Full-Length LRH-1 Reveals Inter-domain Interactions Contribute to Receptor Structure and Function.
Structure (2020)
Seacrist CD, Kuenze G, Hoffmann RM, Moeller BE, Burke JE, Meiler J, Blind RD
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
76 |
nm3 |
|
|