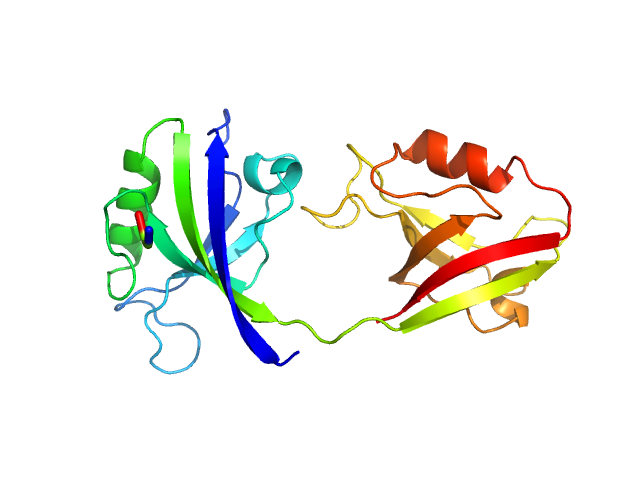
UniProt ID: P78352 (55-249) PDZ1-2 fragment of PSD-95/Disks large homolog 4
|
|
|
|
| Sample: |
PDZ1-2 fragment of PSD-95/Disks large homolog 4 monomer, 21 kDa Homo sapiens protein
|
| Buffer: |
20 mM TRIS/HCl, 150 mM NaCl + 10 mM GSH, pH: 8.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Jan 26
|
The dual PDZ domain from Postsynaptic density protein 95 forms a scaffold with peptide ligand
Biophysical Journal (2020)
Rodzli N, Lockhart-Cairns M, Levy C, Chipperfield J, Bird L, Baldock C, Prince S
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.2 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
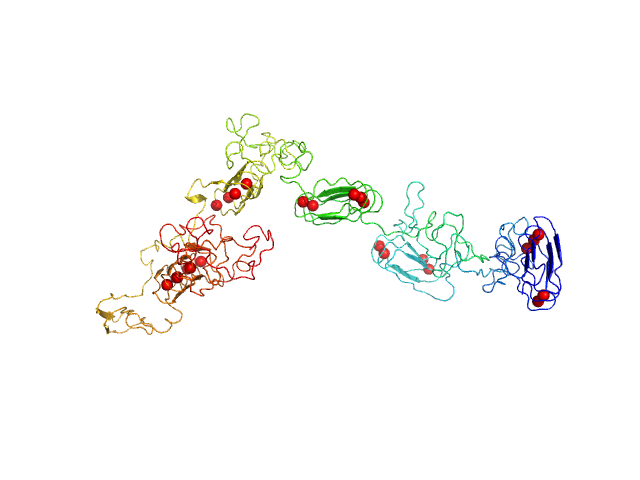
UniProt ID: P0DKX7 (1007-1706) RD domain of B. Pertussis Adenylate Cyclase Toxin (CyaA)
|
|
|
|
| Sample: |
RD domain of B. Pertussis Adenylate Cyclase Toxin (CyaA) monomer, 73 kDa Bordetella pertussis protein
|
| Buffer: |
20 mM Hepes, 150 mM NaCl, 2 mM DTT, 4 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 May 31
|
Structural models of intrinsically disordered and calcium-bound folded states of a protein adapted for secretion.
Sci Rep 5:14223 (2015)
O'Brien DP, Hernandez B, Durand D, Hourdel V, Sotomayor-Pérez AC, Vachette P, Ghomi M, Chamot-Rooke J, Ladant D, Brier S, Chenal A
|
| RgGuinier |
4.4 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
89 |
nm3 |
|
|
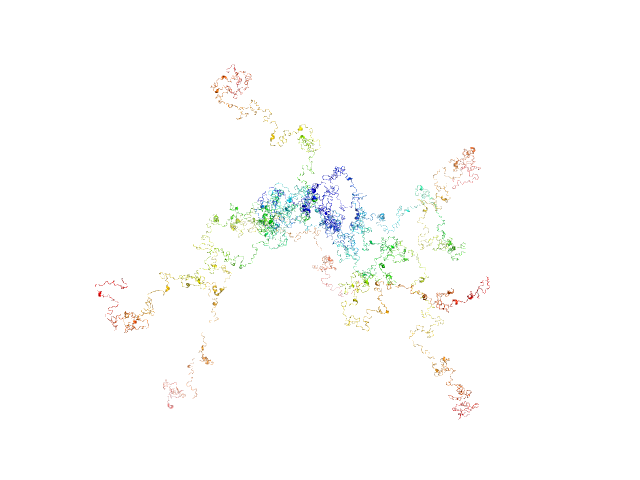
UniProt ID: P0DKX7 (1007-1706) RD domain of B. Pertussis Adenylate Cyclase Toxin (CyaA)
|
|
|
|
| Sample: |
RD domain of B. Pertussis Adenylate Cyclase Toxin (CyaA) monomer, 73 kDa Bordetella pertussis protein
|
| Buffer: |
20 mM Hepes, 150 mM NaCl, 2 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 May 31
|
Structural models of intrinsically disordered and calcium-bound folded states of a protein adapted for secretion.
Sci Rep 5:14223 (2015)
O'Brien DP, Hernandez B, Durand D, Hourdel V, Sotomayor-Pérez AC, Vachette P, Ghomi M, Chamot-Rooke J, Ladant D, Brier S, Chenal A
|
| RgGuinier |
8.3 |
nm |
| Dmax |
33.0 |
nm |
|
|
UniProt ID: Q9KPH3 (1-157) DciA
|
|
|
|
| Sample: |
DciA monomer, 18 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM Tris-HCl, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Mar 26
|
Study of the DnaB:DciA interplay reveals insights into the primary mode of loading of the bacterial replicative helicase.
Nucleic Acids Res (2021)
Marsin S, Adam Y, Cargemel C, Andreani J, Baconnais S, Legrand P, Li de la Sierra-Gallay I, Humbert A, Aumont-Nicaise M, Velours C, Ochsenbein F, Durand D, Le Cam E, Walbott H, Possoz C, Quevillon-Cheruel S, Ferat JL
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
UniProt ID: A0A0H3AIB6 (1-468) Replicative DNA helicase (DnaB)
|
|
|
|
| Sample: |
Replicative DNA helicase (DnaB) hexamer, 317 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM Tris-HCl, 100 mM NaCl, 1 mM ATP, pH: 8.8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Oct 4
|
Study of the DnaB:DciA interplay reveals insights into the primary mode of loading of the bacterial replicative helicase.
Nucleic Acids Res (2021)
Marsin S, Adam Y, Cargemel C, Andreani J, Baconnais S, Legrand P, Li de la Sierra-Gallay I, Humbert A, Aumont-Nicaise M, Velours C, Ochsenbein F, Durand D, Le Cam E, Walbott H, Possoz C, Quevillon-Cheruel S, Ferat JL
|
| RgGuinier |
4.6 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
585 |
nm3 |
|
|
UniProt ID: A0A0H3AIB6 (1-468) DnaB helicase complexed with ATP
UniProt ID: Q9KPH3 (1-157) DciA
|
|
|
|
| Sample: |
DnaB helicase complexed with ATP hexamer, 317 kDa Vibrio cholerae serotype … protein
DciA, 55 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM Tris-HCl, 100 mM NaCl, 1 mM ATP, pH: 8.8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Oct 4
|
Study of the DnaB:DciA interplay reveals insights into the primary mode of loading of the bacterial replicative helicase.
Nucleic Acids Res (2021)
Marsin S, Adam Y, Cargemel C, Andreani J, Baconnais S, Legrand P, Li de la Sierra-Gallay I, Humbert A, Aumont-Nicaise M, Velours C, Ochsenbein F, Durand D, Le Cam E, Walbott H, Possoz C, Quevillon-Cheruel S, Ferat JL
|
| RgGuinier |
5.0 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
670 |
nm3 |
|
|
UniProt ID: Q9HW86 (1-124) MvaT(mutant)
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDGS5_fit1_model1.png)
|
| Sample: |
MvaT(mutant) dimer, 28 kDa Pseudomonas aeruginosa protein
|
| Buffer: |
20 mM Bis-Tris 50 mM KCl, pH: 6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 11
|
Structural basis for osmotic regulation of the DNA binding properties of H-NS proteins.
Nucleic Acids Res (2020)
Qin L, Bdira FB, Sterckx YGJ, Volkov AN, Vreede J, Giachin G, van Schaik P, Ubbink M, Dame RT
|
| RgGuinier |
3.6 |
nm |
| Dmax |
14.7 |
nm |
| VolumePorod |
47 |
nm3 |
|
|
UniProt ID: Q9HW86 (1-124) MvaT(mutant)
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDGT5_fit1_model1.png)
|
| Sample: |
MvaT(mutant) dimer, 28 kDa Pseudomonas aeruginosa protein
|
| Buffer: |
20 mM Bis-Tris 300 mM KCl, pH: 6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 11
|
Structural basis for osmotic regulation of the DNA binding properties of H-NS proteins.
Nucleic Acids Res (2020)
Qin L, Bdira FB, Sterckx YGJ, Volkov AN, Vreede J, Giachin G, van Schaik P, Ubbink M, Dame RT
|
| RgGuinier |
3.8 |
nm |
| Dmax |
15.8 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
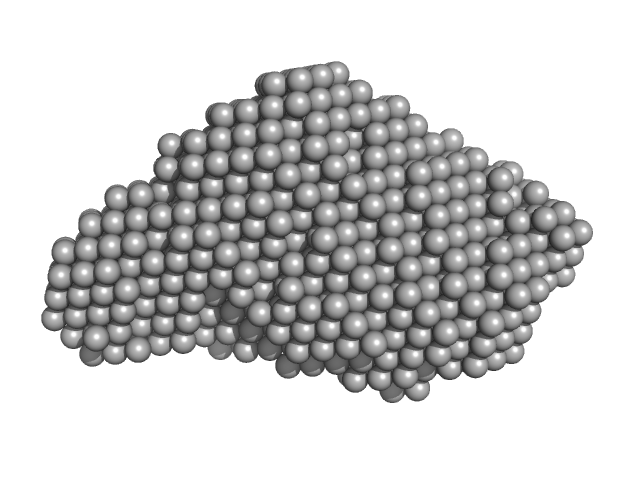
UniProt ID: Q52209 (110-324) PupR protein
UniProt ID: P38047 (None-None) Ferric-pseudobactin BN7/BN8 receptor
|
|
|

|
| Sample: |
PupR protein monomer, 24 kDa Pseudomonas putida protein
Ferric-pseudobactin BN7/BN8 receptor monomer, 8 kDa Pseudomonas putida protein
|
| Buffer: |
25 mM HEPES 400 mM LiCl 10% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Mar 16
|
Structural basis of cell surface signaling by a conserved sigma regulator in Gram-negative bacteria.
J Biol Chem (2020)
Jensen JL, Jernberg BD, Sinha S, Colbert CL
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.7 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: P60752 (338-582) Lipid A export ATP-binding/permease protein MsbA - Nucleotide binding domain
|
|
|
|
| Sample: |
Lipid A export ATP-binding/permease protein MsbA - Nucleotide binding domain monomer, 27 kDa Escherichia coli protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 5 mM MgCl2, 0.45 mM Mg2+-ATP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 8
|
Structural Kinetics of MsbA Investigated by Stopped-Flow Time-Resolved Small-Angle X-Ray Scattering.
Structure (2019)
Josts I, Gao Y, Monteiro DCF, Niebling S, Nitsche J, Veith K, Gräwert TW, Blanchet CE, Schroer MA, Huse N, Pearson AR, Svergun DI, Tidow H
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.8 |
nm |
| VolumePorod |
50 |
nm3 |
|
|