UniProt ID: Q9VG38 (1-468) Suppressor of fused homolog
|
|
|
|
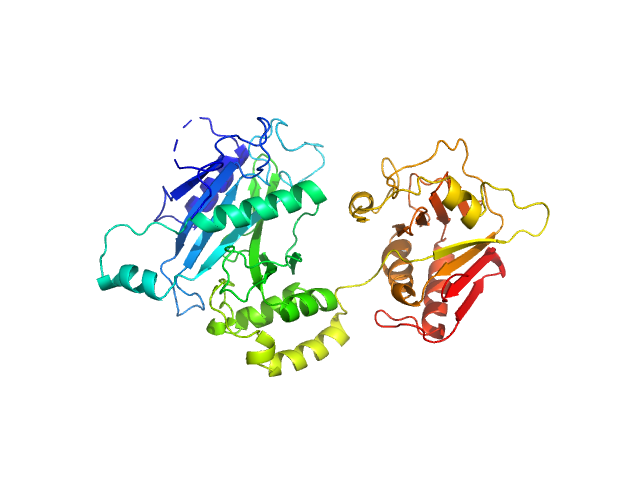
| Sample: |
Suppressor of fused homolog monomer, 53 kDa Drosophila melanogaster protein
|
| Buffer: |
50 mM bis-TRIS pH 5.5, 200 mM NaCl, 10% glycerol, pH: 5.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 May 7
|
Suppressor of Fused
Valerie Biou
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
12 |
nm3 |
|
|
UniProt ID: Q9UMX1 (None-None) Suppressor of fused homolog
|
|
|
|
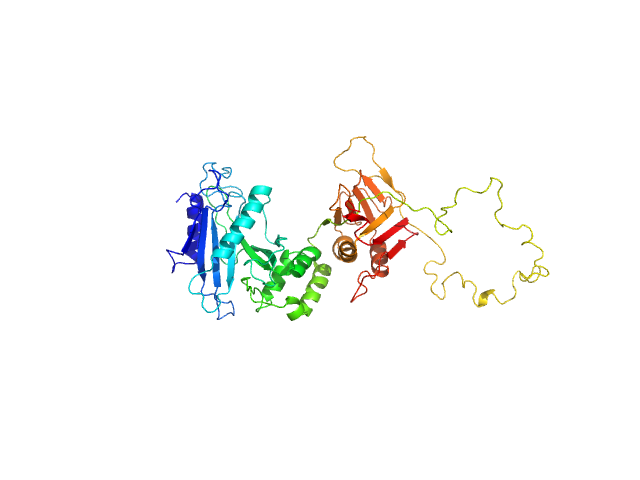
| Sample: |
Suppressor of fused homolog monomer, 51 kDa Homo sapiens protein
|
| Buffer: |
50 mM bis-TRIS pH 5.5, 200 mM NaCl, 10% glycerol, pH: 5.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Jun 30
|
Suppressor of Fused
Valerie Biou
|
| RgGuinier |
3.2 |
nm |
| Dmax |
14.0 |
nm |
|
|
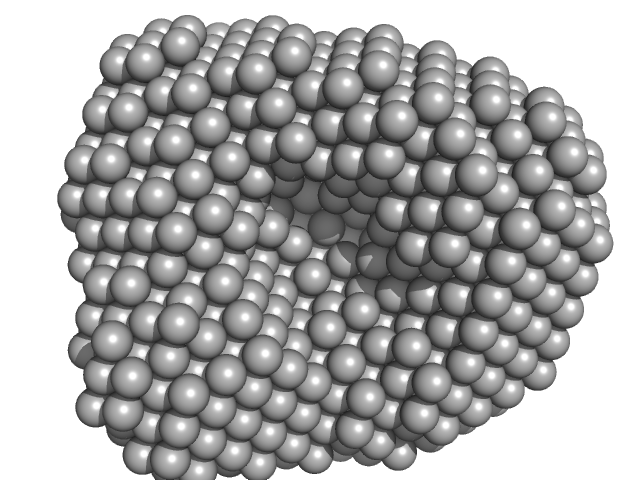
UniProt ID: Q9PPB4 (1-298) 4-hydroxy-tetrahydrodipicolinate synthase
|
|
|

|
| Sample: |
4-hydroxy-tetrahydrodipicolinate synthase tetramer, 131 kDa Campylobacter jejuni protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Aug 2
|
Asparagine-84, a regulatory allosteric site residue, helps maintain the quaternary structure of Campylobacter jejuni dihydrodipicolinate synthase.
J Struct Biol :107409 (2019)
Majdi Yazdi M, Saran S, Mrozowich T, Lehnert C, Patel TR, Sanders DAR, Palmer DRJ
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
188 |
nm3 |
|
|
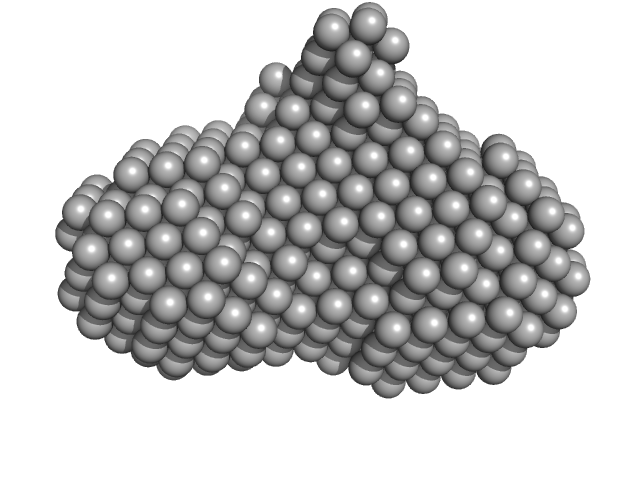
UniProt ID: Q9PPB4 (1-298) 4-hydroxy-tetrahydrodipicolinate synthase (N84D mutant)
|
|
|

|
| Sample: |
4-hydroxy-tetrahydrodipicolinate synthase (N84D mutant) dimer, 65 kDa Campylobacter jejuni protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Aug 2
|
Asparagine-84, a regulatory allosteric site residue, helps maintain the quaternary structure of Campylobacter jejuni dihydrodipicolinate synthase.
J Struct Biol :107409 (2019)
Majdi Yazdi M, Saran S, Mrozowich T, Lehnert C, Patel TR, Sanders DAR, Palmer DRJ
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
108 |
nm3 |
|
|
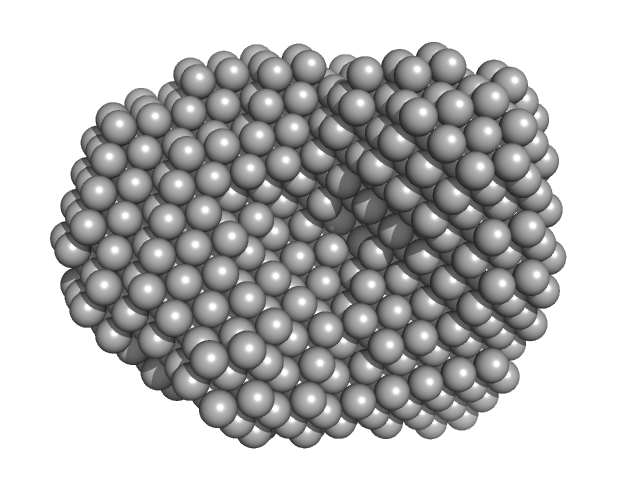
UniProt ID: Q9PPB4 (1-298) 4-hydroxy-tetrahydrodipicolinate synthase (N84A mutant)
|
|
|

|
| Sample: |
4-hydroxy-tetrahydrodipicolinate synthase (N84A mutant), 33 kDa Campylobacter jejuni protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Aug 2
|
Asparagine-84, a regulatory allosteric site residue, helps maintain the quaternary structure of Campylobacter jejuni dihydrodipicolinate synthase.
J Struct Biol :107409 (2019)
Majdi Yazdi M, Saran S, Mrozowich T, Lehnert C, Patel TR, Sanders DAR, Palmer DRJ
|
| RgGuinier |
3.3 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
163 |
nm3 |
|
|
UniProt ID: Q5E9J8 (1-492) Resistance to inhibitors of cholinesterase 8 homolog A
UniProt ID: None (None-None) miniGi
|
|
|
|
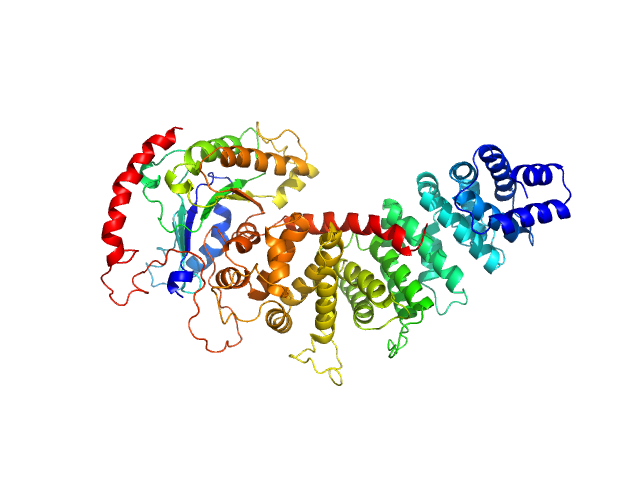
| Sample: |
Resistance to inhibitors of cholinesterase 8 homolog A monomer, 56 kDa Bos taurus protein
MiniGi monomer, 25 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 5 % glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Oct 27
|
Large-scale conformational rearrangement of the α5-helix of Gα subunits in complex with the guanine nucleotide exchange factor Ric8A.
J Biol Chem (2019)
Srivastava D, Artemyev NO
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.7 |
nm |
|
|
UniProt ID: A0A0A7EQR3 (18-474) 4-O-methyl-glucuronoyl methylesterase (Glucuronoyl esterase)
|
|
|
|
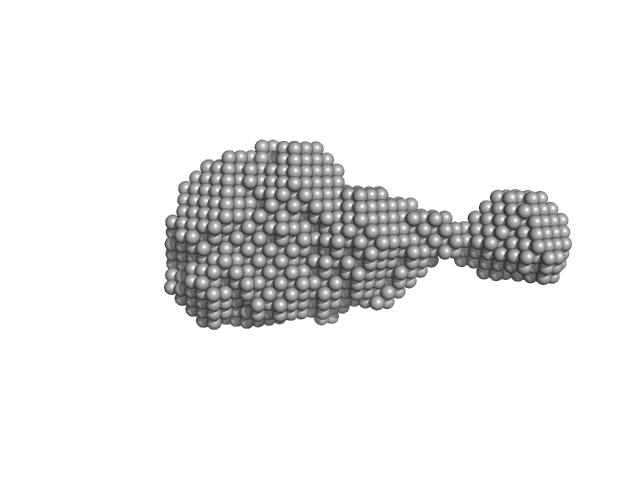
| Sample: |
4-O-methyl-glucuronoyl methylesterase (Glucuronoyl esterase) monomer, 51 kDa Cerrena unicolor protein
|
| Buffer: |
20 mM sodium acetate, pH: 5 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with GeniX3D, University of Copenhagen, Department of Drug Design and Pharmacology on 2018 Oct 10
|
The structural basis of fungal glucuronoyl esterase activity on natural substrates.
Nat Commun 11(1):1026 (2020)
Ernst HA, Mosbech C, Langkilde AE, Westh P, Meyer AS, Agger JW, Larsen S
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
71 |
nm3 |
|
|
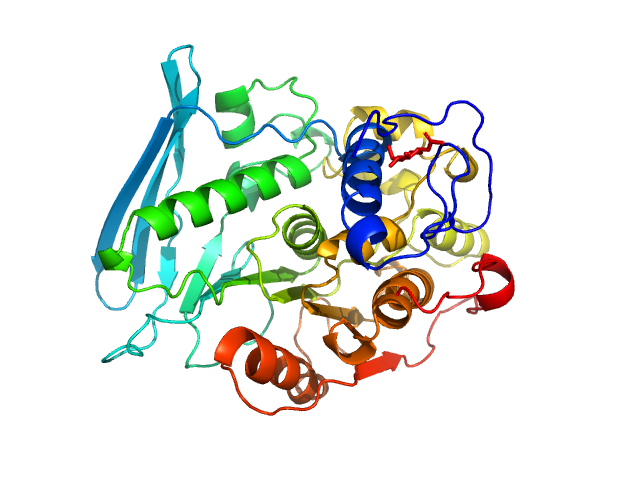
UniProt ID: A0A0A7EQR3 (95-474) 4-O-methyl-glucuronoyl methylesterase (Glucuronoyl esterase, truncated)
|
|
|
|
| Sample: |
4-O-methyl-glucuronoyl methylesterase (Glucuronoyl esterase, truncated) monomer, 43 kDa Cerrena unicolor protein
|
| Buffer: |
20 mM sodium acetate, pH: 5 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with GeniX3D, University of Copenhagen, Department of Drug Design and Pharmacology on 2018 Oct 10
|
The structural basis of fungal glucuronoyl esterase activity on natural substrates.
Nat Commun 11(1):1026 (2020)
Ernst HA, Mosbech C, Langkilde AE, Westh P, Meyer AS, Agger JW, Larsen S
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.1 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
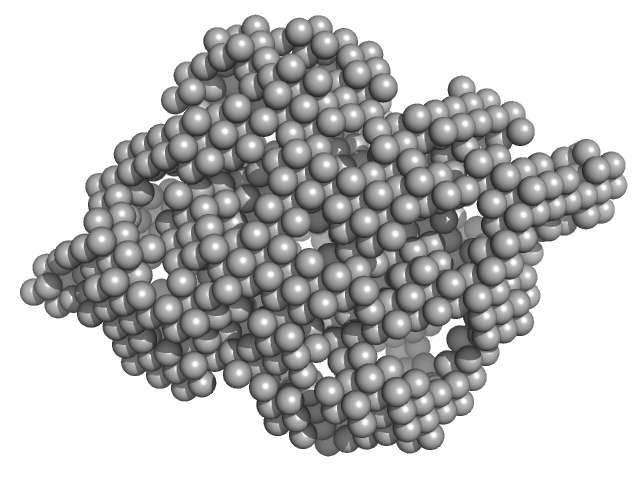
UniProt ID: P0ABK5 (1-323) Cysteine synthase A
|
|
|
|
| Sample: |
Cysteine synthase A dimer, 71 kDa Escherichia coli protein
|
| Buffer: |
20 mM sodium phosphate, 85 mM NaCl, 2 mM EDTA, 10 mM 2-MCE, pH: 7.5 |
| Experiment: |
SAXS
data collected at Austrian SAXS beamline 5.2L, ELETTRA on 2016 Jun 1
|
Combination of SAXS and Protein Painting Discloses the Three-Dimensional Organization of the Bacterial Cysteine Synthase Complex, a Potential Target for Enhancers of Antibiotic Action.
Int J Mol Sci 20(20) (2019)
Rosa B, Marchetti M, Paredi G, Amenitsch H, Franko N, Benoni R, Giabbai B, De Marino MG, Mozzarelli A, Ronda L, Storici P, Campanini B, Bettati S
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
108 |
nm3 |
|
|
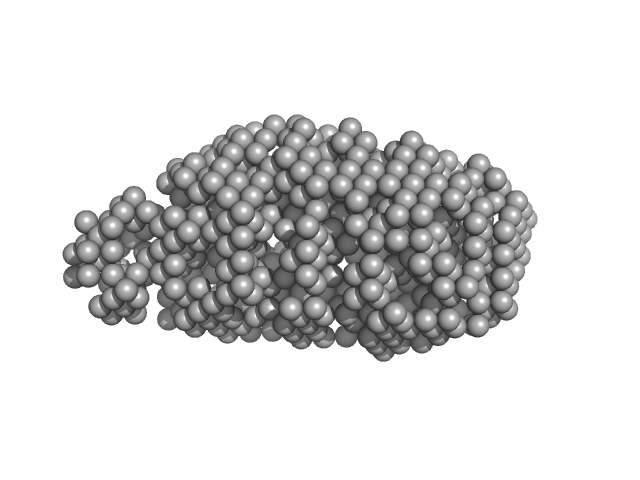
UniProt ID: P0A9D4 (2-273) Serine acetyltransferase
|
|
|
|
| Sample: |
Serine acetyltransferase hexamer, 177 kDa Escherichia coli protein
|
| Buffer: |
20 mM sodium phosphate, 85 mM NaCl, 2 mM EDTA, 10 mM 2-MCE, pH: 7.5 |
| Experiment: |
SAXS
data collected at Austrian SAXS beamline 5.2L, ELETTRA on 2016 Jun 1
|
Combination of SAXS and Protein Painting Discloses the Three-Dimensional Organization of the Bacterial Cysteine Synthase Complex, a Potential Target for Enhancers of Antibiotic Action.
Int J Mol Sci 20(20) (2019)
Rosa B, Marchetti M, Paredi G, Amenitsch H, Franko N, Benoni R, Giabbai B, De Marino MG, Mozzarelli A, Ronda L, Storici P, Campanini B, Bettati S
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
280 |
nm3 |
|
|