UniProt ID: P0ABK5 (1-323) Cysteine synthase A (4-mer)
UniProt ID: P0A9D4 (2-273) Serine acetyltransferase (6-mer)
|
|
|

|
| Sample: |
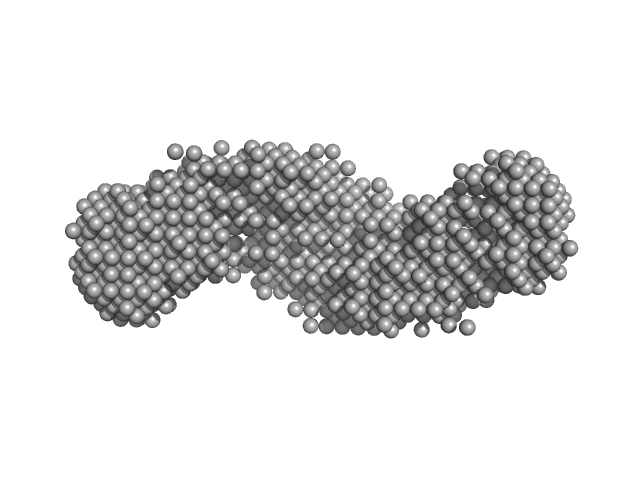
Cysteine synthase A (4-mer) tetramer, 143 kDa Escherichia coli protein
Serine acetyltransferase (6-mer) hexamer, 177 kDa Escherichia coli protein
|
| Buffer: |
20 mM sodium phosphate, 85 mM NaCl, 2 mM EDTA, 10 mM 2-MCE, pH: 7.5 |
| Experiment: |
SAXS
data collected at Austrian SAXS beamline 5.2L, ELETTRA on 2016 Jun 1
|
Combination of SAXS and Protein Painting Discloses the Three-Dimensional Organization of the Bacterial Cysteine Synthase Complex, a Potential Target for Enhancers of Antibiotic Action.
Int J Mol Sci 20(20) (2019)
Rosa B, Marchetti M, Paredi G, Amenitsch H, Franko N, Benoni R, Giabbai B, De Marino MG, Mozzarelli A, Ronda L, Storici P, Campanini B, Bettati S
|
| RgGuinier |
6.1 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
457 |
nm3 |
|
|
UniProt ID: A2WPN7 (1-145) Salt stress-induced protein
|
|
|
|
| Sample: |
Salt stress-induced protein dimer, 37 kDa Oryza sativa Indica … protein
|
| Buffer: |
50 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2017 Mar 6
|
Structural insights into rice SalTol QTL located SALT protein.
Sci Rep 10(1):16589 (2020)
Kaur N, Sagar A, Sharma P, Ashish, Pati PK
|
| RgGuinier |
2.5 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: A2WPN7 (1-145) Salt stress-induced protein
|
|
|
|
| Sample: |
Salt stress-induced protein dimer, 37 kDa Oryza sativa Indica … protein
|
| Buffer: |
50 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2017 Mar 6
|
Structural insights into rice SalTol QTL located SALT protein.
Sci Rep 10(1):16589 (2020)
Kaur N, Sagar A, Sharma P, Ashish, Pati PK
|
| RgGuinier |
2.4 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
UniProt ID: A2WPN7 (1-145) Salt stress-induced protein
|
|
|
|
| Sample: |
Salt stress-induced protein dimer, 37 kDa Oryza sativa Indica … protein
|
| Buffer: |
50 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2017 Mar 6
|
Structural insights into rice SalTol QTL located SALT protein.
Sci Rep 10(1):16589 (2020)
Kaur N, Sagar A, Sharma P, Ashish, Pati PK
|
| RgGuinier |
2.5 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
UniProt ID: A0A110B4Q7 (1-296) Tegument protein UL7
UniProt ID: D3YPL0 (8-142) Tegument protein UL51
|
|
|
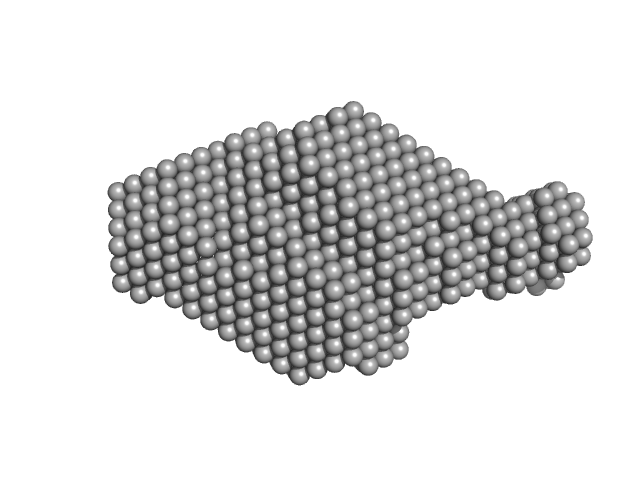

|
| Sample: |
Tegument protein UL7 monomer, 34 kDa Human alphaherpesvirus 1 … protein
Tegument protein UL51 dimer, 30 kDa Human alphaherpesvirus 1 … protein
|
| Buffer: |
20 mM tris, 200 mM NaCl, 3% (v/v) glycerol, 0.25 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 May 16
|
Insights into herpesvirus assembly from the structure of the pUL7:pUL51 complex.
Elife 9 (2020)
Butt BG, Owen DJ, Jeffries CM, Ivanova L, Hill CH, Houghton JW, Ahmed MF, Antrobus R, Svergun DI, Welch JJ, Crump CM, Graham SC
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
116 |
nm3 |
|
|
UniProt ID: A0A110B4Q7 (1-296) Tegument protein UL7
UniProt ID: D3YPL0 (1-244) Tegument protein UL51
|
|
|
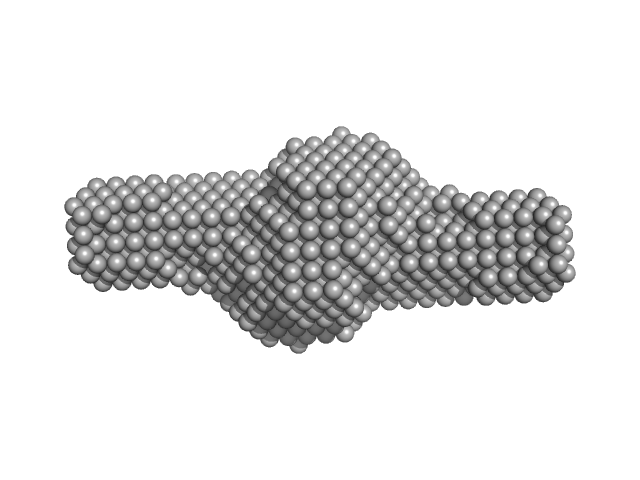

|
| Sample: |
Tegument protein UL7 dimer, 68 kDa Human alphaherpesvirus 1 … protein
Tegument protein UL51 tetramer, 102 kDa Human alphaherpesvirus 1 … protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 3% (v/v) glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 May 16
|
Insights into herpesvirus assembly from the structure of the pUL7:pUL51 complex.
Elife 9 (2020)
Butt BG, Owen DJ, Jeffries CM, Ivanova L, Hill CH, Houghton JW, Ahmed MF, Antrobus R, Svergun DI, Welch JJ, Crump CM, Graham SC
|
| RgGuinier |
4.6 |
nm |
| Dmax |
19.7 |
nm |
| VolumePorod |
340 |
nm3 |
|
|
UniProt ID: A0A110B4Q7 (1-296) Tegument protein UL7
UniProt ID: D3YPL0 (1-244) Tegument protein UL51
|
|
|
|
| Sample: |
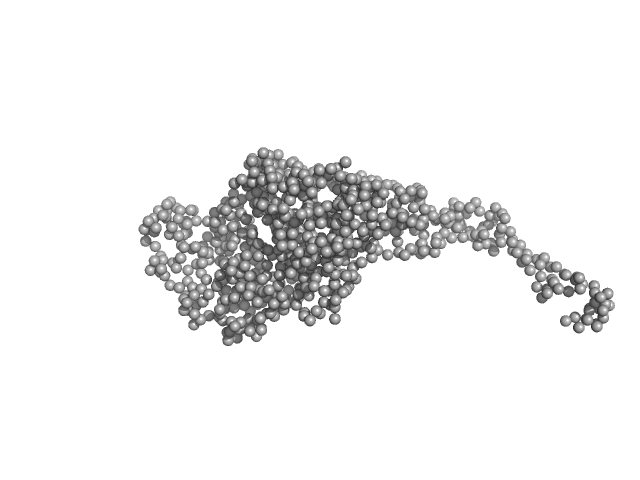
Tegument protein UL7 monomer, 34 kDa Human alphaherpesvirus 1 … protein
Tegument protein UL51 dimer, 51 kDa Human alphaherpesvirus 1 protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 3% (v/v) glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 May 16
|
Insights into herpesvirus assembly from the structure of the pUL7:pUL51 complex.
Elife 9 (2020)
Butt BG, Owen DJ, Jeffries CM, Ivanova L, Hill CH, Houghton JW, Ahmed MF, Antrobus R, Svergun DI, Welch JJ, Crump CM, Graham SC
|
| RgGuinier |
4.0 |
nm |
| Dmax |
18.2 |
nm |
| VolumePorod |
160 |
nm3 |
|
|
UniProt ID: Q587D3 (26-358) Trypanosoma brucei Membrane Occupation and Recognition Nexus MORN repeats 2-15
|
|
|
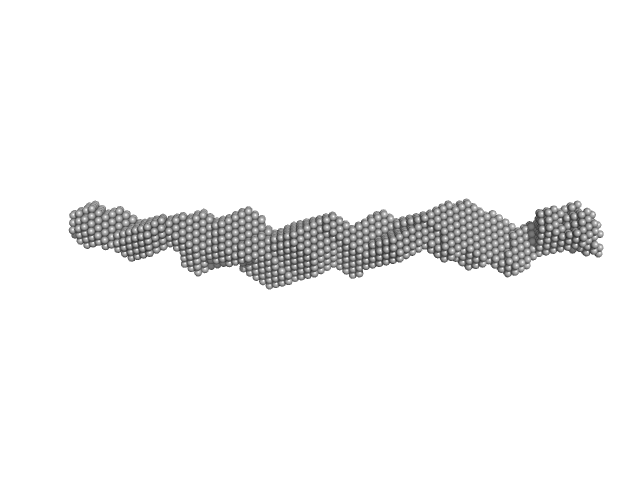

|
| Sample: |
Trypanosoma brucei Membrane Occupation and Recognition Nexus MORN repeats 2-15 dimer, 78 kDa Trypanosoma brucei protein
|
| Buffer: |
20 mM Tris-HCl, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jan 27
|
Structures of three MORN repeat proteins and a re-evaluation of the proposed lipid-binding properties of MORN repeats.
PLoS One 15(12):e0242677 (2020)
Sajko S, Grishkovskaya I, Kostan J, Graewert M, Setiawan K, Trübestein L, Niedermüller K, Gehin C, Sponga A, Puchinger M, Gavin AC, Leonard TA, Svergun DI, Smith TK, Morriswood B, Djinovic-Carugo K
|
| RgGuinier |
6.5 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
187 |
nm3 |
|
|
UniProt ID: Q587D3 (143-358) Trypanosoma brucei Membrane Occupation and Recognition Nexus MORN (7-15)
|
|
|
|
| Sample: |
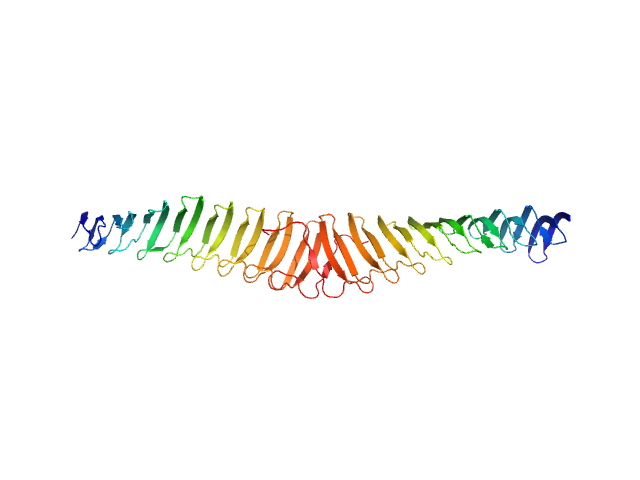
Trypanosoma brucei Membrane Occupation and Recognition Nexus MORN (7-15) dimer, 52 kDa Trypanosoma brucei protein
|
| Buffer: |
20 mM Tris-HCl, 200 mM NaCl, 2% (w/v) glycerol, 0.5 mM DTT, pH: 8.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jun 11
|
Structures of three MORN repeat proteins and a re-evaluation of the proposed lipid-binding properties of MORN repeats.
PLoS One 15(12):e0242677 (2020)
Sajko S, Grishkovskaya I, Kostan J, Graewert M, Setiawan K, Trübestein L, Niedermüller K, Gehin C, Sponga A, Puchinger M, Gavin AC, Leonard TA, Svergun DI, Smith TK, Morriswood B, Djinovic-Carugo K
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: A0A0F7VBC6 (48-363) Toxoplasma gondii Membrane Occupation and Recognition Nexus (MORN1) repeats 7-15
|
|
|
|
| Sample: |
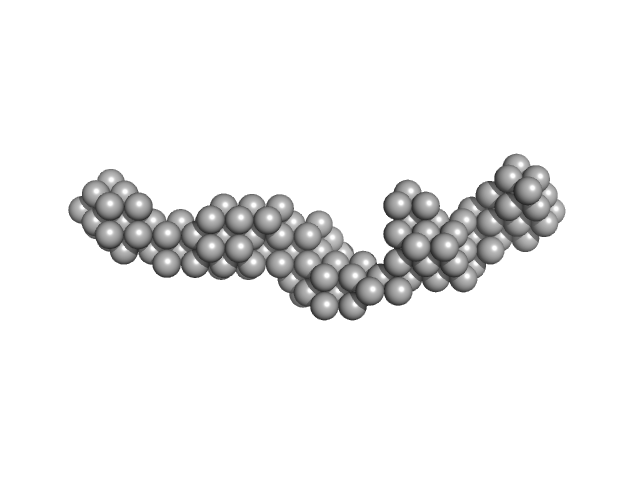
Toxoplasma gondii Membrane Occupation and Recognition Nexus (MORN1) repeats 7-15 dimer, 49 kDa Toxoplasma gondii protein
|
| Buffer: |
20 mM Tris-HCl, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jun 10
|
Structures of three MORN repeat proteins and a re-evaluation of the proposed lipid-binding properties of MORN repeats.
PLoS One 15(12):e0242677 (2020)
Sajko S, Grishkovskaya I, Kostan J, Graewert M, Setiawan K, Trübestein L, Niedermüller K, Gehin C, Sponga A, Puchinger M, Gavin AC, Leonard TA, Svergun DI, Smith TK, Morriswood B, Djinovic-Carugo K
|
| RgGuinier |
4.0 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
55 |
nm3 |
|
|