UniProt ID: P20936 (174-1047) Ras GTPase-activating protein 1
UniProt ID: Q9NRY4 (1083-1111) Rho GTPase-activating protein 35
|
|
|

|
| Sample: |
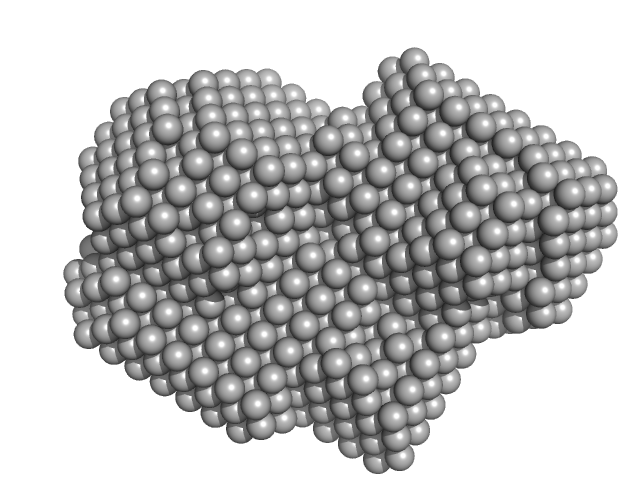
Ras GTPase-activating protein 1 monomer, 101 kDa Homo sapiens protein
Rho GTPase-activating protein 35 monomer, 3 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8 350 mM NaCl 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 11
|
Tandem engagement of phosphotyrosines by the dual SH2 domains of p120RasGAP.
Structure (2022)
Stiegler AL, Vish KJ, Boggon TJ
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
41 |
nm3 |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|

|
| Sample: |
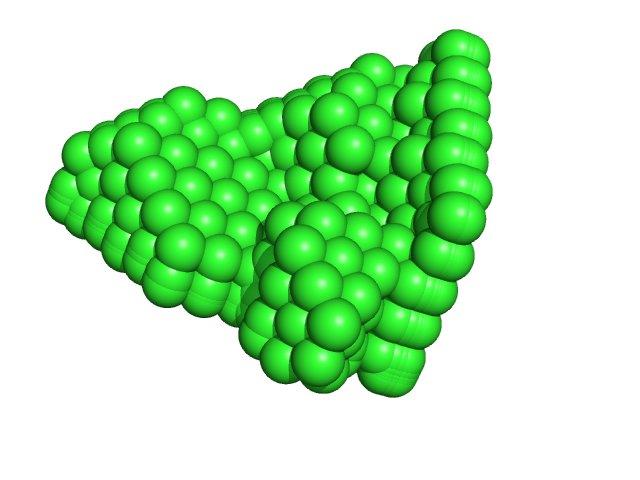
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.9 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
UniProt ID: Q00511 (2-302) Uricase
|
|
|

|
| Sample: |
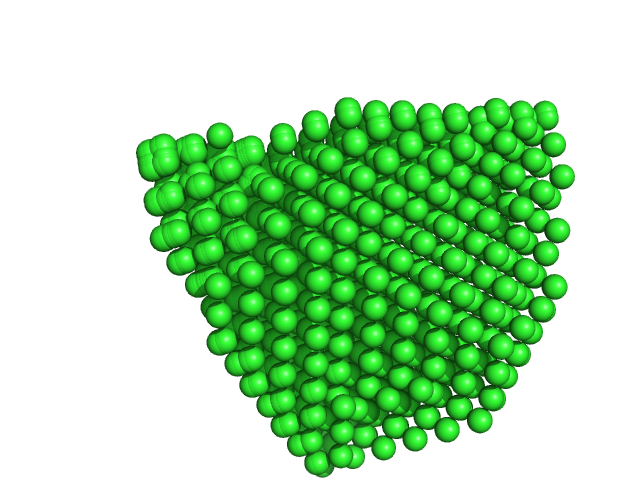
Uricase tetramer, 136 kDa Aspergillus flavus protein
|
| Buffer: |
100 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
220 |
nm3 |
|
|
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|

|
| Sample: |
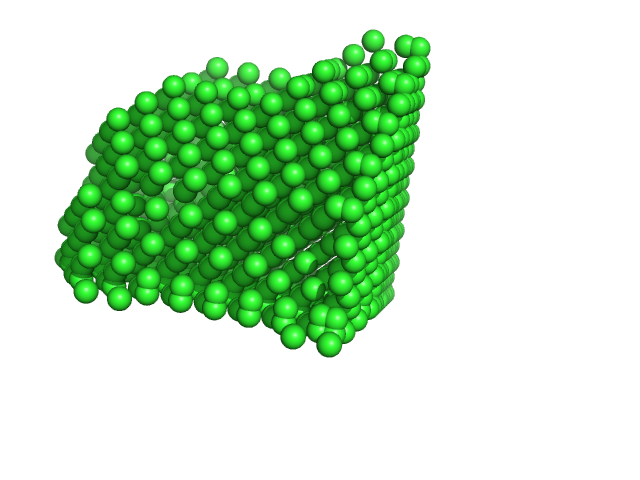
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
ConsensusBuffer_50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
243 |
nm3 |
|
|
UniProt ID: F8W669 (1-190) Endo-1,4-beta-xylanase
|
|
|

|
| Sample: |
Endo-1,4-beta-xylanase monomer, 21 kDa Trichoderma longibrachiatum protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.1 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
UniProt ID: P00698 (19-147) Lysozyme C
|
|
|

|
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
50 mM sodium citrate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.8 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
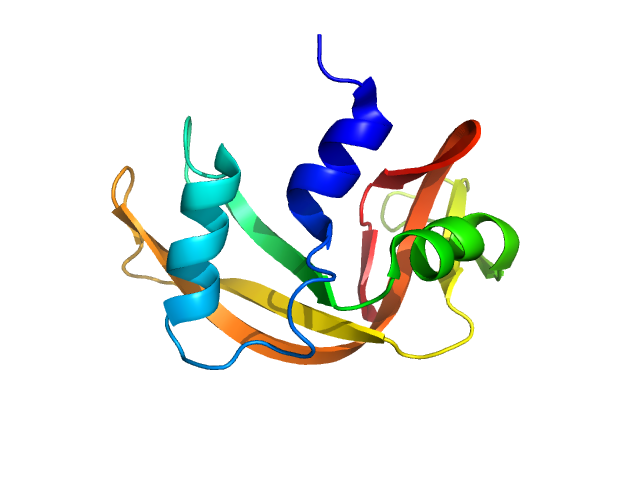
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|
|
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.4 |
nm |
|
|
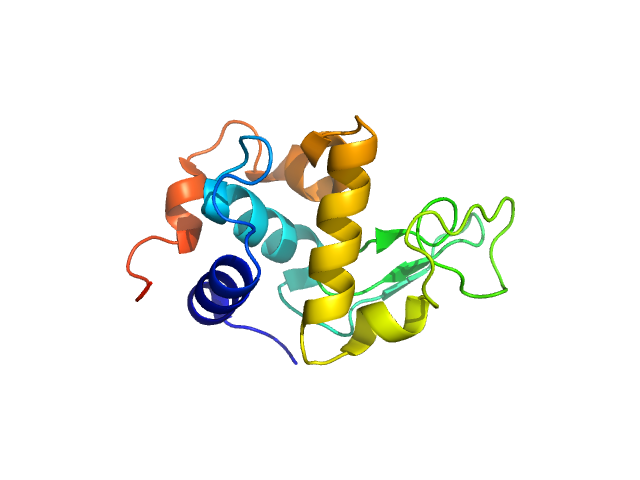
UniProt ID: P00698 (19-147) Lysozyme C
|
|
|
|
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
50 mM sodium citrate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.2 |
nm |
| Dmax |
3.8 |
nm |
|
|
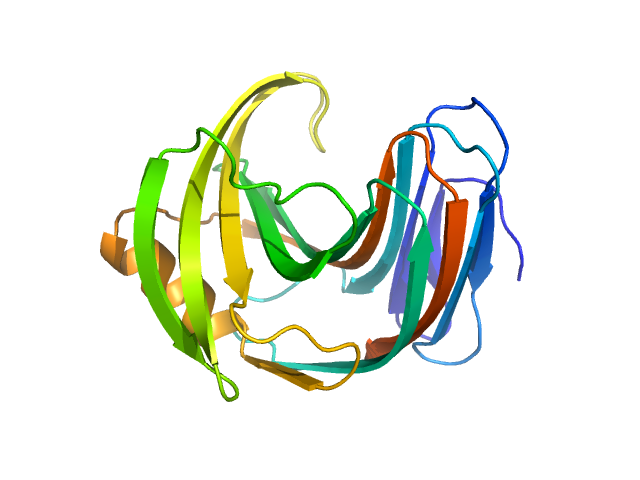
UniProt ID: F8W669 (1-190) Endo-1,4-beta-xylanase
|
|
|
|
| Sample: |
Endo-1,4-beta-xylanase monomer, 21 kDa Trichoderma longibrachiatum protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.4 |
nm |
|
|
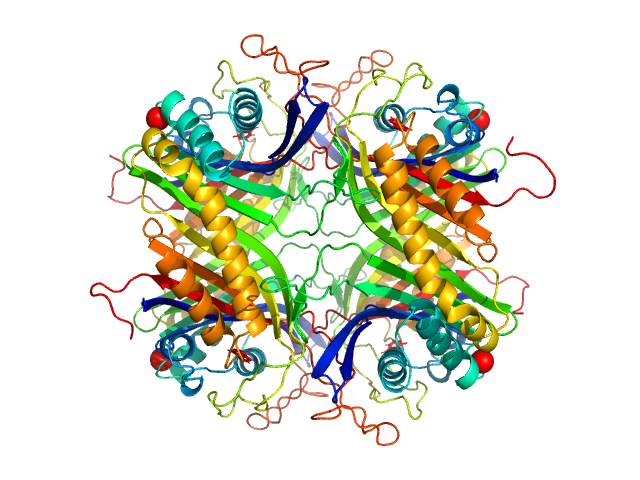
UniProt ID: Q00511 (2-302) Uricase
|
|
|
|
| Sample: |
Uricase tetramer, 136 kDa Aspergillus flavus protein
|
| Buffer: |
100 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.3 |
nm |
|
|