UniProt ID: P15795 (201-294) Cholera toxin transcriptional activator
UniProt ID: P24003 (25-173) Transmembrane regulatory protein ToxS
UniProt ID: None (None-None) bile acid: sodium cholate hydrate
|
|
|
|
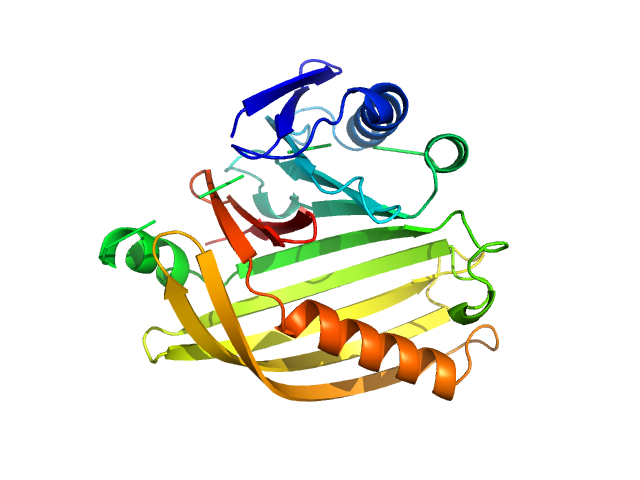
| Sample: |
Cholera toxin transcriptional activator monomer, 12 kDa Vibrio cholerae serotype … protein
Transmembrane regulatory protein ToxS dimer, 37 kDa Vibrio cholerae serotype … protein
Bile acid: sodium cholate hydrate monomer, 0 kDa
|
| Buffer: |
50 mM Na2HPO4, 300 mM NaCl, 3% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Jul 7
|
Vibrio cholerae's ToxRS bile sensing system.
Elife 12 (2023)
Gubensäk N, Sagmeister T, Buhlheller C, Geronimo BD, Wagner GE, Petrowitsch L, Gräwert MA, Rotzinger M, Berger TMI, Schäfer J, Usón I, Reidl J, Sánchez-Murcia PA, Zangger K, Pavkov-Keller T
|
| RgGuinier |
2.4 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
UniProt ID: O00571 (1-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 1-580)
|
|
|
|
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 1-580) monomer, 67 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
116 |
nm3 |
|
|
UniProt ID: O00571 (50-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 50-580)
|
|
|
|
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 50-580) monomer, 62 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.9 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
UniProt ID: O00571 (100-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 100-580)
|
|
|
|
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 100-580) monomer, 57 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
88 |
nm3 |
|
|
UniProt ID: O00571 (135-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 135-580)
|
|
|

|
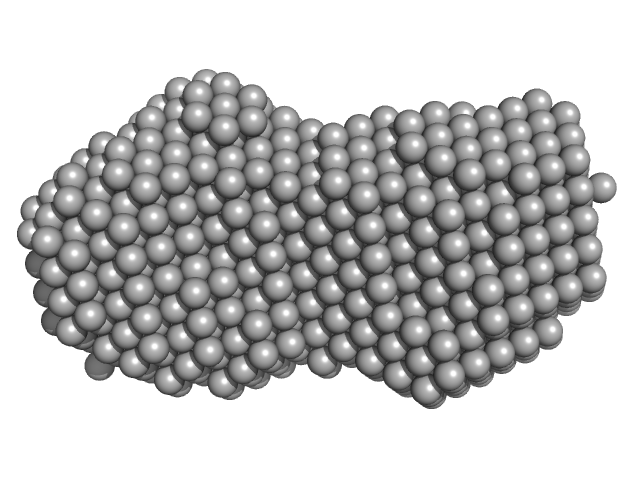
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 135-580) monomer, 52 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: O00571 (1-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 1-580)
UniProt ID: None (None-None) 15 nucleotide RNA duplex (ATP-dependent RNA helicase DDX3X binding target)
|
|
|

|
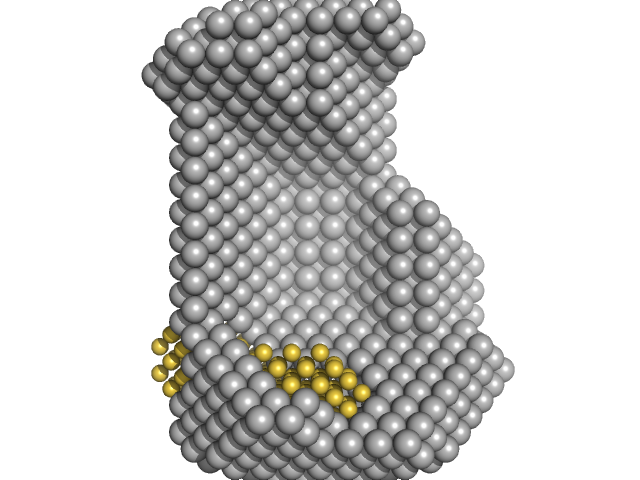
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 1-580) monomer, 67 kDa Homo sapiens protein
15 nucleotide RNA duplex (ATP-dependent RNA helicase DDX3X binding target) dimer, 10 kDa RNA
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
3.6 |
nm |
| Dmax |
11.2 |
nm |
| VolumePorod |
105 |
nm3 |
|
|
UniProt ID: O00571 (135-580) ATP-dependent RNA helicase DDX3X (truncation; amino acids 135-580)
UniProt ID: None (None-None) 15 nucleotide RNA duplex (ATP-dependent RNA helicase DDX3X binding target)
|
|
|

|
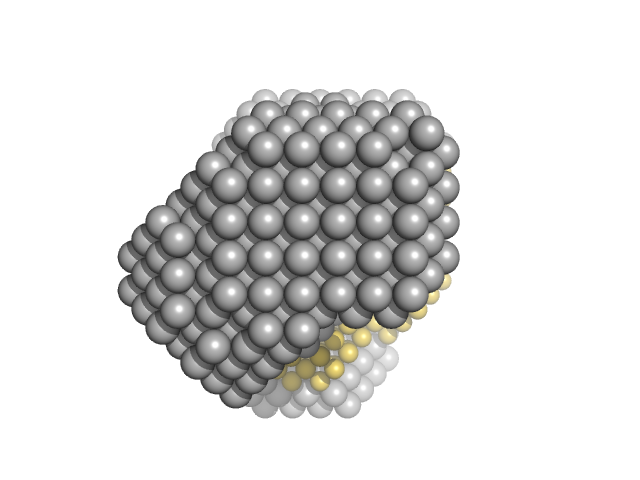
| Sample: |
ATP-dependent RNA helicase DDX3X (truncation; amino acids 135-580) monomer, 52 kDa Homo sapiens protein
15 nucleotide RNA duplex (ATP-dependent RNA helicase DDX3X binding target) dimer, 10 kDa RNA
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.5 |
nm |
|
|
UniProt ID: Q9LZ65 (1-222) AT5g04600/T32M21_200
|
|
|
|
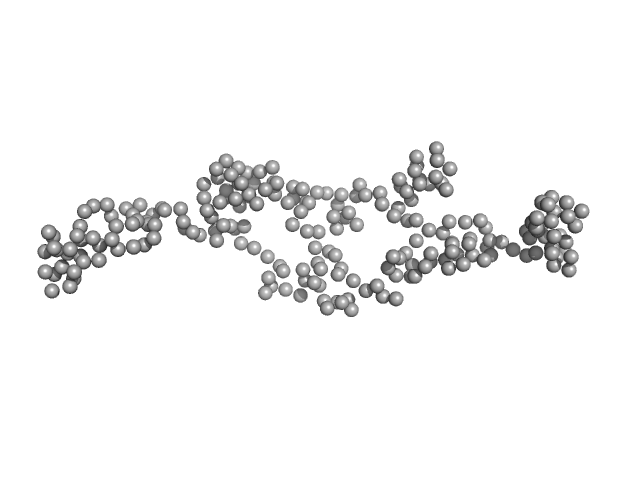
| Sample: |
AT5g04600/T32M21_200 monomer, 25 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HNa2PO4, 300 mM NaCl, 5% glycerol (v/v), 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 22
|
Structural and functional analysis of a plant nucleolar RNA chaperone-like protein.
Sci Rep 13(1):9656 (2023)
Fernandes R, Ostendorp A, Ostendorp S, Mehrmann J, Falke S, Graewert MA, Weingartner M, Kehr J, Hoth S
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
UniProt ID: Q16637 (1-294) Survival motor neuron protein (fusion construct with the protein transduction domain of the human immunodeficiency virus type 1 Tat protein)
|
|
|
|
| Sample: |
Survival motor neuron protein (fusion construct with the protein transduction domain of the human immunodeficiency virus type 1 Tat protein), 533 kDa synthetic construct protein
|
| Buffer: |
50 mM Tris-HCl, 500 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jun 13
|
Recombinant flSMN
Federico Forneris
|
| RgGuinier |
8.0 |
nm |
| Dmax |
29.7 |
nm |
| VolumePorod |
897 |
nm3 |
|
|
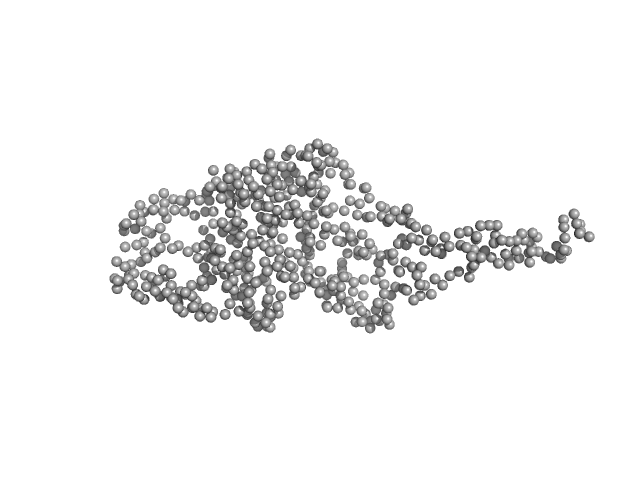
UniProt ID: Q5EAH9 (57-578) At3g55760
|
|
|

|
| Sample: |
At3g55760 monomer, 60 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM Tris, 150 mM NaCl, 10% glycerol, 2 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2016 Dec 8
|
LIKE EARLY STARVATION 1 and EARLY STARVATION 1 promote and stabilize amylopectin phase transition in starch biosynthesis.
Sci Adv 9(21):eadg7448 (2023)
Liu C, Pfister B, Osman R, Ritter M, Heutinck A, Sharma M, Eicke S, Fischer-Stettler M, Seung D, Bompard C, Abt MR, Zeeman SC
|
| RgGuinier |
3.7 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
121 |
nm3 |
|
|