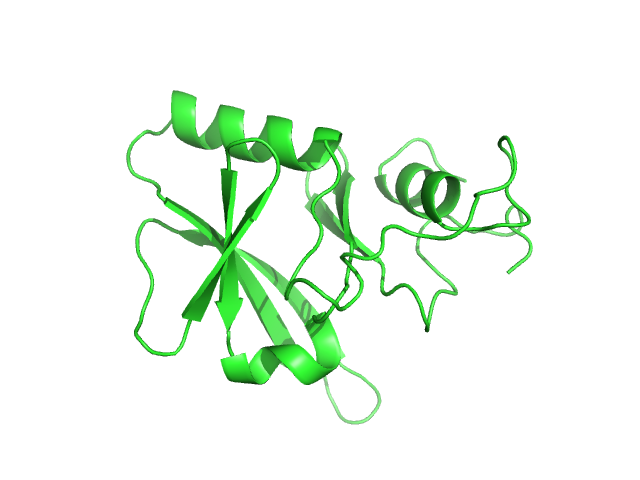
UniProt ID: P0DN75 (2-132) Iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Feb 25
|
Magnetic field effects on the structure and molecular behavior of pigeon iron–sulfur protein
Protein Science 31(6) (2022)
Arai S, Shimizu R, Adachi M, Hirai M
|
|
|
UniProt ID: E0IW31 (90-234) Accessory colonization factor
|
|
|

|
| Sample: |
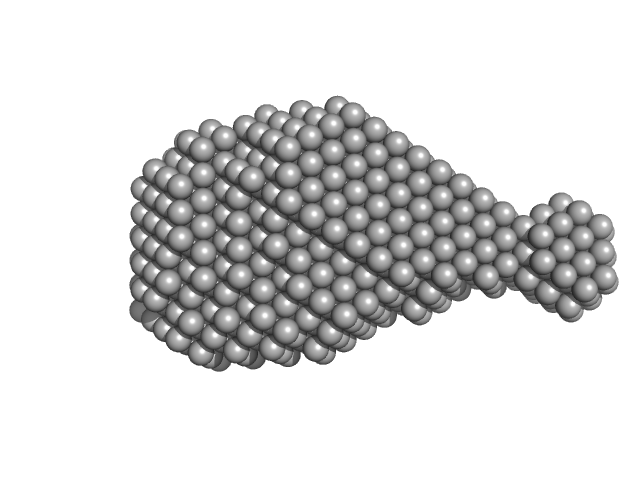
Accessory colonization factor monomer, 17 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Apr 21
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari V, Garnett J
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
UniProt ID: P02787 (1-698) Serotransferrin
|
|
|

|
| Sample: |
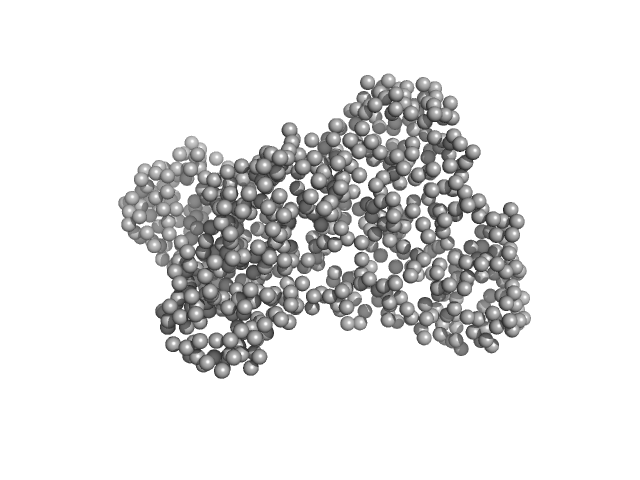
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl, (APO Buffer), pH: 5.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
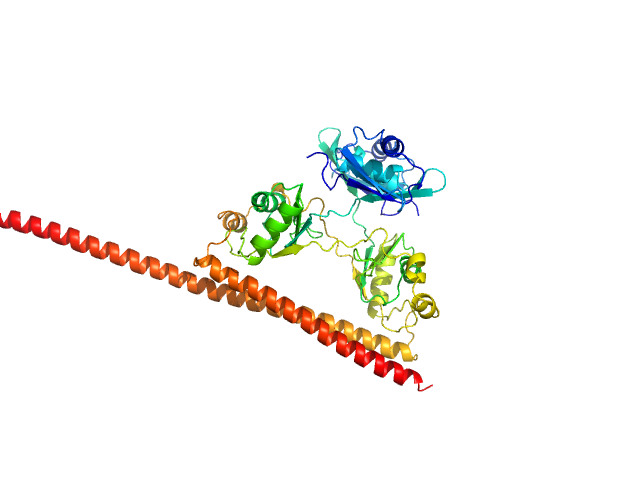
UniProt ID: P23246-1 (276-565) Splicing factor, proline- and glutamine-rich (276-565)
UniProt ID: Q15233-1 (53-312) Non-POU domain-containing octamer-binding protein (53-312)
|
|
|
|
| Sample: |
Splicing factor, proline- and glutamine-rich (276-565) monomer, 34 kDa Homo sapiens protein
Non-POU domain-containing octamer-binding protein (53-312) monomer, 30 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 250 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Apr 28
|
Structural plasticity of the coiled-coil interactions in human SFPQ.
Nucleic Acids Res (2024)
Koning HJ, Lai JY, Marshall AC, Stroeher E, Monahan G, Pullakhandam A, Knott GJ, Ryan TM, Fox AH, Whitten A, Lee M, Bond CS
|
| RgGuinier |
3.0 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
UniProt ID: None (None-None) Homo sapiens RNA component of 7SK nuclear ribonucleoprotein (RN7SK), small nuclear RNA
UniProt ID: O89943 (44-60) Protein Tat
|
|
|
|
| Sample: |
Homo sapiens RNA component of 7SK nuclear ribonucleoprotein (RN7SK), small nuclear RNA monomer, 18 kDa Homo sapiens RNA
Protein Tat monomer, 2 kDa Human immunodeficiency virus … protein
|
| Buffer: |
10 mM phosphate, 70 mM NaCl, 0.1 mM EDTA, pH: 5.6 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Nov 27
|
A structure-based mechanism for displacement of the HEXIM adapter from 7SK small nuclear RNA.
Commun Biol 5(1):819 (2022)
Pham VV, Gao M, Meagher JL, Smith JL, D'Souza VM
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
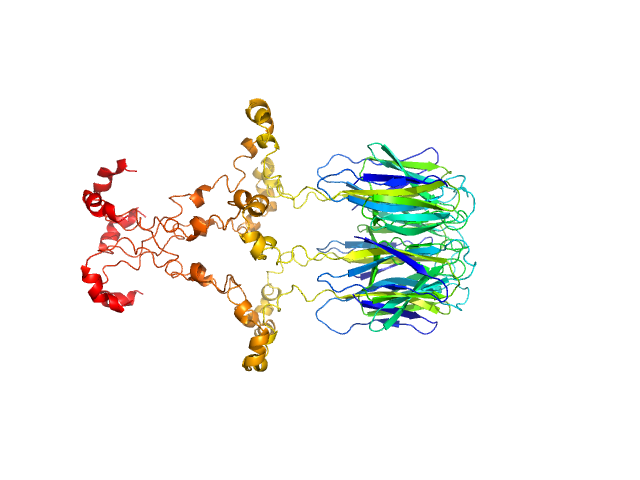
UniProt ID: F4J9Q6 (1-164) Peptidyl-prolyl cis-trans isomerase FKBP43
|
|
|
|
| Sample: |
Peptidyl-prolyl cis-trans isomerase FKBP43 pentamer, 96 kDa Arabidopsis thaliana protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 28
|
The plant nucleoplasmin AtFKBP43 needs its extended arms for histone interaction.
Biochim Biophys Acta Gene Regul Mech 1865(7):194872 (2022)
Singh AK, Saharan K, Baral S, Vasudevan D
|
| RgGuinier |
4.0 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
180 |
nm3 |
|
|
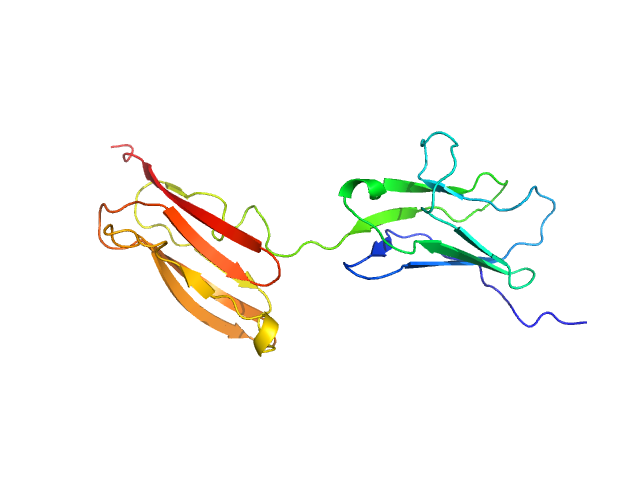
UniProt ID: Q5VST9 (1071-1251) Obscurin Ig domains 12/13 short
|
|
|
|
| Sample: |
Obscurin Ig domains 12/13 short monomer, 21 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 0.35 mM NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Oct 20
|
The N-terminus of obscurin is flexible in solution.
Proteins (2022)
Mauriello GE, Moncure GE, Nowzari RA, Miller CJ, Wright NT
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
UniProt ID: Q63HQ2 (784-1017) Pikachurin
|
|
|

|
| Sample: |
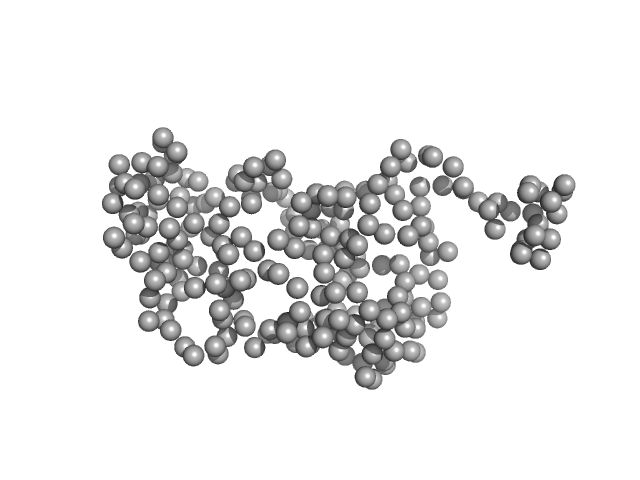
Pikachurin monomer, 26 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Mar 3
|
Structure of the photoreceptor synaptic assembly of the extracellular matrix protein pikachurin with the orphan receptor GPR179
Science Signaling 16(795) (2023)
Patil D, Pantalone S, Cao Y, Laboute T, Novick S, Singh S, Savino S, Faravelli S, Magnani F, Griffin P, Singh A, Forneris F, Martemyanov K
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
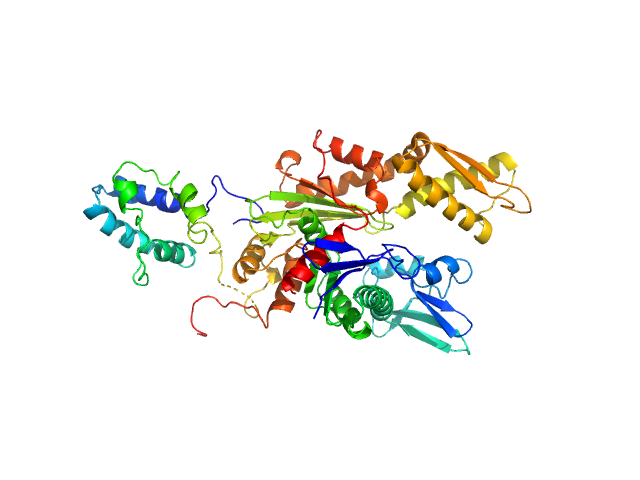
UniProt ID: P11021 (26-405) Endoplasmic reticulum chaperone BiP
UniProt ID: Q49AH0 (1-187) Cerebral dopamine neurotrophic factor
|
|
|
|
| Sample: |
Endoplasmic reticulum chaperone BiP monomer, 42 kDa Homo sapiens protein
Cerebral dopamine neurotrophic factor monomer, 21 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 21
|
Structural basis of CDNF interaction with the UPR regulator GRP78.
Nat Commun 15(1):8175 (2024)
Graewert MA, Volkova M, Jonasson K, Määttä JAE, Gräwert T, Mamidi S, Kulesskaya N, Evenäs J, Johnsson RE, Svergun D, Bhattacharjee A, Huttunen HJ
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
UniProt ID: P74102 (2-317) Orange carotenoid-binding protein
|
|
|

|
| Sample: |
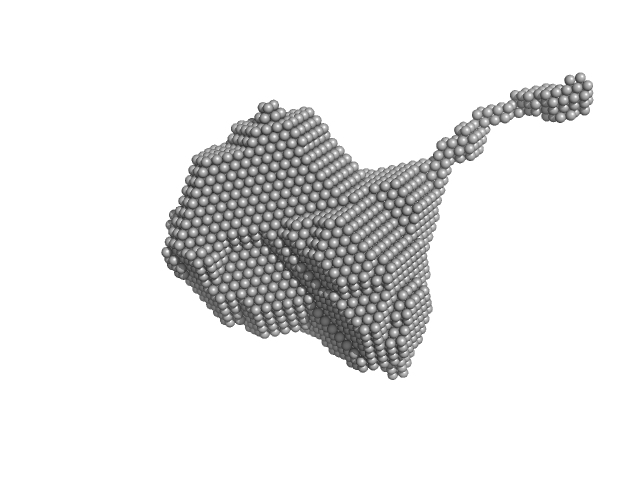
Orange carotenoid-binding protein dimer, 70 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM NaCL, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 23
|
Oligomerization processes limit photoactivation and recovery of the orange carotenoid protein.
Biophys J 121(15):2849-2872 (2022)
Andreeva EA, Niziński S, Wilson A, Levantino M, De Zitter E, Munro R, Muzzopappa F, Thureau A, Zala N, Burdzinski G, Sliwa M, Kirilovsky D, Schirò G, Colletier JP
|
| RgGuinier |
2.8 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
85 |
nm3 |
|
|