UniProt ID: P68135 (1-377) Actin, alpha skeletal muscle
|
|
|

|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 0.1 mM CaCl2, 1 mM NaN3, 1.0 mM ATP, 50 mM KCl, 2 mM MgCl2, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 1
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
15.7 |
nm |
| Dmax |
60.0 |
nm |
| VolumePorod |
4710 |
nm3 |
|
|
UniProt ID: P68135 (1-377) Actin, alpha skeletal muscle
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDTZ4_fit1_model1.png)
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 2.0 mM EDTA, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Nov 16
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
4.9 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
549 |
nm3 |
|
|
UniProt ID: P68135 (1-377) Actin, alpha skeletal muscle
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDT25_fit1_model1.png)
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris, 0.1 mM CaCl2, 1 mM NaN3, 0.2 mM ATP, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 5
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
3.1 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
UniProt ID: P15116 (160-714) Cadherin-2
|
|
|
|
| Sample: |
Cadherin-2 dimer, 123 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3 mM CaCl2, 0.02% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 May 1
|
Rapid simulation of glycoprotein structures by grafting and steric exclusion of glycan conformer libraries.
Cell 187(5):1296-1311.e26 (2024)
Tsai YX, Chang NE, Reuter K, Chang HT, Yang TJ, von Bülow S, Sehrawat V, Zerrouki N, Tuffery M, Gecht M, Grothaus IL, Colombi Ciacchi L, Wang YS, Hsu MF, Khoo KH, Hummer G, Hsu SD, Hanus C, Sikora M
|
| RgGuinier |
8.8 |
nm |
| Dmax |
40.0 |
nm |
| VolumePorod |
496 |
nm3 |
|
|
UniProt ID: P15116 (490-702) Cadherin-2
|
|
|
|
| Sample: |
Cadherin-2 monomer, 24 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3 mM CaCl2, 0.02% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 May 1
|
Rapid simulation of glycoprotein structures by grafting and steric exclusion of glycan conformer libraries.
Cell 187(5):1296-1311.e26 (2024)
Tsai YX, Chang NE, Reuter K, Chang HT, Yang TJ, von Bülow S, Sehrawat V, Zerrouki N, Tuffery M, Gecht M, Grothaus IL, Colombi Ciacchi L, Wang YS, Hsu MF, Khoo KH, Hummer G, Hsu SD, Hanus C, Sikora M
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
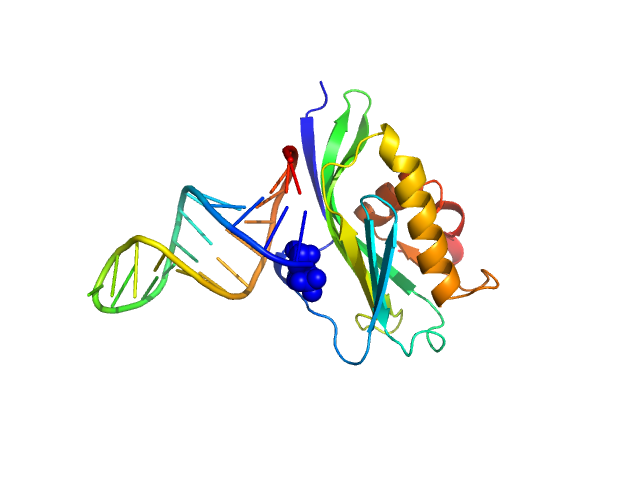
UniProt ID: X2C382 (141-281) Non-structural protein 1
UniProt ID: None (None-None) 5’ppp ds10 HP RNA
|
|
|
|
| Sample: |
Non-structural protein 1 monomer, 16 kDa Influenza B virus protein
5’ppp ds10 HP RNA monomer, 8 kDa RNA
|
| Buffer: |
40 mM ammonium acetate, pH 5.5, 225 mM NaCl, 5 mM CaCl2, 0.02% NaN3, 50 mM arginine, pH: 5.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Aug 6
|
The NS1 protein of influenza B virus binds 5’-triphosphorylated dsRNA to suppress RIG-I activation and the host antiviral response
(2024)
Woltz R, Schweibenz B, Tsutakawa S, Zhao C, Ma L, Shurina B, Hura G, John R, Vorobiev S, Swapna G, Solotchi M, Tainer J, Krug R, Patel S, Montelione G
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.9 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
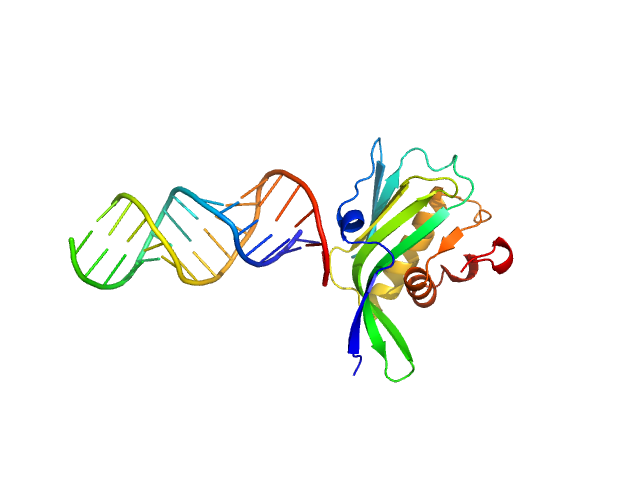
UniProt ID: X2C382 (141-281) Non-structural protein 1
UniProt ID: None (None-None) RNA top strand
UniProt ID: None (None-None) RNA bottom strand
|
|
|
|
| Sample: |
Non-structural protein 1 monomer, 16 kDa Influenza B virus protein
RNA top strand monomer, 5 kDa RNA
RNA bottom strand monomer, 5 kDa RNA
|
| Buffer: |
40 mM ammonium acetate, pH 5.5, 450 mM NaCl, 5 mM CaCl2, 0.02% NaN3, 50 mM arginine, pH: 5.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2017 May 18
|
The NS1 protein of influenza B virus binds 5’-triphosphorylated dsRNA to suppress RIG-I activation and the host antiviral response
(2024)
Woltz R, Schweibenz B, Tsutakawa S, Zhao C, Ma L, Shurina B, Hura G, John R, Vorobiev S, Swapna G, Solotchi M, Tainer J, Krug R, Patel S, Montelione G
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
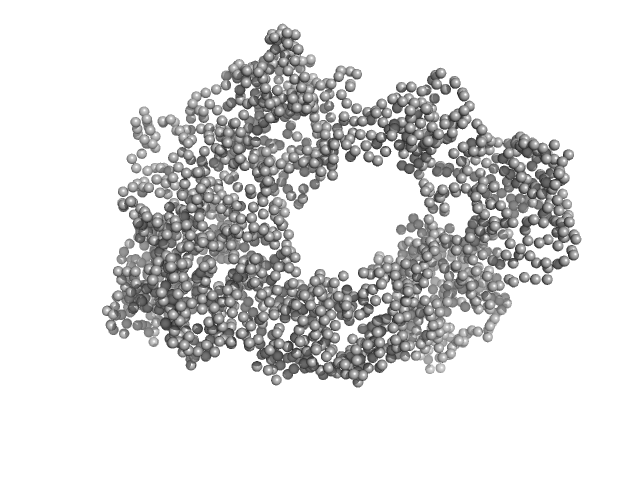
UniProt ID: Q8IYU2 (1-909) E3 ubiquitin-protein ligase HACE1
|
|
|

|
| Sample: |
E3 ubiquitin-protein ligase HACE1 dimer, 205 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 50 mM NaCl, 5 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Apr 21
|
Structural mechanisms of autoinhibition and substrate recognition by the ubiquitin ligase HACE1
Nature Structural & Molecular Biology (2024)
Düring J, Wolter M, Toplak J, Torres C, Dybkov O, Fokkens T, Bohnsack K, Urlaub H, Steinchen W, Dienemann C, Lorenz S
|
| RgGuinier |
5.2 |
nm |
| Dmax |
16.4 |
nm |
| VolumePorod |
379 |
nm3 |
|
|
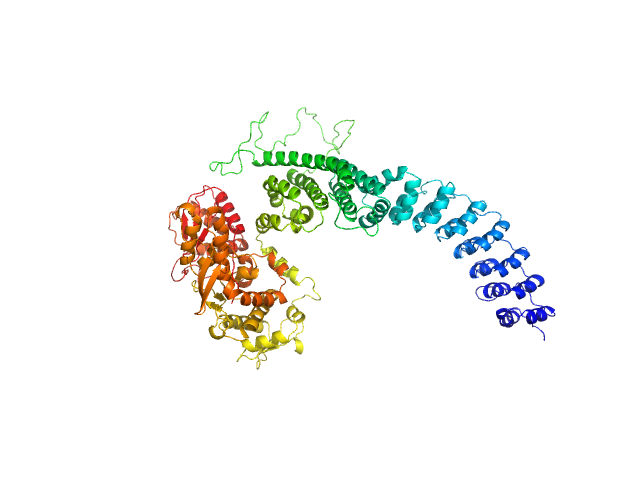
UniProt ID: Q8IYU2 (22-909) E3 ubiquitin-protein ligase HACE1
|
|
|
|
| Sample: |
E3 ubiquitin-protein ligase HACE1 monomer, 100 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 50 mM NaCl, 5 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Apr 21
|
Structural mechanisms of autoinhibition and substrate recognition by the ubiquitin ligase HACE1
Nature Structural & Molecular Biology (2024)
Düring J, Wolter M, Toplak J, Torres C, Dybkov O, Fokkens T, Bohnsack K, Urlaub H, Steinchen W, Dienemann C, Lorenz S
|
| RgGuinier |
4.6 |
nm |
| Dmax |
17.8 |
nm |
| VolumePorod |
219 |
nm3 |
|
|
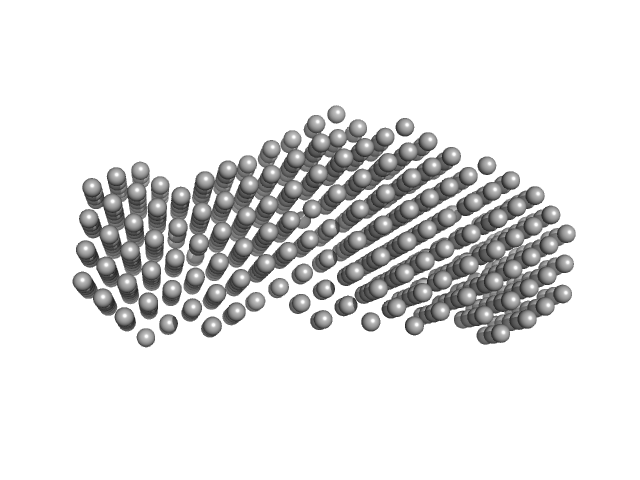
UniProt ID: Q4K5N8 (1-183) Heme acquisition protein HasAp
|
|
|

|
| Sample: |
Heme acquisition protein HasAp dimer, 38 kDa Pseudomonas fluorescens (strain … protein
|
| Buffer: |
50 mM CHES, 5 % glycerol, pH: 9.5 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2023 Feb 24
|
Heme-substituted protein assembly bridged by synthetic porphyrin: achieving controlled configuration while maintaining rotational freedom
RSC Advances 14(13):8829-8836 (2024)
Inaba H, Shisaka Y, Ariyasu S, Sakakibara E, Ueda G, Aiba Y, Shimizu N, Sugimoto H, Shoji O
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
54 |
nm3 |
|
|