UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2 (R580K)
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2 (R580K), 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
5.9 |
nm |
| Dmax |
27.0 |
nm |
| VolumePorod |
381 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2 (R580K)
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2 (R580K), 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
5.7 |
nm |
| Dmax |
30.0 |
nm |
| VolumePorod |
283 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2 (R580K)
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2 (R580K), 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
5.3 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
264 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2 (R580K)
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2 (R580K), 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
5.2 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
243 |
nm3 |
|
|
UniProt ID: J3QMY9 (1-554) Type 2 DNA topoisomerase 6 subunit B-like
|
|
|
|
| Sample: |
Type 2 DNA topoisomerase 6 subunit B-like monomer, 61 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 500 mM KCl, 5% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2023 Apr 12
|
The TOPOVIBL meiotic DSB formation protein: new insights from its biochemical and structural characterization
Nucleic Acids Research (2024)
Diagouraga B, Tambones I, Carivenc C, Bechara C, Nadal M, de Massy B, le Maire A, Robert T
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
114 |
nm3 |
|
|
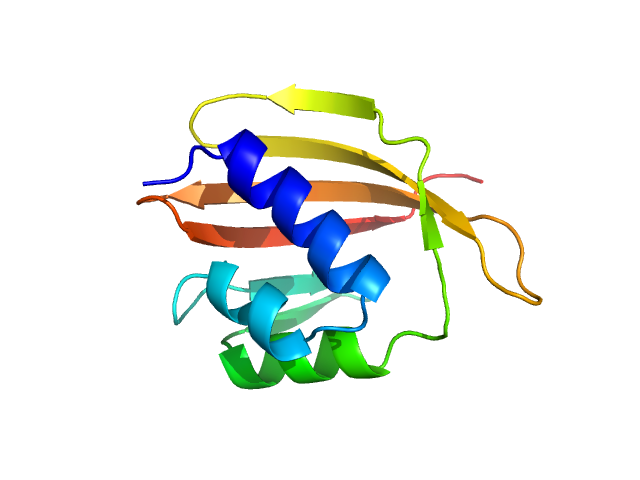
UniProt ID: D1C7H4 (1-108) SnoaL-like domain-containing protein
|
|
|
|
| Sample: |
SnoaL-like domain-containing protein monomer, 12 kDa Sphaerobacter thermophilus (strain … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 3% glycerol, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Nov 6
|
Elements of the C-terminal tail of a C-terminal domain homolog of the Orange Carotenoid Protein determining xanthophyll uptake from liposomes.
Biochim Biophys Acta Bioenerg 1865(3):149043 (2024)
Likkei K, Moldenhauer M, Tavraz NN, Egorkin NA, Slonimskiy YB, Maksimov EG, Sluchanko NN, Friedrich T
|
| RgGuinier |
1.5 |
nm |
| Dmax |
5.0 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
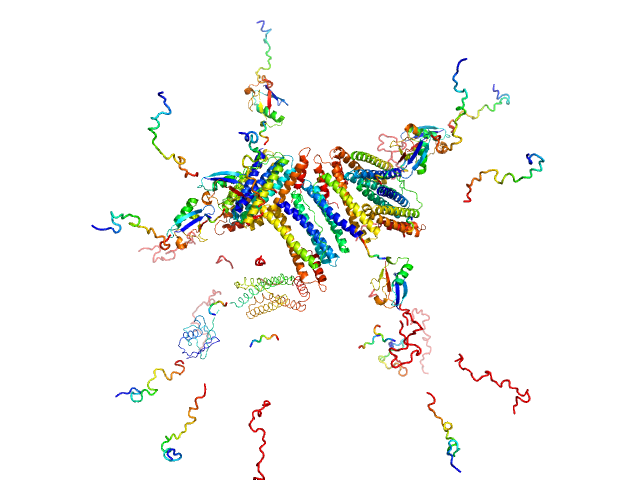
UniProt ID: Q9ZLI1 (5-167) Bacterial non-heme ferritin (N19Q, I59V, N-terminal His-SUMO fusion)
|
|
|
|
| Sample: |
Bacterial non-heme ferritin (N19Q, I59V, N-terminal His-SUMO fusion) 24-mer, 755 kDa Helicobacter pylori (strain … protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2022 Jul 30
|
Ferritin-based fusion protein shows octameric deadlock state of self-assembly
Biochemical and Biophysical Research Communications 690:149276 (2024)
Sudarev V, Gette M, Bazhenov S, Tilinova O, Zinovev E, Manukhov I, Kuklin A, Ryzhykau Y, Vlasov A
|
| RgGuinier |
7.6 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
2010 |
nm3 |
|
|
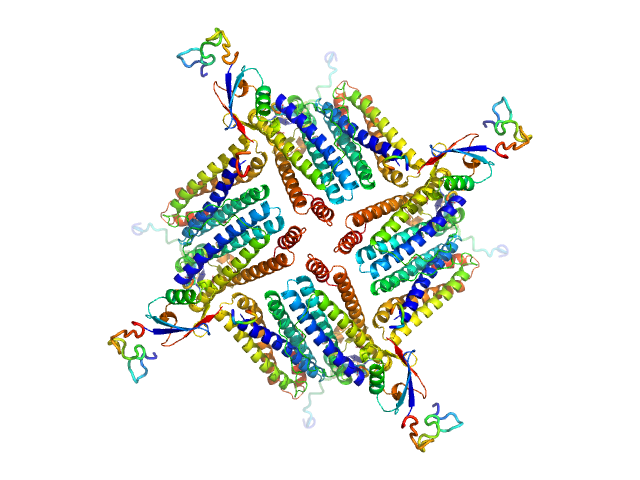
UniProt ID: Q9ZLI1 (5-167) Bacterial non-heme ferritin (N19Q, I59V, N-terminal His-SUMO fusion)
|
|
|

|
| Sample: |
Bacterial non-heme ferritin (N19Q, I59V, N-terminal His-SUMO fusion) octamer, 252 kDa Helicobacter pylori (strain … protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2022 Jul 17
|
Ferritin-based fusion protein shows octameric deadlock state of self-assembly
Biochemical and Biophysical Research Communications 690:149276 (2024)
Sudarev V, Gette M, Bazhenov S, Tilinova O, Zinovev E, Manukhov I, Kuklin A, Ryzhykau Y, Vlasov A
|
| RgGuinier |
5.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
618 |
nm3 |
|
|
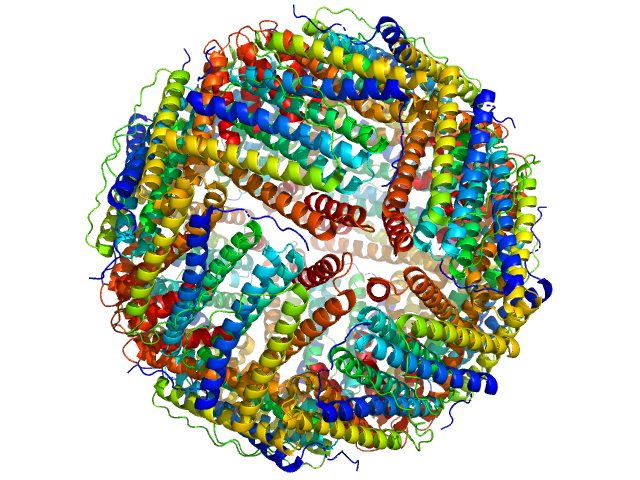
UniProt ID: Q9ZLI1 (5-167) Bacterial non-heme ferritin (N19Q, I59V)
|
|
|
|
| Sample: |
Bacterial non-heme ferritin (N19Q, I59V) 24-mer, 480 kDa Helicobacter pylori (strain … protein
|
| Buffer: |
20 mM Tris, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2022 Dec 14
|
Ferritin-based fusion protein shows octameric deadlock state of self-assembly
Biochemical and Biophysical Research Communications 690:149276 (2024)
Sudarev V, Gette M, Bazhenov S, Tilinova O, Zinovev E, Manukhov I, Kuklin A, Ryzhykau Y, Vlasov A
|
| RgGuinier |
5.6 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
674 |
nm3 |
|
|
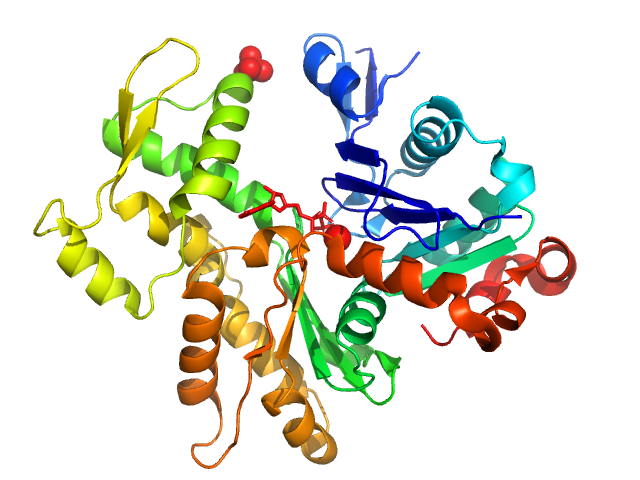
UniProt ID: P68135 (1-377) Actin, alpha skeletal muscle
|
|
|
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 0.1 mM CaCl2, 1 mM NaN3, 0.2 mM ATP, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 1
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
58 |
nm3 |
|
|