UniProt ID: Q13426 (201-336) DNA repair protein XRCC4
|
|
|
|
| Sample: |
DNA repair protein XRCC4 monomer, 15 kDa Homo sapiens protein
|
| Buffer: |
20 mM Bis-tris, 150 mM KCl, 1 mM EDTA, 1 mM DTT, pH: 6.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Oct 20
|
Multivalent interactions of the disordered regions of XLF and XRCC4 foster robust cellular NHEJ and drive the formation of ligation-boosting condensates in vitro.
Nat Struct Mol Biol (2024)
Vu DD, Bonucci A, Brenière M, Cisneros-Aguirre M, Pelupessy P, Wang Z, Carlier L, Bouvignies G, Cortes P, Aggarwal AK, Blackledge M, Gueroui Z, Belle V, Stark JM, Modesti M, Ferrage F
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
41 |
nm3 |
|
|
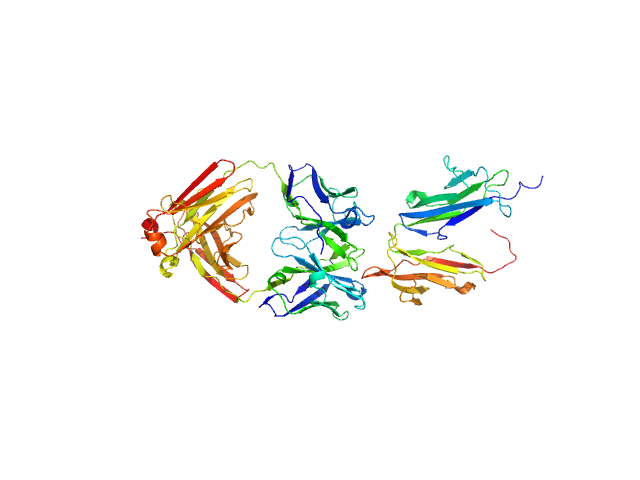
UniProt ID: P31994 (43-217) Low affinity immunoglobulin gamma Fc region receptor II-b
UniProt ID: None (None-None) Human IgG1 F(ab') 5C05
|
|
|
|
| Sample: |
Low affinity immunoglobulin gamma Fc region receptor II-b monomer, 20 kDa Homo sapiens protein
Human IgG1 F(ab') 5C05 monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 May 18
|
Antibody binding geometry and affinity control inhibitory FcγRIIB receptor signaling
Hayden Fisher
|
| RgGuinier |
3.5 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
95 |
nm3 |
|
|
UniProt ID: P31994 (43-217) Low affinity immunoglobulin gamma Fc region receptor II-b
UniProt ID: None (None-None) Human IgG1 F(ab') 6C11
|
|
|
|
| Sample: |
Low affinity immunoglobulin gamma Fc region receptor II-b monomer, 20 kDa Homo sapiens protein
Human IgG1 F(ab') 6C11 monomer, 24 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 May 9
|
Antibody binding geometry and affinity control inhibitory FcγRIIB receptor signaling
Hayden Fisher
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
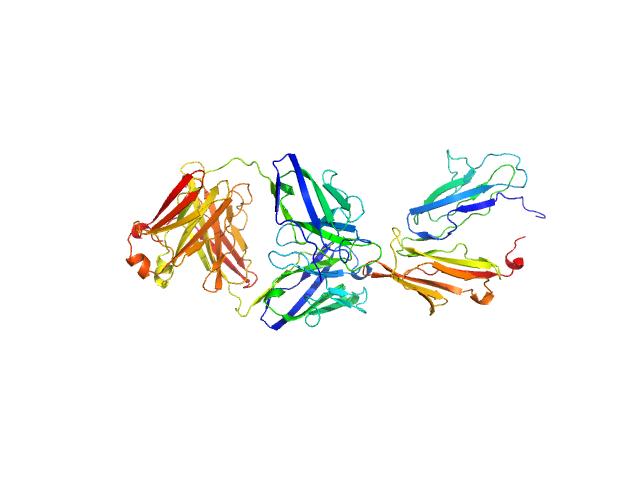
UniProt ID: P31994 (43-217) Low affinity immunoglobulin gamma Fc region receptor II-b
UniProt ID: None (None-None) Human IgG1 F(ab') 6G11
|
|
|
|
| Sample: |
Low affinity immunoglobulin gamma Fc region receptor II-b monomer, 20 kDa Homo sapiens protein
Human IgG1 F(ab') 6G11 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Apr 9
|
Antibody binding geometry and affinity control inhibitory FcγRIIB receptor signaling
Hayden Fisher
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
97 |
nm3 |
|
|
UniProt ID: P31994 (43-217) Low affinity immunoglobulin gamma Fc region receptor II-b
UniProt ID: None (None-None) Human IgG1 F(ab') 7C07
|
|
|
|
| Sample: |
Low affinity immunoglobulin gamma Fc region receptor II-b monomer, 20 kDa Homo sapiens protein
Human IgG1 F(ab') 7C07 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Dec 5
|
Antibody binding geometry and affinity control inhibitory FcγRIIB receptor signaling
Hayden Fisher
|
| RgGuinier |
3.6 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
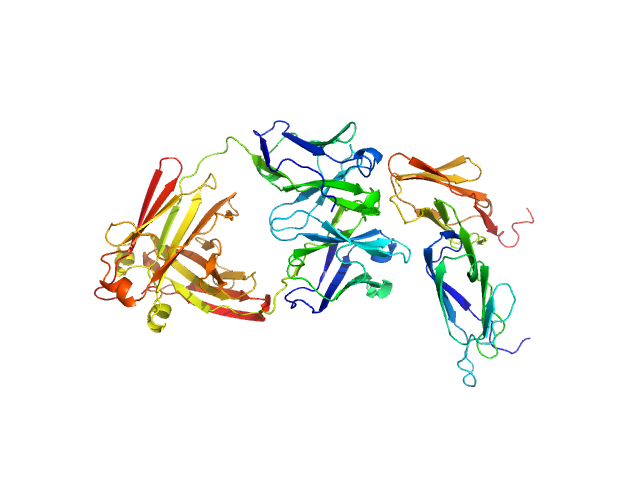
UniProt ID: P31994 (43-217) Low affinity immunoglobulin gamma Fc region receptor II-b
UniProt ID: None (None-None) Human IgG1 F(ab') 6G08
|
|
|
|
| Sample: |
Low affinity immunoglobulin gamma Fc region receptor II-b monomer, 20 kDa Homo sapiens protein
Human IgG1 F(ab') 6G08 monomer, 24 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Dec 14
|
Antibody binding geometry and affinity control inhibitory FcγRIIB receptor signaling
Hayden Fisher
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
96 |
nm3 |
|
|
UniProt ID: E7BLH6 (387-536) HbP1
|
|
|
|
| Sample: |
HbP1 trimer, 56 kDa Legionella pneumophila protein
|
| Buffer: |
20 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM EDTA, pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 29
|
The Legionella collagen-like protein employs a distinct binding mechanism for the recognition of host glycosaminoglycans.
Nat Commun 15(1):4912 (2024)
Rehman S, Antonovic AK, McIntire IE, Zheng H, Cleaver L, Baczynska M, Adams CO, Portlock T, Richardson K, Shaw R, Oregioni A, Mastroianni G, Whittaker SB, Kelly G, Lorenz CD, Fornili A, Cianciotto NP, Garnett JA
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
105 |
nm3 |
|
|
UniProt ID: E7BLH6 (387-536) HbP1 (R477A)
|
|
|
|
| Sample: |
HbP1 (R477A) trimer, 55 kDa Legionella pneumophila protein
|
| Buffer: |
20 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM EDTA, pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 29
|
The Legionella collagen-like protein employs a distinct binding mechanism for the recognition of host glycosaminoglycans.
Nat Commun 15(1):4912 (2024)
Rehman S, Antonovic AK, McIntire IE, Zheng H, Cleaver L, Baczynska M, Adams CO, Portlock T, Richardson K, Shaw R, Oregioni A, Mastroianni G, Whittaker SB, Kelly G, Lorenz CD, Fornili A, Cianciotto NP, Garnett JA
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
79 |
nm3 |
|
|
UniProt ID: E7BLH6 (387-536) HbP1 (R477A)
|
|
|
|
| Sample: |
HbP1 (R477A) monomer, 18 kDa Legionella pneumophila protein
|
| Buffer: |
20 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM EDTA, pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 29
|
The Legionella collagen-like protein employs a distinct binding mechanism for the recognition of host glycosaminoglycans.
Nat Commun 15(1):4912 (2024)
Rehman S, Antonovic AK, McIntire IE, Zheng H, Cleaver L, Baczynska M, Adams CO, Portlock T, Richardson K, Shaw R, Oregioni A, Mastroianni G, Whittaker SB, Kelly G, Lorenz CD, Fornili A, Cianciotto NP, Garnett JA
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.4 |
nm |
| VolumePorod |
33 |
nm3 |
|
|
UniProt ID: E7BLH6 (387-536) HbP1 (E503A)
|
|
|
|
| Sample: |
HbP1 (E503A) trimer, 56 kDa Legionella pneumophila protein
|
| Buffer: |
20 mM Tris–HCl pH 8.0, 200 mM NaCl, 5 mM EDTA, pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 29
|
The Legionella collagen-like protein employs a distinct binding mechanism for the recognition of host glycosaminoglycans.
Nat Commun 15(1):4912 (2024)
Rehman S, Antonovic AK, McIntire IE, Zheng H, Cleaver L, Baczynska M, Adams CO, Portlock T, Richardson K, Shaw R, Oregioni A, Mastroianni G, Whittaker SB, Kelly G, Lorenz CD, Fornili A, Cianciotto NP, Garnett JA
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
106 |
nm3 |
|
|