UniProt ID: Q89E26 (None-None) Proline dehydrogenase
|
|
|
|
| Sample: |
Proline dehydrogenase tetramer, 430 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 12
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
5.2 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
553 |
nm3 |
|
|
UniProt ID: Q89E26 (None-None) Proline dehydrogenase
|
|
|

|
| Sample: |
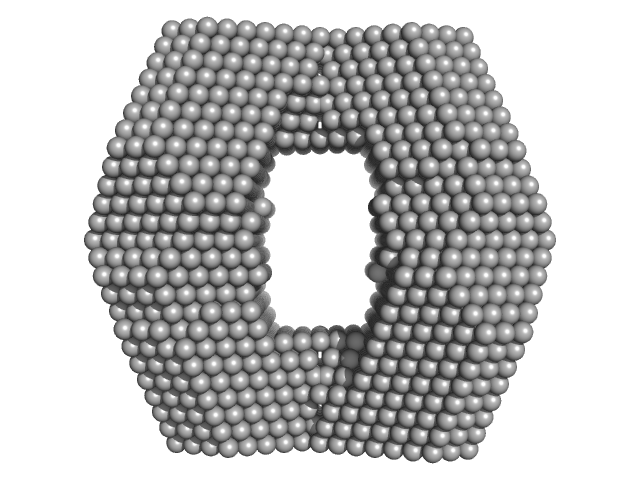
Proline dehydrogenase tetramer, 430 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 12
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
5.2 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
560 |
nm3 |
|
|
UniProt ID: Q5ZUU6 (None-None) Bifunctional protein PutA
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 238 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 20
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
297 |
nm3 |
|
|
UniProt ID: Q5ZUU6 (None-None) Bifunctional protein PutA
|
|
|

|
| Sample: |
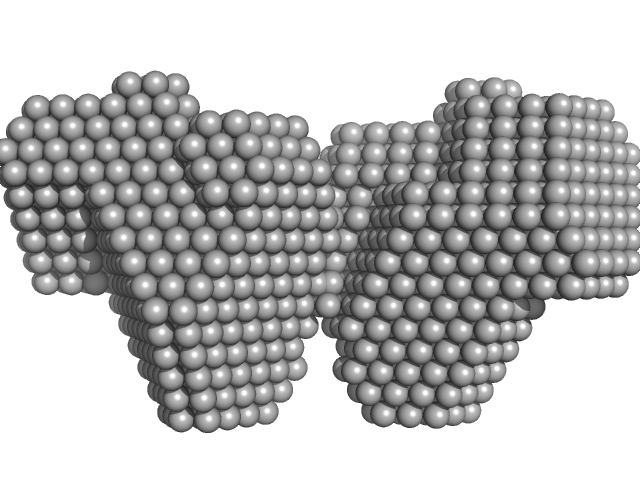
Bifunctional protein PutA dimer, 238 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 20
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
295 |
nm3 |
|
|
UniProt ID: A0A0E0T6R2 (None-None) Bifunctional protein PutA
|
|
|

|
| Sample: |
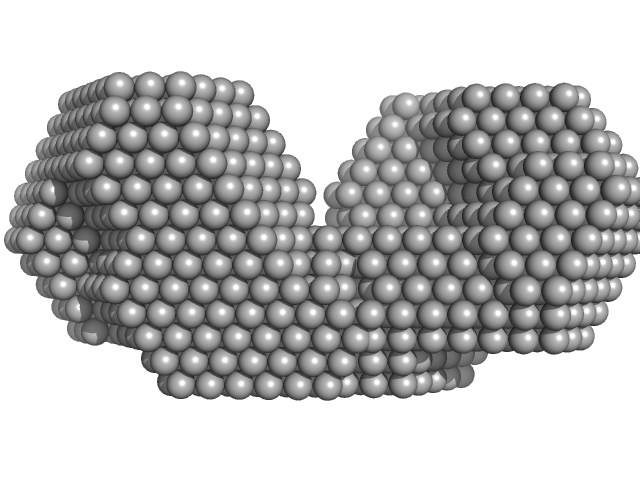
Bifunctional protein PutA dimer, 229 kDa Desulfovibrio vulgaris protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
295 |
nm3 |
|
|
UniProt ID: A0A0E0T6R2 (None-None) Bifunctional protein PutA
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 229 kDa Desulfovibrio vulgaris protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
294 |
nm3 |
|
|
UniProt ID: Q89E26 (None-None) Proline dehydrogenase
|
|
|
|
| Sample: |
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
281 |
nm3 |
|
|
UniProt ID: Q89E26 (None-None) Proline dehydrogenase
|
|
|
|
| Sample: |
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.5 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
283 |
nm3 |
|
|
UniProt ID: Q89E26 (None-None) Proline dehydrogenase
|
|
|

|
| Sample: |
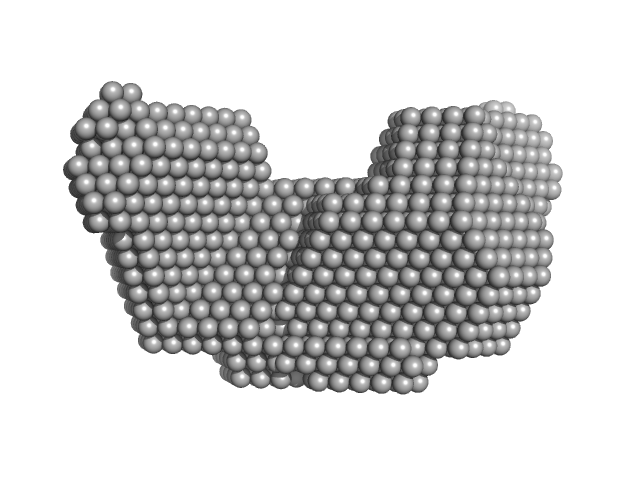
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
FEBS J 284(18):3029-3049 (2017)
Korasick DA, Singh H, Pemberton TA, Luo M, Dhatwalia R, Tanner JJ
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
289 |
nm3 |
|
|
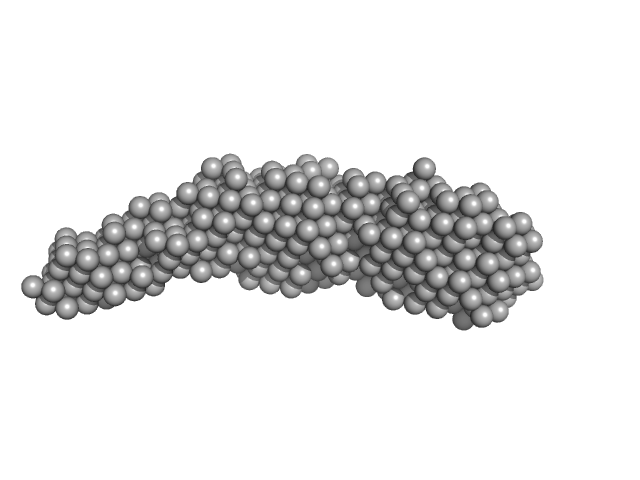
UniProt ID: Q46085 (806-1021) ColH protein
UniProt ID: (None-None) Collagenous Peptide model [(PPG)10]
|
|
|
|
| Sample: |
ColH protein monomer, 24 kDa Hathewaya histolytica protein
Collagenous Peptide model [(PPG)10] trimer, 9 kDa synthetic construct protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 5 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
Ca2+ -induced orientation of tandem collagen binding domains from clostridial collagenase ColG permits two opposing functions of collagen fibril formation and retardation.
FEBS J 285(17):3254-3269 (2018)
Caviness P, Bauer R, Tanaka K, Janowska K, Roeser JR, Harter D, Sanders J, Ruth C, Matsushita O, Sakon J
|
| RgGuinier |
2.7 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|