UniProt ID: Q13263 (23-418) Transcription intermediary factor 1-beta, TIF1b, KAP1, TRIM28, Fragment 23-418, RBCC domain
|
|
|

|
| Sample: |
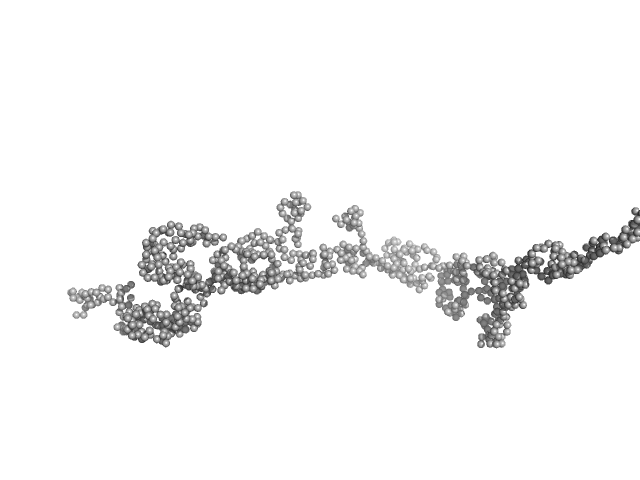
Transcription intermediary factor 1-beta, TIF1b, KAP1, TRIM28, Fragment 23-418, RBCC domain dimer, 92 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 500 mM NaCl, 10 % Glycerol, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jul 6
|
KAP1 is an antiparallel dimer with a functional asymmetry.
Life Sci Alliance 2(4) (2019)
Fonti G, Marcaida MJ, Bryan LC, Träger S, Kalantzi AS, Helleboid PJ, Demurtas D, Tully MD, Grudinin S, Trono D, Fierz B, Dal Peraro M
|
| RgGuinier |
8.3 |
nm |
| Dmax |
37.0 |
nm |
| VolumePorod |
381 |
nm3 |
|
|
UniProt ID: Q4J9K7 (32-151) Stator protein FlaG soluble domain
UniProt ID: Q4J9K8 (35-164) Conserved flagellar protein FlaF-I96Y soluble domain
|
|
|
|
| Sample: |
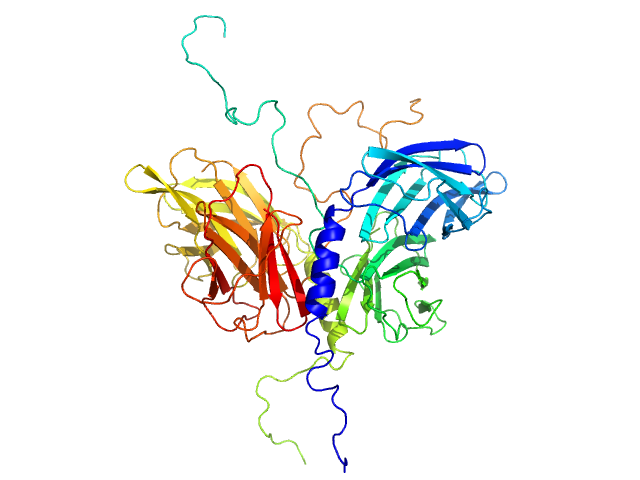
Stator protein FlaG soluble domain dimer, 30 kDa Sulfolobus acidocaldarius protein
Conserved flagellar protein FlaF-I96Y soluble domain dimer, 33 kDa Sulfolobus acidocaldarius protein
|
| Buffer: |
25 mM citric acid/sodium citrate, 150mM NaCl, 3% Glycerol, pH: 3 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Nov 10
|
The structure of the periplasmic FlaG-FlaF complex and its essential role for archaellar swimming motility.
Nat Microbiol (2019)
Tsai CL, Tripp P, Sivabalasarma S, Zhang C, Rodriguez-Franco M, Wipfler RL, Chaudhury P, Banerjee A, Beeby M, Whitaker RJ, Tainer JA, Albers SV
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
90 |
nm3 |
|
|
UniProt ID: P53893 (None-None) Ribosome assembly protein 1
|
|
|

|
| Sample: |
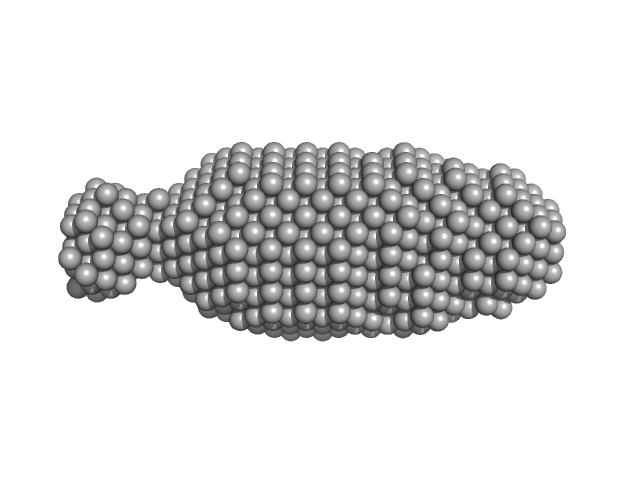
Ribosome assembly protein 1 monomer, 124 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Sep 21
|
Interaction of the GTPase Elongation Factor Like-1 with the Shwachman-Diamond Syndrome Protein and Its Missense Mutations.
Int J Mol Sci 19(12) (2018)
Gijsbers A, Montagut DC, Méndez-Godoy A, Altamura D, Saviano M, Siliqi D, Sánchez-Puig N
|
| RgGuinier |
4.7 |
nm |
| Dmax |
15.8 |
nm |
| VolumePorod |
258 |
nm3 |
|
|
UniProt ID: Q13478 (22-329) Interleukin-18 receptor 1
|
|
|
|
| Sample: |
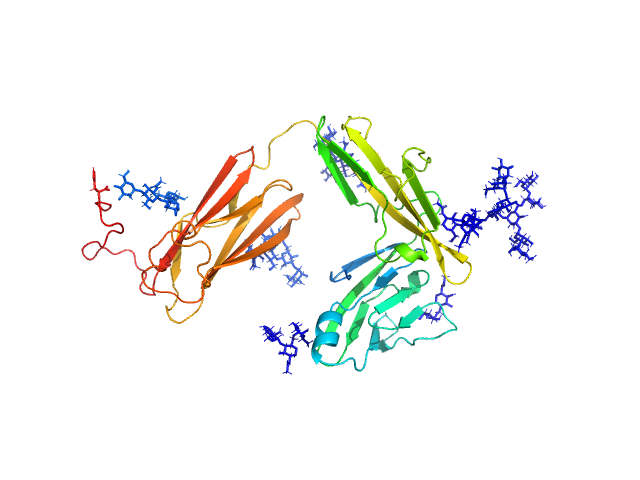
Interleukin-18 receptor 1 monomer, 36 kDa Homo sapiens protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 3% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Oct 3
|
Functional Relevance of Interleukin-1 Receptor Inter-domain Flexibility for Cytokine Binding and Signaling.
Structure 27(8):1296-1307.e5 (2019)
Ge J, Remesh SG, Hammel M, Pan S, Mahan AD, Wang S, Wang X
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.9 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: P11532 (1461-1973) Dystrophin (R11-15 human dystrophin fragment)
|
|
|
|
| Sample: |
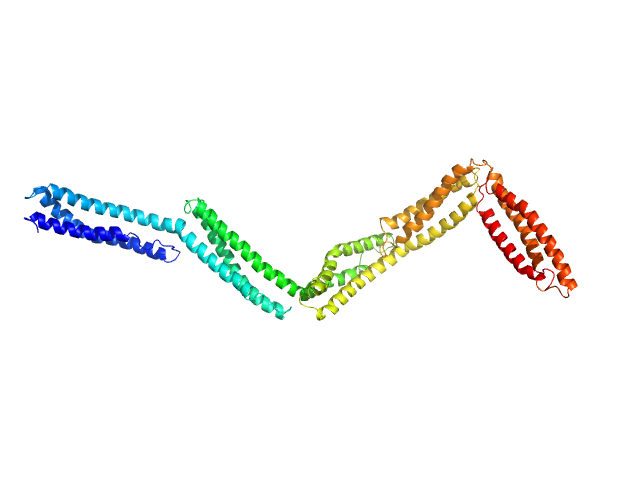
Dystrophin (R11-15 human dystrophin fragment) monomer, 60 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-d11, 150 mM NaCl, 0.1 mM EDTA-d16, in 100% v/v D2O, pH: 7.1 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Nov 1
|
How the central domain of dystrophin acts to bridge F-actin to sarcolemmal lipids.
J Struct Biol :107411 (2019)
Mias-Lucquin D, Dos Santos Morais R, Chéron A, Lagarrigue M, Winder SJ, Chenuel T, Pérez J, Appavou MS, Martel A, Alviset G, Le Rumeur E, Combet S, Hubert JF, Delalande O
|
| RgGuinier |
6.2 |
nm |
| Dmax |
27.4 |
nm |
| VolumePorod |
146 |
nm3 |
|
|
UniProt ID: Q9Y265 (1-456) RuvB-like 1
UniProt ID: Q9Y230 (1-463) RuvB-like 2
|
|
|

|
| Sample: |
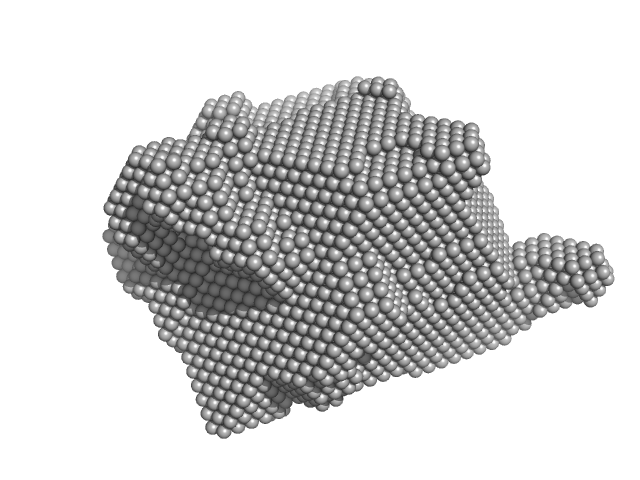
RuvB-like 1 hexamer, 311 kDa Homo sapiens protein
RuvB-like 2 hexamer, 311 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1% glycerol, 5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 21
|
Deciphering cellular and molecular determinants of human DPCD protein in complex with RUVBL1/RUVBL2 AAA-ATPases.
J Mol Biol :167760 (2022)
Dos Santos Morais R, Santo PE, Ley M, Schelcher C, Abel Y, Plassart L, Deslignière E, Chagot ME, Quinternet M, Paiva ACF, Hessmann S, Morellet N, M F Sousa P, Vandermoere F, Bertrand E, Charpentier B, Bandeiras TM, Plisson-Chastang C, Verheggen C, Cianférani S, Manival X
|
| RgGuinier |
6.2 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
1490 |
nm3 |
|
|
UniProt ID: O67206 (44-527) Aquifex aeolicus McoA metaloxidase ∆328-352 (MCoA∆328-352)
|
|
|

|
| Sample: |
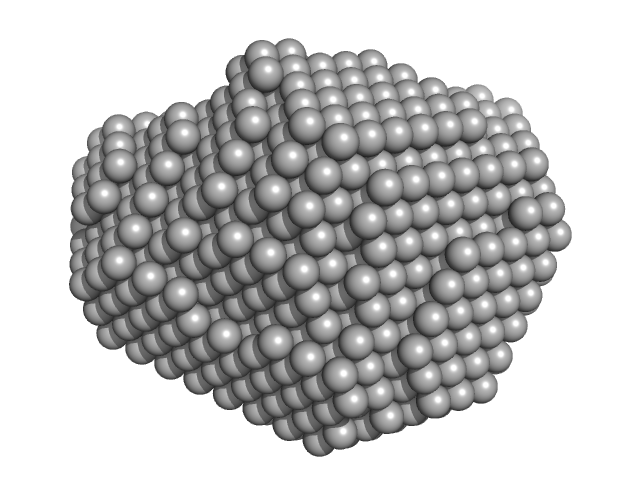
Aquifex aeolicus McoA metaloxidase ∆328-352 (MCoA∆328-352) monomer, 53 kDa Aquifex aeolicus protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jul 13
|
The Methionine-Rich Loop of Multicopper Oxidase McoA follows Open-To-Close Transitions with a Role in Enzyme Catalysis
ACS Catalysis (2020)
Borges P, Brissos V, Hernandez G, Masgrau L, Lucas M, Monza E, Frazão C, Cordeiro T, Martins L
|
| RgGuinier |
2.3 |
nm |
| Dmax |
6.9 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: P02769 (25-607) Bovine serum albumin
|
|
|

|
| Sample: |
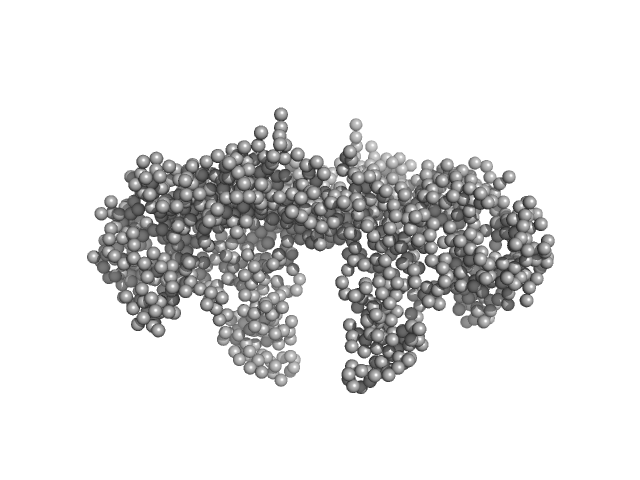
Bovine serum albumin dimer, 133 kDa Bos taurus protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 2% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Apr 5
|
Adding Size Exclusion Chromatography (SEC) and Light Scattering (LS) Devices to Obtain High-Quality Small Angle X-Ray Scattering (SAXS) Data
Crystals 10(11):975 (2020)
Graewert M, Da Vela S, Gräwert T, Molodenskiy D, Blanchet C, Svergun D, Jeffries C
|
| RgGuinier |
4.0 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
211 |
nm3 |
|
|
UniProt ID: O08967 (14-399) Cytohesin-3
|
|
|
|
| Sample: |
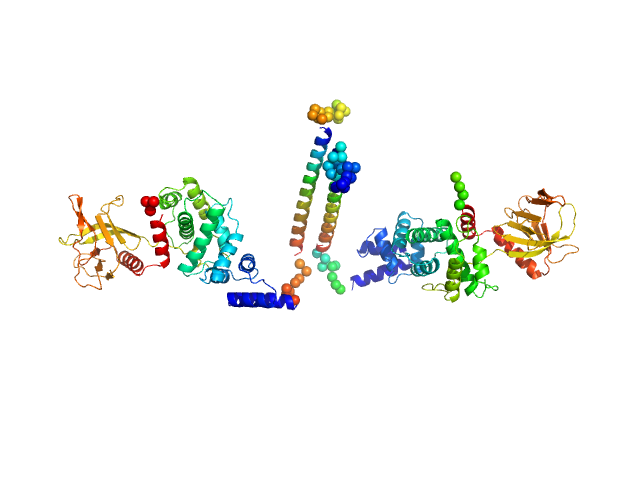
Cytohesin-3 dimer, 93 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2 mM MgCl2, 0.1% 2-mercaptoethanol, 5% glycerol, 0.001 mM insitol 1,3,4,5-tetrakis phosphate, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2013 Nov 15
|
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Structure (2019)
Das S, Malaby AW, Nawrotek A, Zhang W, Zeghouf M, Maslen S, Skehel M, Chakravarthy S, Irving TC, Bilsel O, Cherfils J, Lambright DG
|
| RgGuinier |
5.5 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
194 |
nm3 |
|
|
UniProt ID: P9WNV1 (1-328) DNA ligase A
|
|
|

|
| Sample: |
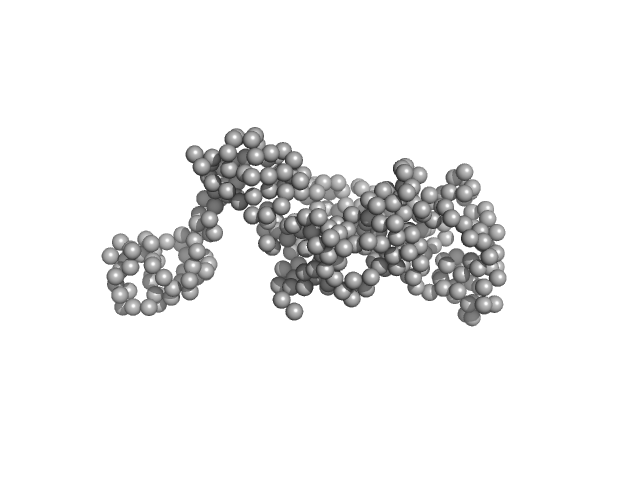
DNA ligase A monomer, 37 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl 200 mM NaCl 2mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2019 May 21
|
Salt bridges at the subdomain interfaces of the adenylation domain and active-site residues of Mycobacterium tuberculosis
NAD +
-dependent DNA ligase A (MtbLigA) are important for the initial steps of nick-sealing activity
Acta Crystallographica Section D Structural Biology 77(6) (2021)
Afsar M, Shukla A, Kumar N, Ramachandran R
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
68 |
nm3 |
|
|