UniProt ID: Q9BRS8 (1-83) La-related protein 6
|
|
|
|
| Sample: |
La-related protein 6 monomer, 9 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM KCl, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 1
|
An intrinsically disordered region mediates RNA-binding selectivity and cellular activities of LARP6.
Nat Commun (2026)
Capraro F, Abis G, Incocciati A, Simpson PJ, Karimzadeh M, Masino L, Barley A, Bui TTT, Kelly G, Goodarzi H, Conte MR, Mardakheh FK
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
21 |
nm3 |
|
|
UniProt ID: Q9BRS8 (70-300) La-related protein 6
UniProt ID: None (None-None) 3'-untranslated region of catenin alpha 1 mRNA
|
|
|
|
| Sample: |
La-related protein 6 monomer, 35 kDa Homo sapiens protein
3'-untranslated region of catenin alpha 1 mRNA, 8 kDa RNA
|
| Buffer: |
25 mM Tris-HCl, 100 mM KCl, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 1
|
An intrinsically disordered region mediates RNA-binding selectivity and cellular activities of LARP6.
Nat Commun (2026)
Capraro F, Abis G, Incocciati A, Simpson PJ, Karimzadeh M, Masino L, Barley A, Bui TTT, Kelly G, Goodarzi H, Conte MR, Mardakheh FK
|
| RgGuinier |
3.0 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
UniProt ID: Q9BRS8 (1-300) La-related protein 6
UniProt ID: None (None-None) 3'-untranslated region of catenin alpha 1 mRNA
|
|
|
|
| Sample: |
La-related protein 6 monomer, 42 kDa Homo sapiens protein
3'-untranslated region of catenin alpha 1 mRNA, 8 kDa RNA
|
| Buffer: |
25 mM Tris-HCl, 100 mM KCl, 5 mM MgCl2, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 1
|
An intrinsically disordered region mediates RNA-binding selectivity and cellular activities of LARP6.
Nat Commun (2026)
Capraro F, Abis G, Incocciati A, Simpson PJ, Karimzadeh M, Masino L, Barley A, Bui TTT, Kelly G, Goodarzi H, Conte MR, Mardakheh FK
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
138 |
nm3 |
|
|
UniProt ID: Q13283 (1-466) Ras GTPase-activating protein-binding protein 1
|
|
|

|
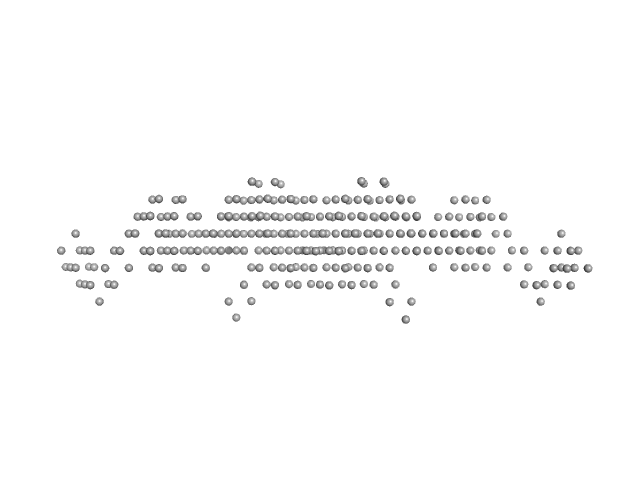
| Sample: |
Ras GTPase-activating protein-binding protein 1 dimer, 105 kDa Homo sapiens protein
|
| Buffer: |
50 mM MES, 150 mM NaCl, pH: 6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2025 Dec 27
|
Full-length G3BP1 (residues 1-466) at pH 6.0, 150 mM NaCl
|
| RgGuinier |
5.0 |
nm |
| Dmax |
22.5 |
nm |
| VolumePorod |
159 |
nm3 |
|
|
UniProt ID: Q13283 (1-413) RGG deletion mutant of Ras GTPase-activating protein-binding protein 1
|
|
|

|
| Sample: |
RGG deletion mutant of Ras GTPase-activating protein-binding protein 1 dimer, 94 kDa Homo sapiens protein
|
| Buffer: |
50 mM MES, 150 mM NaCl, pH: 6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2025 Dec 27
|
Full-length G3BP1 (residues 1-466) at pH 6.0, 150 mM NaCl
|
| RgGuinier |
6.0 |
nm |
| Dmax |
31.4 |
nm |
| VolumePorod |
193 |
nm3 |
|
|
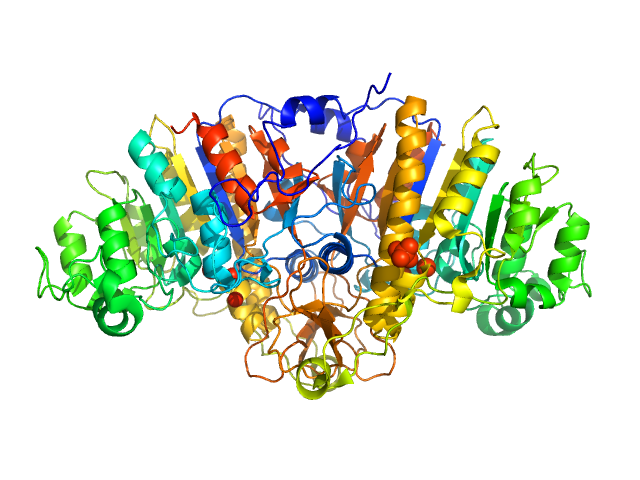
UniProt ID: P00634 (None-None) Wild Alkaline Phosphatase under vis
|
|
|
|
| Sample: |
Wild Alkaline Phosphatase under vis dimer, 97 kDa Escherichia coli protein
|
| Buffer: |
25 mM Tris, 100 mM NaCl, 2 mM TCEP, 3% (w/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 Dec 23
|
Single-engineered-residue solvation perturbations regulate global protein architecture and function.
Nat Commun (2026)
Liu Y, Zhai J, Cao S, Guo H, Zhang H, Song B, Guo J, Men D, He X, Li D
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
UniProt ID: Q815N5 (40-338) Ferric anguibactin-binding protein
|
|
|
|
| Sample: |
Ferric anguibactin-binding protein monomer, 32 kDa Bacillus cereus (strain … protein
|
| Buffer: |
20mM Tris-HCl, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2025 Apr 7
|
Structural basis of FatB-mediated iron uptake via tyrosine/histidine direct coordination accompanying long-distance domain reorganization.
Nat Commun (2026)
Lee H, Kim SO, You S, Segalina A, Noh T, Ihee H
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
UniProt ID: Q815N5 (40-338) Ferric anguibactin-binding protein
|
|
|
|
| Sample: |
Ferric anguibactin-binding protein monomer, 32 kDa Bacillus cereus (strain … protein
|
| Buffer: |
20mM Tris-HCl, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2025 Apr 7
|
Structural basis of FatB-mediated iron uptake via tyrosine/histidine direct coordination accompanying long-distance domain reorganization.
Nat Commun (2026)
Lee H, Kim SO, You S, Segalina A, Noh T, Ihee H
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
65 |
nm3 |
|
|
UniProt ID: Q815N5 (40-338) Ferric anguibactin-binding protein
|
|
|
|
| Sample: |
Ferric anguibactin-binding protein monomer, 32 kDa Bacillus cereus (strain … protein
|
| Buffer: |
20mM Tris-HCl, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2025 Apr 7
|
Structural basis of FatB-mediated iron uptake via tyrosine/histidine direct coordination accompanying long-distance domain reorganization.
Nat Commun (2026)
Lee H, Kim SO, You S, Segalina A, Noh T, Ihee H
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: Q815N5 (None-None) Ferric anguibactin-binding protein
|
|
|
|
| Sample: |
Ferric anguibactin-binding protein monomer, 32 kDa Bacillus cereus (strain … protein
|
| Buffer: |
20mM Tris-HCl, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2025 Apr 7
|
Structural basis of FatB-mediated iron uptake via tyrosine/histidine direct coordination accompanying long-distance domain reorganization.
Nat Commun (2026)
Lee H, Kim SO, You S, Segalina A, Noh T, Ihee H
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
57 |
nm3 |
|
|