UniProt ID: Q815N5 (None-None) Ferric anguibactin-binding protein
|
|
|
|
| Sample: |
Ferric anguibactin-binding protein monomer, 32 kDa Bacillus cereus (strain … protein
|
| Buffer: |
20mM Tris-HCl, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2025 Apr 7
|
Structural basis of FatB-mediated iron uptake via tyrosine/histidine direct coordination accompanying long-distance domain reorganization.
Nat Commun (2026)
Lee H, Kim SO, You S, Segalina A, Noh T, Ihee H
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
42 |
nm3 |
|
|
UniProt ID: Q9WYC4 (None-None) ABC transporter TM288 subunit
UniProt ID: Q9WYC3 (None-None) ABC transporter, ATP-binding protein
|
|
|
|
| Sample: |
ABC transporter TM288 subunit monomer, 68 kDa Thermotoga maritima (strain … protein
ABC transporter, ATP-binding protein monomer, 64 kDa ABC transporter TM287 … protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 5 mM MgCl2, 20 μM DMM, 1 mM Mg2+ ATP, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Sep 10
|
Capturing transient states of heterodimeric ABC transporter TM287/288 by Time-Resolved Small-Angle X-ray Scattering.
Biophys J (2026)
Schröder L, De Vecchis D, Gruzinov A, Schäfer LV, Blanchet CE, Seeger MA, Tidow H, Josts I
|
|
|
UniProt ID: Q9WYC4 (None-None) ABC transporter TM288 subunit
UniProt ID: Q9WYC3 (None-None) ABC transporter, ATP-binding protein
UniProt ID: None (None-None) Nanobody Nb_TM#1 bound to TM287/288 ABC transporter
|
|
|
|
| Sample: |
ABC transporter TM288 subunit monomer, 68 kDa Thermotoga maritima (strain … protein
ABC transporter, ATP-binding protein monomer, 64 kDa ABC transporter TM287 … protein
Nanobody Nb_TM#1 bound to TM287/288 ABC transporter monomer, 14 kDa Vicugna pacos (alpaca)
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 5 mM MgCl2, 20 μM DMM, 1 mM Mg2+ ATP, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Sep 10
|
Capturing transient states of heterodimeric ABC transporter TM287/288 by Time-Resolved Small-Angle X-ray Scattering.
Biophys J (2026)
Schröder L, De Vecchis D, Gruzinov A, Schäfer LV, Blanchet CE, Seeger MA, Tidow H, Josts I
|
|
|
UniProt ID: Q9WYC4 (None-None) ABC transporter TM288 subunit
UniProt ID: Q9WYC3 (None-None) ABC transporter, ATP-binding protein
UniProt ID: None (None-None) Synthetic nanobody Sb_TM#35 bound to TM287/288 ABC transporter
|
|
|
|
| Sample: |
ABC transporter TM288 subunit monomer, 68 kDa Thermotoga maritima (strain … protein
ABC transporter, ATP-binding protein monomer, 64 kDa ABC transporter TM287 … protein
Synthetic nanobody Sb_TM#35 bound to TM287/288 ABC transporter monomer, 14 kDa synthetic (in vitro …
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 5 mM MgCl2, 20 μM DMM, 1 mM Mg2+ ATP, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Jun 6
|
Capturing transient states of heterodimeric ABC transporter TM287/288 by Time-Resolved Small-Angle X-ray Scattering.
Biophys J (2026)
Schröder L, De Vecchis D, Gruzinov A, Schäfer LV, Blanchet CE, Seeger MA, Tidow H, Josts I
|
|
|
UniProt ID: A0A0H2URK1 (187-385) Functional binding region (187-385) of the pneumococcal serine-rich repeat protein
|
|
|
|
| Sample: |
Functional binding region (187-385) of the pneumococcal serine-rich repeat protein monomer, 22 kDa Streptococcus pneumoniae protein
|
| Buffer: |
PBS 5 % Glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2013 Feb 27
|
The BR domain of PsrP interacts with extracellular DNA to promote bacterial aggregation; structural insights into pneumococcal biofilm formation.
Sci Rep 6:32371 (2016)
Schulte T, Mikaelsson C, Beaussart A, Kikhney A, Deshmukh M, Wolniak S, Pathak A, Ebel C, Löfling J, Fogolari F, Henriques-Normark B, Dufrêne YF, Svergun D, Nygren PÅ, Achour A
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
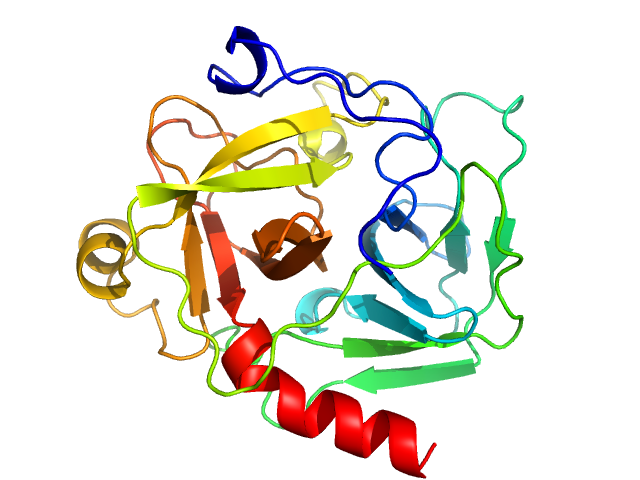
UniProt ID: P00766 (None-None) Chymotrypsinogen A
|
|
|
|
| Sample: |
Chymotrypsinogen A monomer, 26 kDa Bos taurus protein
|
| Buffer: |
100 mM Tris/HCl 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 May 19
|
Accuracy of molecular mass determination of proteins in solution by small-angle X-ray scattering
Journal of Applied Crystallography 40(s1):s245-s249 (2007)
Mylonas E, Svergun D
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.0 |
nm |
|
|
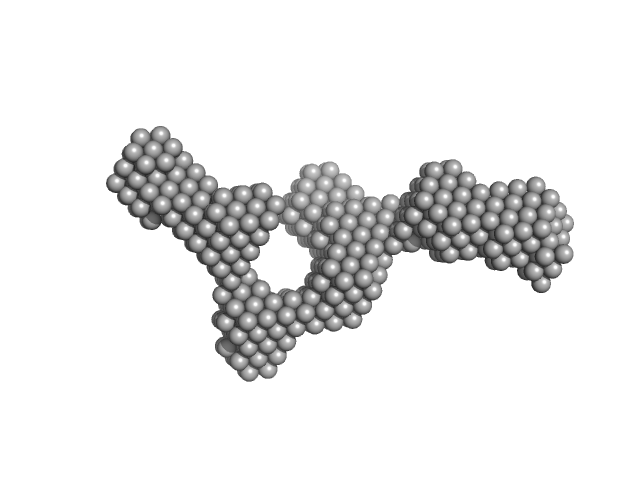
UniProt ID: M4N8T6 (None-None) S-layer protein
|
|
|

|
| Sample: |
S-layer protein monomer, 116 kDa Lysinibacillus sphaericus protein
|
| Buffer: |
Water, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Jun 2
|
Analysis of self-assembly of S-layer protein slp-B53 from Lysinibacillus sphaericus.
Eur Biophys J 46(1):77-89 (2017)
Liu J, Falke S, Drobot B, Oberthuer D, Kikhney A, Guenther T, Fahmy K, Svergun D, Betzel C, Raff J
|
| RgGuinier |
5.8 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
495 |
nm3 |
|
|
UniProt ID: A0A0D7C2L1 (1-92) Plasmid stabilization protein ParE
UniProt ID: A0A0F6F6Q9 (14-75) Uncharacterized protein (Antitoxin)
|
|
|
|
| Sample: |
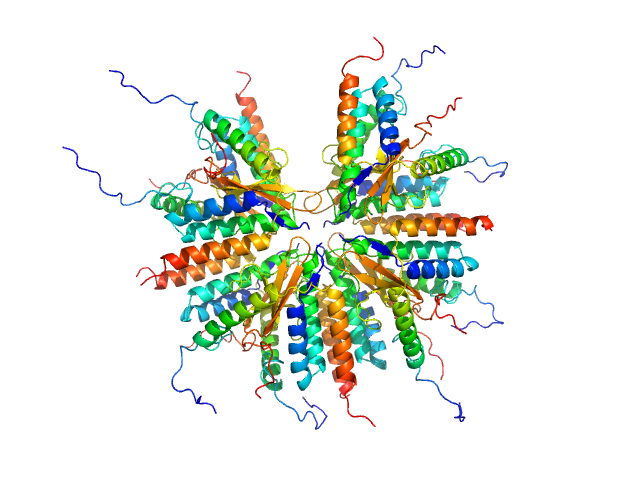
Plasmid stabilization protein ParE 16-mer, 188 kDa Escherichia coli protein
Uncharacterized protein (Antitoxin) 16-mer, 135 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Dec 9
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
3.8 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
312 |
nm3 |
|
|
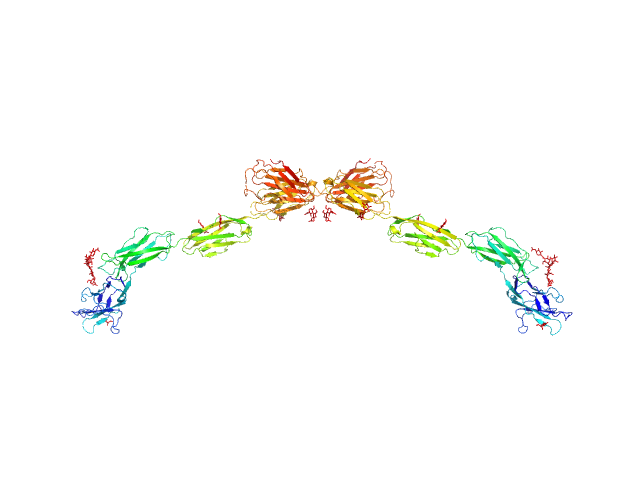
UniProt ID: P20917 (20-508) Myelin-associated glycoprotein Ig domains 1-5
|
|
|
|
| Sample: |
Myelin-associated glycoprotein Ig domains 1-5 dimer, 108 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 11
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
6.8 |
nm |
| Dmax |
23.8 |
nm |
| VolumePorod |
177 |
nm3 |
|
|
UniProt ID: Q15185 (1-160) Prostaglandin E synthase 3
|
|
|
|
| Sample: |
Prostaglandin E synthase 3 monomer, 19 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM NaCl, 5 mM B-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS1 Beamline, Brazilian Synchrotron Light Laboratory on 2012 Jun 22
|
The C-terminal region of the human p23 chaperone modulates its structure and function.
Arch Biochem Biophys 565:57-67 (2015)
Seraphim TV, Gava LM, Mokry DZ, Cagliari TC, Barbosa LR, Ramos CH, Borges JC
|
| RgGuinier |
2.5 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
40 |
nm3 |
|
|