UniProt ID: P31483-2 (93-274) Nucleolysin TIA-1 isoform p40
UniProt ID: (None-None) RNA (UUUUUACU)
|
|
|
|
| Sample: |
Nucleolysin TIA-1 isoform p40 monomer, 21 kDa Homo sapiens protein
RNA (UUUUUACU) monomer, 1 kDa RNA
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 May 27
|
TIA-1 RRM23 binding and recognition of target oligonucleotides.
Nucleic Acids Res 45(8):4944-4957 (2017)
Waris S, García-Mauriño SM, Sivakumaran A, Beckham SA, Loughlin FE, Gorospe M, Díaz-Moreno I, Wilce MCJ, Wilce JA
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
UniProt ID: P12004 (None-None) Proliferating cell nuclear antigen
|
|
|
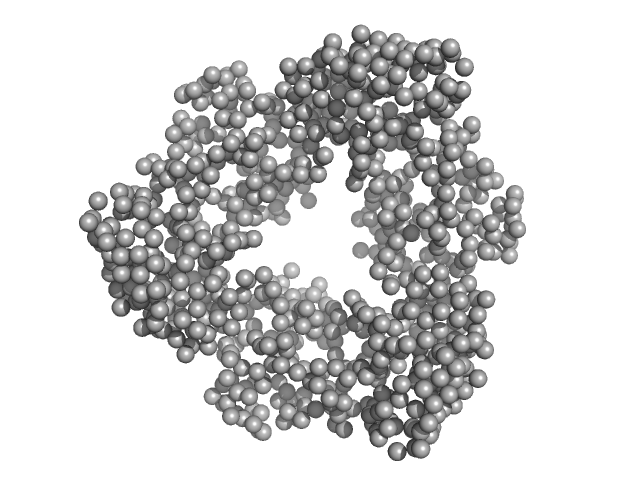
|
| Sample: |
Proliferating cell nuclear antigen trimer, 89 kDa Homo sapiens protein
|
| Buffer: |
25mM HEPES 100mM NaCl 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Jan 20
|
Destabilization of the PCNA trimer mediated by its interaction with the NEIL1 DNA glycosylase.
Nucleic Acids Res 45(5):2897-2909 (2017)
Prakash A, Moharana K, Wallace SS, Doublié S
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
128 |
nm3 |
|
|
UniProt ID: Q9RS64 (51-207) N-terminal truncated DNA protection during starvation protein 1
|
|
|

|
| Sample: |
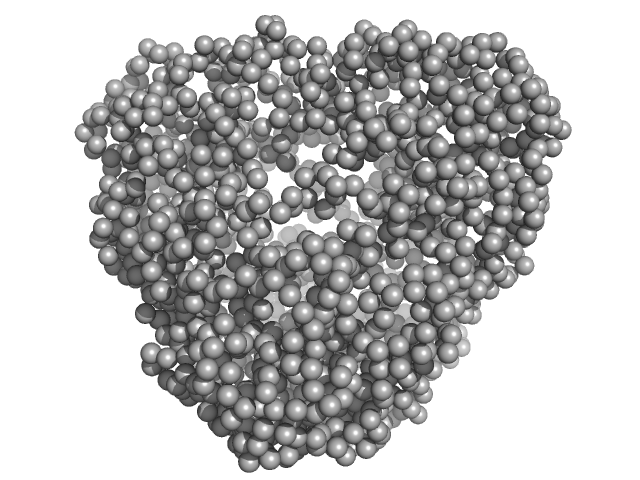
N-terminal truncated DNA protection during starvation protein 1 dodecamer, 216 kDa Deinococcus radiodurans R1 protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2013 Nov 22
|
SAXS Structural Studies of Dps from Deinococcus radiodurans Highlights the Conformation of the Mobile N-Terminal Extensions.
J Mol Biol 429(5):667-687 (2017)
Santos SP, Cuypers MG, Round A, Finet S, Narayanan T, Mitchell EP, Romão CV
|
| RgGuinier |
3.9 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
291 |
nm3 |
|
|
UniProt ID: P08392 (258-487) Major viral transcription factor ICP4
|
|
|
|
| Sample: |
Major viral transcription factor ICP4 dimer, 49 kDa Human alphaherpesvirus 1 protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2015 Jul 27
|
The herpes viral transcription factor ICP4 forms a novel DNA recognition complex.
Nucleic Acids Res 45(13):8064-8078 (2017)
Tunnicliffe RB, Lockhart-Cairns MP, Levy C, Mould AP, Jowitt TA, Sito H, Baldock C, Sandri-Goldin RM, Golovanov AP
|
| RgGuinier |
2.9 |
nm |
| Dmax |
12.7 |
nm |
|
|
UniProt ID: P39685 (None-None) Nucleoporin POM152
|
|
|

|
| Sample: |
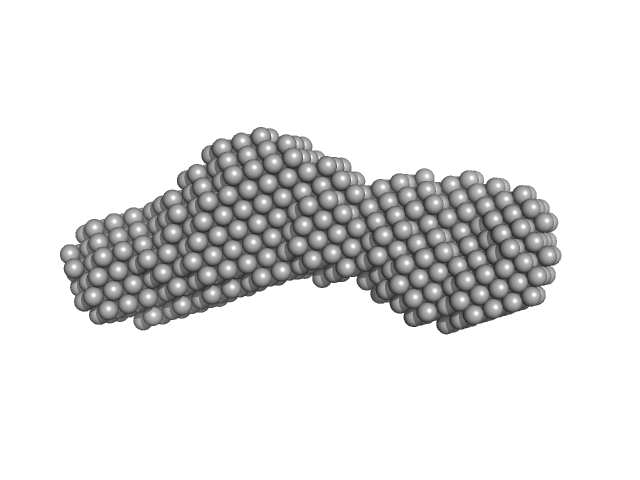
Nucleoporin POM152 monomer, 24 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 10%(v/v) glycerol, 5mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2015 Apr 12
|
Molecular Architecture of the Major Membrane Ring Component of the Nuclear Pore Complex.
Structure 25(3):434-445 (2017)
Upla P, Kim SJ, Sampathkumar P, Dutta K, Cahill SM, Chemmama IE, Williams R, Bonanno JB, Rice WJ, Stokes DL, Cowburn D, Almo SC, Sali A, Rout MP, Fernandez-Martinez J
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
UniProt ID: P0A9Q8 (451-891) Aldehyde-alcohol dehydrogenase
|
|
|

|
| Sample: |
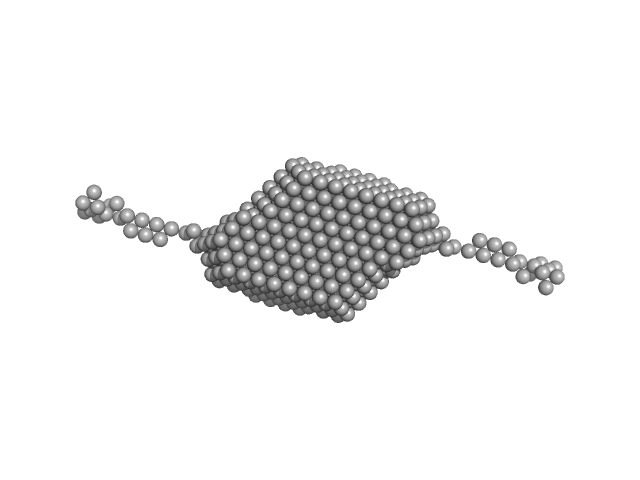
Aldehyde-alcohol dehydrogenase dimer, 97 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
20 mM Tris 400 mM NaCl 5% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2015 Nov 27
|
High-resolution structure of the alcohol dehydrogenase domain of the bifunctional bacterial enzyme AdhE.
Acta Crystallogr F Struct Biol Commun 76(Pt 9):414-421 (2020)
Azmi L, Bragginton EC, Cadby IT, Byron O, Roe AJ, Lovering AL, Gabrielsen M
|
| RgGuinier |
3.2 |
nm |
| Dmax |
16.9 |
nm |
| VolumePorod |
146 |
nm3 |
|
|
UniProt ID: Q15059 (20-418) Bromodomain-containing protein 3
|
|
|
|
| Sample: |
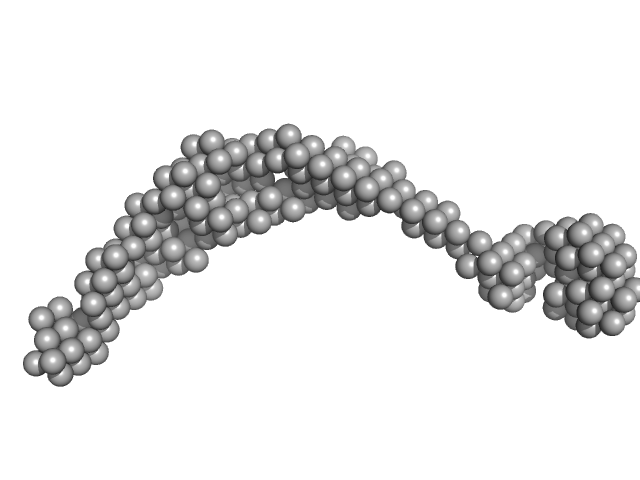
Bromodomain-containing protein 3 monomer, 44 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2% glycerol, 0.5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Jan 13
|
Interactome Rewiring Following Pharmacological Targeting of BET Bromodomains.
Mol Cell (2018)
Lambert JP, Picaud S, Fujisawa T, Hou H, Savitsky P, Uusküla-Reimand L, Gupta GD, Abdouni H, Lin ZY, Tucholska M, Knight JDR, Gonzalez-Badillo B, St-Denis N, Newman JA, Stucki M, Pelletier L, Bandeira N, Wilson MD, Filippakopoulos P, Gingras AC
|
| RgGuinier |
6.2 |
nm |
| Dmax |
21.5 |
nm |
| VolumePorod |
210 |
nm3 |
|
|
UniProt ID: Q9X721 (787-1118) Collagenase ColG segement s2s3as3b
|
|
|

|
| Sample: |
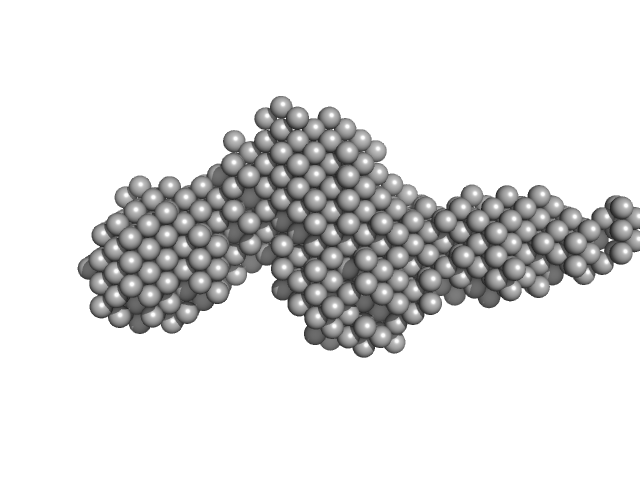
Collagenase ColG segement s2s3as3b monomer, 37 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 11
|
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
3.0 |
nm |
| Dmax |
14.4 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
UniProt ID: P9WNV1 (1-691) DNA ligase A
|
|
|

|
| Sample: |
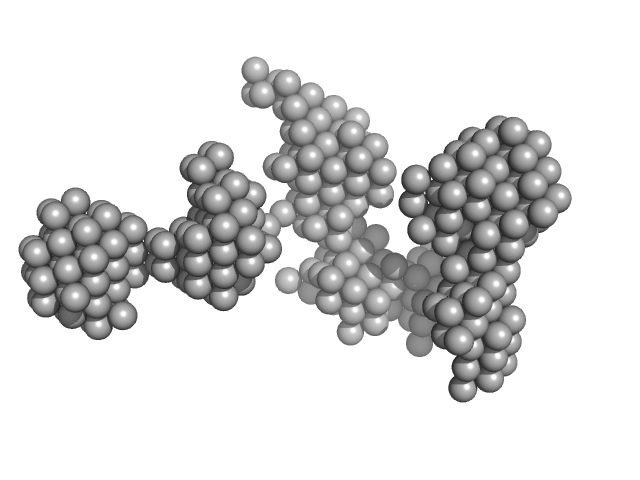
DNA ligase A monomer, 76 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 13
|
M. tuberculosis class II apurinic/ apyrimidinic-endonuclease/3'-5' exonuclease (XthA) engages with NAD+-dependent DNA ligase A (LigA) to counter futile cleavage and ligation cycles in base excision repair.
Nucleic Acids Res (2020)
Khanam T, Afsar M, Shukla A, Alam F, Kumar S, Soyar H, Dolma K, Pasupuleti M, Srivastava KK, Ampapathi RS, Ramachandran R
|
| RgGuinier |
5.2 |
nm |
| Dmax |
16.7 |
nm |
| VolumePorod |
870 |
nm3 |
|
|
UniProt ID: Q911W0 (478-919) Tegument protein UL37
|
|
|
|
| Sample: |
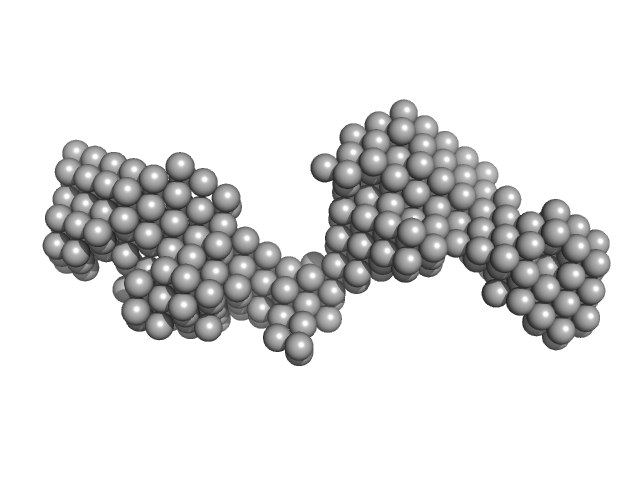
Tegument protein UL37 monomer, 49 kDa Suid alphaherpesvirus 1 protein
|
| Buffer: |
100 mM HEPES 150 mM NaCl 5% glycerol 0.1 mM tris(2-carboxyethyl)phosphine (TCEP), pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2017 Jun 3
|
The dynamic nature of the conserved tegument protein UL37 of herpesviruses.
J Biol Chem 293(41):15827-15839 (2018)
Koenigsberg AL, Heldwein EE
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
71 |
nm3 |
|
|