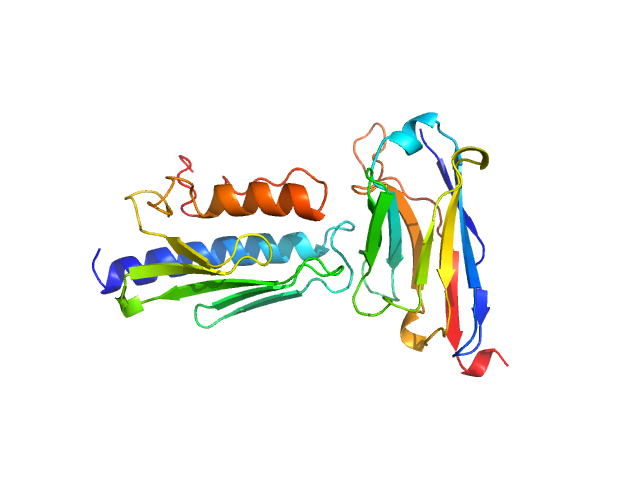
UniProt ID: Q16595 (90-210) Frataxin, mitochondrial
UniProt ID: None (1-139) Nanobody 16C10
|
|
|
|
| Sample: |
Frataxin, mitochondrial monomer, 14 kDa Homo sapiens protein
Nanobody 16C10 monomer, 15 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Sep 18
|
Nanobodies as tools for studying human frataxin biology.
Commun Biol (2026)
Pignataro MF, Fernández NB, Garay-Alvarez A, Pavan MF, Molina R, Muñoz IG, Grossi J, Noguera M, Vila A, García AE, Gentili HG, Rodríguez NA, Aran M, Parreño V, Bok M, Hermoso JA, Ibañez LI, Santos J
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
37 |
nm3 |
|
|
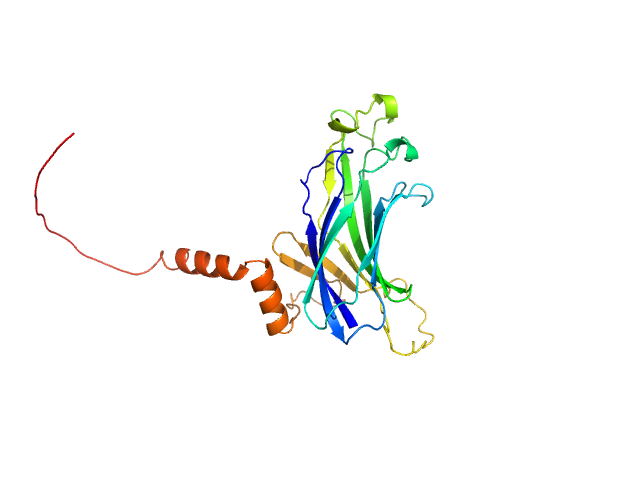
UniProt ID: C4LX08 (1-234) Anti-Silencing Function 1
|
|
|
|
| Sample: |
Anti-Silencing Function 1 monomer, 28 kDa Entamoeba histolytica strain … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, and 2 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Oct 1
|
Characterisation of Entamoeba histolytica Anti-Silencing Function 1 as a histone chaperone.
Biochimie (2025)
Gandhi S, Vasudevan D
|
| RgGuinier |
3.5 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
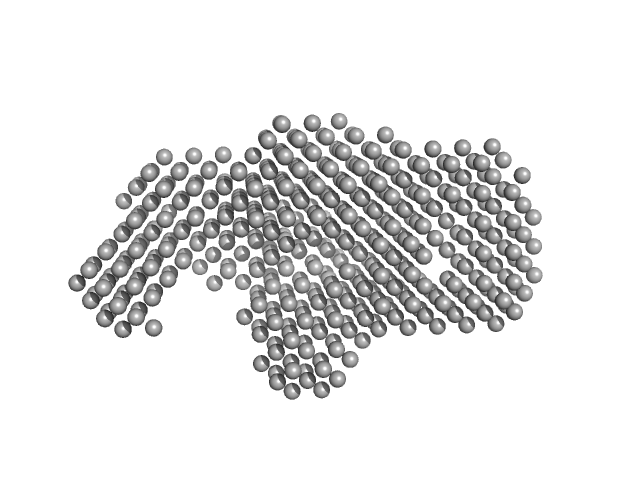
UniProt ID: P18472 (19-210) TraW (Δ1-18)
|
|
|

|
| Sample: |
TraW (Δ1-18) monomer, 26 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 5% glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Feb 23
|
Solution characterization of TraW, a regulatory protein of the F plasmid type 4 secretion system
Structural Dynamics 13(2) (2026)
Rodriguez C, Audette G
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
|
|
UniProt ID: P53893 (1-1110) Ribosome assembly protein 1 with C-terminal tag (RSRSGSENLYFQGSHHHHHHHH)
|
|
|
|
| Sample: |
Ribosome assembly protein 1 with C-terminal tag (RSRSGSENLYFQGSHHHHHHHH) monomer, 127 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM HEPES, 300 mM NaCl, 5 mM MgCl2, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 3
|
Hydroxyl radical footprinting modification reveals an intradomain communication pathway in EFL1 disrupted by a Shwachman-Diamond syndrome-associated mutation.
Protein Sci 35(4):e70504 (2026)
Zúñiga-Domínguez JA, Jain R, González-Andrade M, Farquhar ER, Chance MR, Gijsbers A, Sánchez-Puig N
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.7 |
nm |
| VolumePorod |
368 |
nm3 |
|
|
UniProt ID: A0A0H2URK1 (120-395) Functional binding region (120-395) of the pneumococcal serine-rich repeat protein
|
|
|
|
| Sample: |
Functional binding region (120-395) of the pneumococcal serine-rich repeat protein monomer, 30 kDa Streptococcus pneumoniae protein
|
| Buffer: |
PBS 5 % Glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2013 Feb 27
|
The BR domain of PsrP interacts with extracellular DNA to promote bacterial aggregation; structural insights into pneumococcal biofilm formation.
Sci Rep 6:32371 (2016)
Schulte T, Mikaelsson C, Beaussart A, Kikhney A, Deshmukh M, Wolniak S, Pathak A, Ebel C, Löfling J, Fogolari F, Henriques-Normark B, Dufrêne YF, Svergun D, Nygren PÅ, Achour A
|
| RgGuinier |
2.9 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
41 |
nm3 |
|
|
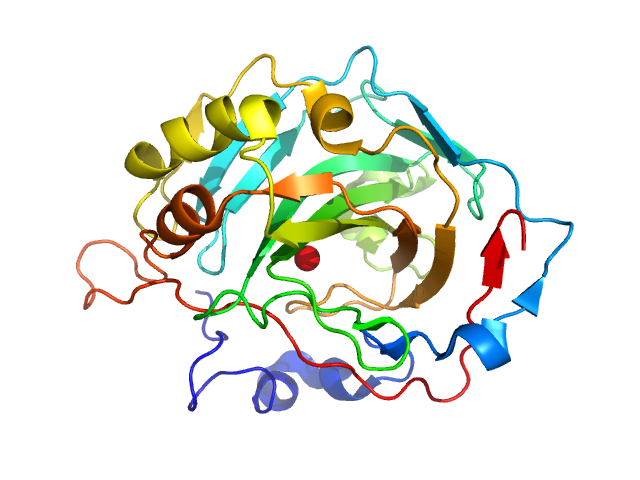
UniProt ID: P00921 (None-None) Carbonic anhydrase 2
|
|
|
|
| Sample: |
Carbonic anhydrase 2 monomer, 29 kDa Bos taurus protein
|
| Buffer: |
100 mM Tris/HCl 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 May 19
|
Accuracy of molecular mass determination of proteins in solution by small-angle X-ray scattering
Journal of Applied Crystallography 40(s1):s245-s249 (2007)
Mylonas E, Svergun D
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.0 |
nm |
|
|
UniProt ID: M4N8T6 (None-None) S-layer protein
|
|
|
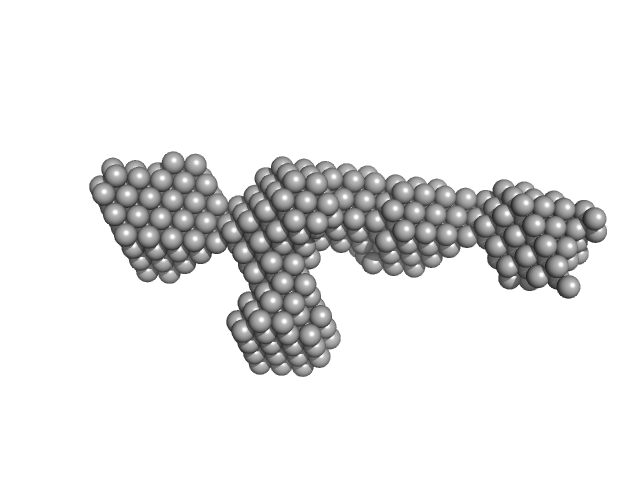
|
| Sample: |
S-layer protein monomer, 116 kDa Lysinibacillus sphaericus protein
|
| Buffer: |
Water with Ca2+, pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Jun 2
|
Analysis of self-assembly of S-layer protein slp-B53 from Lysinibacillus sphaericus.
Eur Biophys J 46(1):77-89 (2017)
Liu J, Falke S, Drobot B, Oberthuer D, Kikhney A, Guenther T, Fahmy K, Svergun D, Betzel C, Raff J
|
| RgGuinier |
6.4 |
nm |
| Dmax |
28.1 |
nm |
| VolumePorod |
609 |
nm3 |
|
|
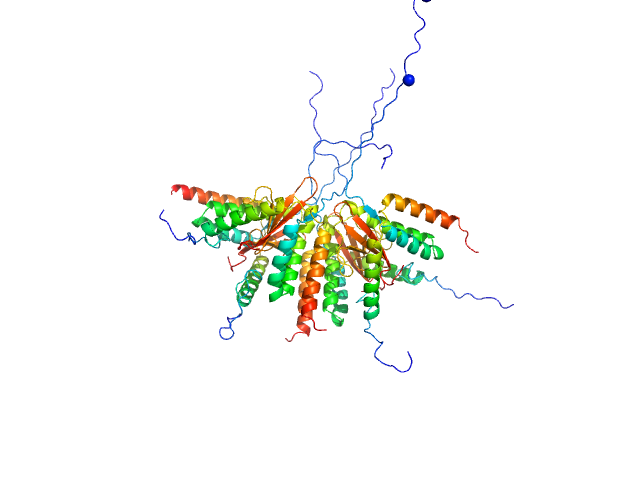
UniProt ID: A0A0D7C2L1 (2-92) Plasmid stabilization protein ParE
UniProt ID: A0A0F6F6Q9 (14-75) Uncharacterized protein (Antitoxin)
|
|
|
|
| Sample: |
Plasmid stabilization protein ParE octamer, 102 kDa Escherichia coli protein
Uncharacterized protein (Antitoxin) octamer, 68 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 Feb 5
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
3.3 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
166 |
nm3 |
|
|
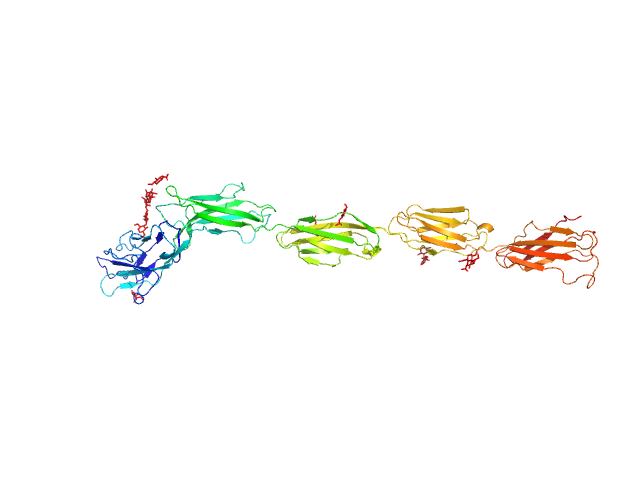
UniProt ID: P20917 (20-508) Myelin-associated glycoprotein (20-508; I473E mutant)
|
|
|
|
| Sample: |
Myelin-associated glycoprotein (20-508; I473E mutant) monomer, 54 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 11
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
6.0 |
nm |
| Dmax |
20.6 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
UniProt ID: Q15185 (1-142) Prostaglandin E synthase 3 (1-142)
|
|
|
|
| Sample: |
Prostaglandin E synthase 3 (1-142) monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM NaCl, 5 mM B-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS1 Beamline, Brazilian Synchrotron Light Laboratory on 2012 Jun 22
|
The C-terminal region of the human p23 chaperone modulates its structure and function.
Arch Biochem Biophys 565:57-67 (2015)
Seraphim TV, Gava LM, Mokry DZ, Cagliari TC, Barbosa LR, Ramos CH, Borges JC
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
36 |
nm3 |
|
|