UniProt ID: P10408 (None-None) Protein translocase subunit SecA
|
|
|
|
| Sample: |
Protein translocase subunit SecA dimer, 199 kDa Escherichia coli protein
|
| Buffer: |
20mM HEPES, 100mM NaCl, 1mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jul 18
|
The C-terminal tail of the bacterial translocation ATPase SecA modulates its activity.
Elife 8 (2019)
Jamshad M, Knowles TJ, White SA, Ward DG, Mohammed F, Rahman KF, Wynne M, Hughes GW, Kramer G, Bukau B, Huber D
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
380 |
nm3 |
|
|
UniProt ID: Q9LM76 (None-None) Senescence-associated E3 ubiquitin ligase 1
|
|
|
|
| Sample: |
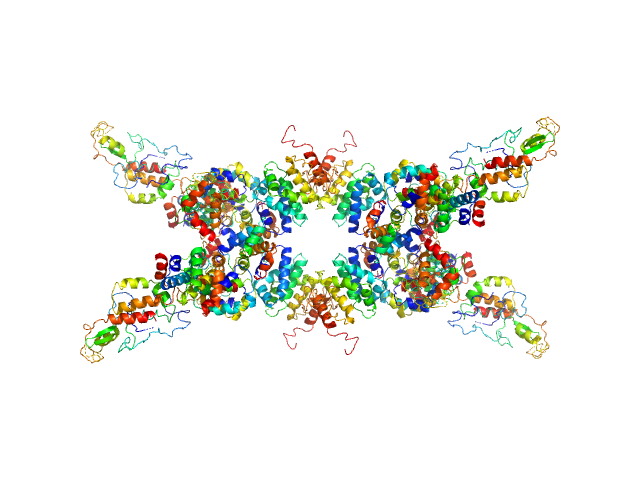
Senescence-associated E3 ubiquitin ligase 1 tetramer, 355 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM Tris, 250 mM NaCl, pH: 9 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Jun 2
|
Senescence-associated ubiquitin ligase 1 (SAUL1)
Haifa El Kilani,
Al Kikhney
|
| RgGuinier |
5.9 |
nm |
| Dmax |
22.0 |
nm |
|
|
UniProt ID: A0A1Z1SYD5 (22-243) Suppressor of Copper Sensitivity C protein
|
|
|
|
| Sample: |
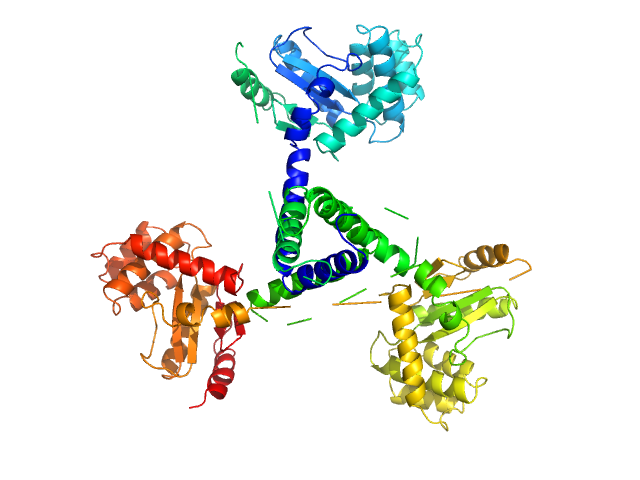
Suppressor of Copper Sensitivity C protein trimer, 74 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Acta Crystallogr D Struct Biol 75(Pt 3):296-307 (2019)
Furlong EJ, Kurth F, Premkumar L, Whitten AE, Martin JL
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
UniProt ID: P01308 (25-110) Insulin detemir (Levemir(R), Novo Nordisk A/S)
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
|
|
|
|
| Sample: |
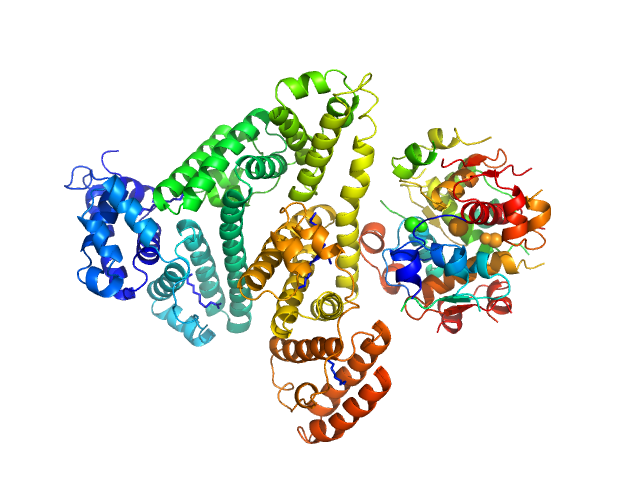
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
UniProt ID: P9WNV1 (2-328) DNA ligase A
|
|
|

|
| Sample: |
DNA ligase A monomer, 38 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2018 Oct 6
|
Salt bridges at the subdomain interfaces of the adenylation domain and active-site residues of Mycobacterium tuberculosis
NAD +
-dependent DNA ligase A (MtbLigA) are important for the initial steps of nick-sealing activity
Acta Crystallographica Section D Structural Biology 77(6) (2021)
Afsar M, Shukla A, Kumar N, Ramachandran R
|
| RgGuinier |
2.4 |
nm |
| Dmax |
6.2 |
nm |
| VolumePorod |
94 |
nm3 |
|
|
UniProt ID: P21333 (478-766) Filamin A Ig.like domains 3-5 P637Q mutant
|
|
|
|
| Sample: |
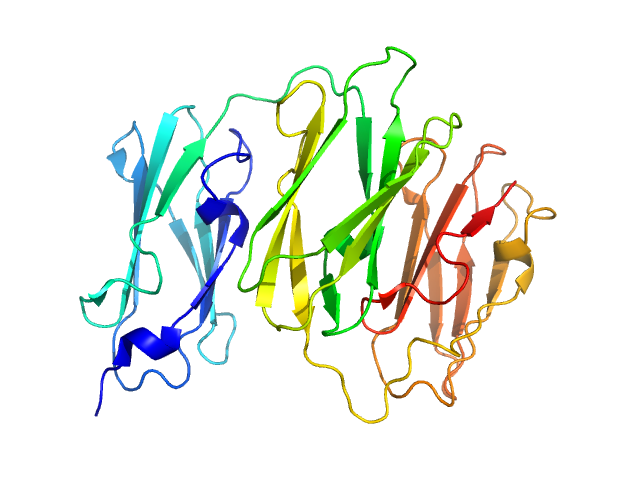
Filamin A Ig.like domains 3-5 P637Q mutant monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Non-syndromic Mitral Valve Dysplasia Mutation Changes the Force Resilience and Interaction of Human Filamin A.
Structure (2018)
Haataja TJK, Bernardi RC, Lecointe S, Capoulade R, Merot J, Pentikäinen U
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
UniProt ID: Q13263 (23-812) Transcription intermediary factor 1-beta, TIF1b, KAP1, TRIM28; amino acids 23-812
|
|
|
|
| Sample: |
Transcription intermediary factor 1-beta, TIF1b, KAP1, TRIM28; amino acids 23-812 dimer, 175 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 500 mM NaCl, 10 % Glycerol, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jul 6
|
KAP1 is an antiparallel dimer with a functional asymmetry.
Life Sci Alliance 2(4) (2019)
Fonti G, Marcaida MJ, Bryan LC, Träger S, Kalantzi AS, Helleboid PJ, Demurtas D, Tully MD, Grudinin S, Trono D, Fierz B, Dal Peraro M
|
| RgGuinier |
8.9 |
nm |
| Dmax |
38.0 |
nm |
| VolumePorod |
853 |
nm3 |
|
|
UniProt ID: Q4J9K8 (35-164) Conserved flagellar protein F
UniProt ID: Q4J9K7 (32-151) Stator protein FlaG soluble domain
|
|
|
|
| Sample: |
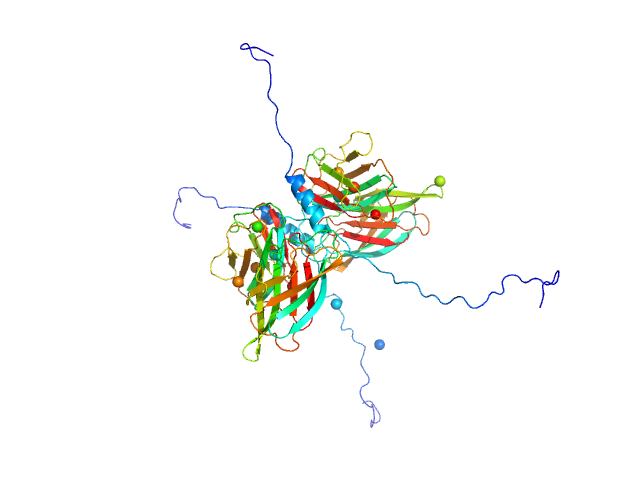
Conserved flagellar protein F dimer, 32 kDa Sulfolobus acidocaldarius protein
Stator protein FlaG soluble domain dimer, 30 kDa Sulfolobus acidocaldarius protein
|
| Buffer: |
25 mM citric acid/sodium citrate, 150mM NaCl, 3% Glycerol, pH: 3 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Nov 10
|
The structure of the periplasmic FlaG-FlaF complex and its essential role for archaellar swimming motility.
Nat Microbiol (2019)
Tsai CL, Tripp P, Sivabalasarma S, Zhang C, Rodriguez-Franco M, Wipfler RL, Chaudhury P, Banerjee A, Beeby M, Whitaker RJ, Tainer JA, Albers SV
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
109 |
nm3 |
|
|
UniProt ID: Q07953 (None-None) Ribosome maturation protein SDO1
|
|
|
|
| Sample: |
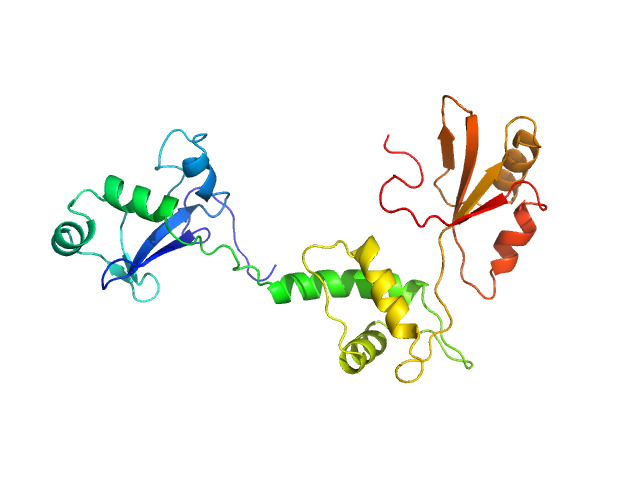
Ribosome maturation protein SDO1 monomer, 28 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Sep 21
|
Interaction of the GTPase Elongation Factor Like-1 with the Shwachman-Diamond Syndrome Protein and Its Missense Mutations.
Int J Mol Sci 19(12) (2018)
Gijsbers A, Montagut DC, Méndez-Godoy A, Altamura D, Saviano M, Siliqi D, Sánchez-Puig N
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.5 |
nm |
|
|
UniProt ID: P27930 (14-341) Interleukin-1 receptor type 2
|
|
|
|
| Sample: |
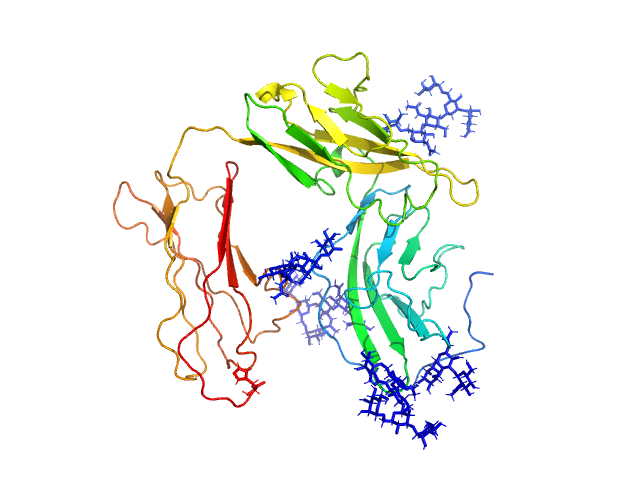
Interleukin-1 receptor type 2 monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 3% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2017 Jul 24
|
Functional Relevance of Interleukin-1 Receptor Inter-domain Flexibility for Cytokine Binding and Signaling.
Structure 27(8):1296-1307.e5 (2019)
Ge J, Remesh SG, Hammel M, Pan S, Mahan AD, Wang S, Wang X
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
80 |
nm3 |
|
|