UniProt ID: P61981 (1-234) 14-3-3 protein gamma
UniProt ID: Q96RR4 (93-517) Calcium/calmodulin-dependent protein kinase kinase 2
|
|
|
|
| Sample: |
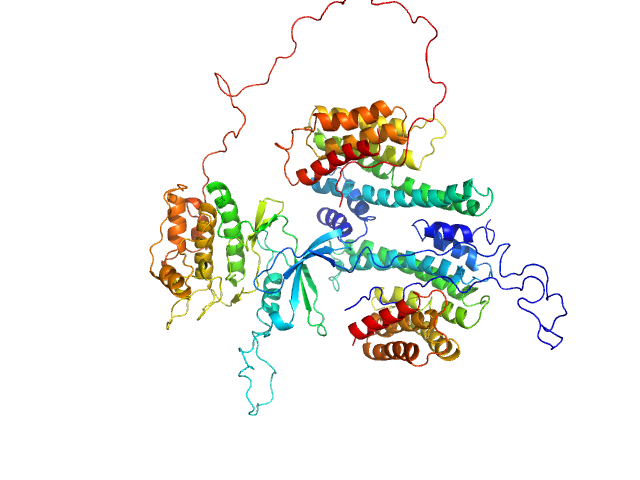
14-3-3 protein gamma dimer, 54 kDa Homo sapiens protein
Calcium/calmodulin-dependent protein kinase kinase 2 monomer, 48 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM TCEP, 3% (w/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Oct 23
|
14-3-3 protein inhibits CaMKK1 by blocking the kinase active site with its last two C-terminal helices.
Protein Sci :e4805 (2023)
Petrvalska O, Honzejkova K, Koupilova N, Herman P, Obsilova V, Obsil T
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
157 |
nm3 |
|
|
UniProt ID: Q9WTS6-2 (343-2699) Isoform A0B0 of Teneurin-3
|
|
|
|
| Sample: |
Isoform A0B0 of Teneurin-3 dimer, 526 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM EDTA, pH: 7.8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Sep 10
|
Alternative splicing controls teneurin-3 compact dimer formation for neuronal recognition
Nature Communications 15(1) (2024)
Gogou C, Beugelink J, Frias C, Kresik L, Jaroszynska N, Drescher U, Janssen B, Hindges R, Meijer D
|
| RgGuinier |
8.6 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
990 |
nm3 |
|
|
UniProt ID: P0DTC9 (1-419) Nucleoprotein
|
|
|
|
| Sample: |
Nucleoprotein tetramer, 187 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 13
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
6.9 |
nm |
| Dmax |
28.5 |
nm |
| VolumePorod |
571 |
nm3 |
|
|
UniProt ID: Q9BUE6 (2-129) Human Iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Human Iron-sulfur cluster assembly 1 homolog, mitochondrial, 14 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2023 Dec 1
|
A hidden property of the iron-sulfur protein in the mononuclear iron-bound state: species-dependent structural ordering induced by magnetic fields.
FEBS J (2025)
Arai S, Soga S, Hirai M, Kobayashi R, Masai H, Kimura K, Maeda K, Nagashima H
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
55 |
nm3 |
|
|
UniProt ID: Q99QS1 (266-476) Protein map
|
|
|
|
| Sample: |
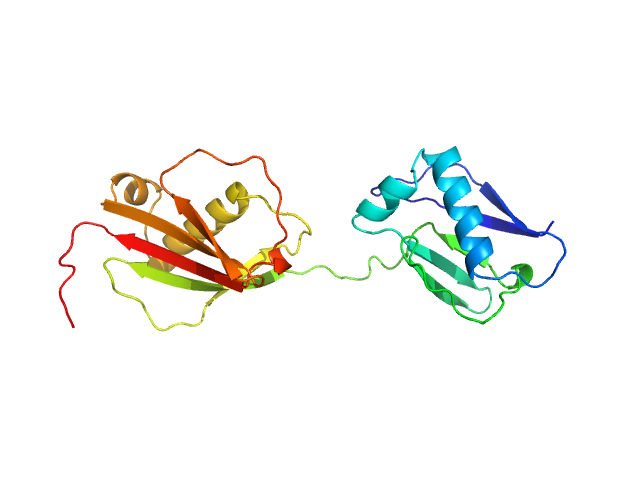
Protein map monomer, 24 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Nov 17
|
S. aureus Eap is a polyvalent inhibitor of neutrophil serine proteases.
J Biol Chem 300(9):107627 (2024)
Mishra N, Gido CD, Herdendorf TJ, Hammel M, Hura GL, Fu ZQ, Geisbrecht BV
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
GlcNAc-binding protein A monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
UniProt ID: P20917 (20-325) Myelin-associated glycoprotein Ig domains 1-3
|
|
|
|
| Sample: |
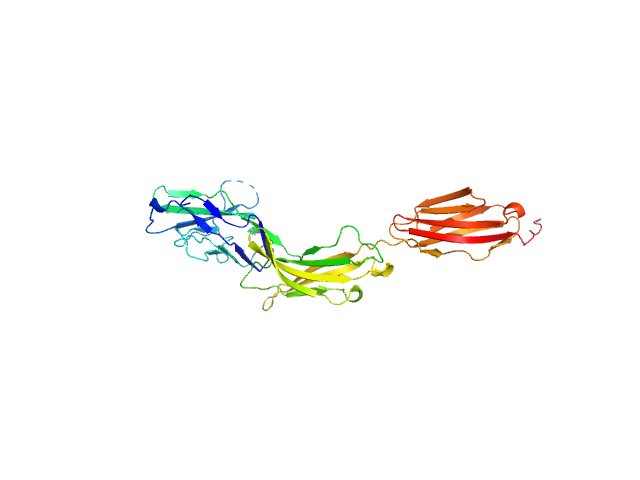
Myelin-associated glycoprotein Ig domains 1-3 monomer, 35 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 11
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
49 |
nm3 |
|
|
UniProt ID: Q46085 (718-1021) Collagenase ColH segement s2as2bs3
|
|
|

|
| Sample: |
Collagenase ColH segement s2as2bs3 monomer, 34 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
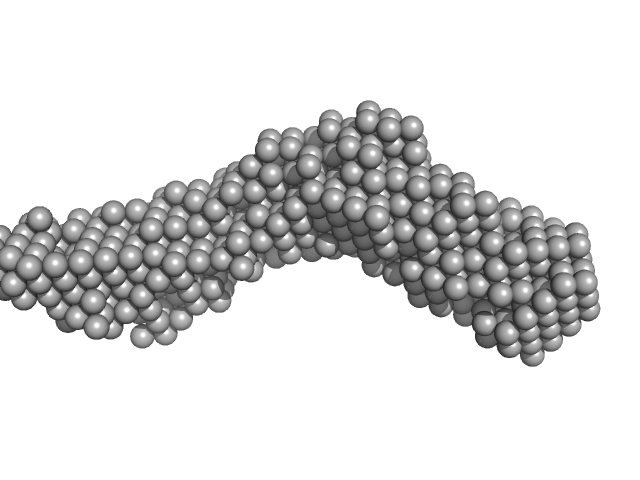
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
3.3 |
nm |
| Dmax |
14.2 |
nm |
| VolumePorod |
37 |
nm3 |
|
|
UniProt ID: O95256 (20-364) Interleukin-18 receptor accessory protein ectodomain with Rα linker
|
|
|
|
| Sample: |
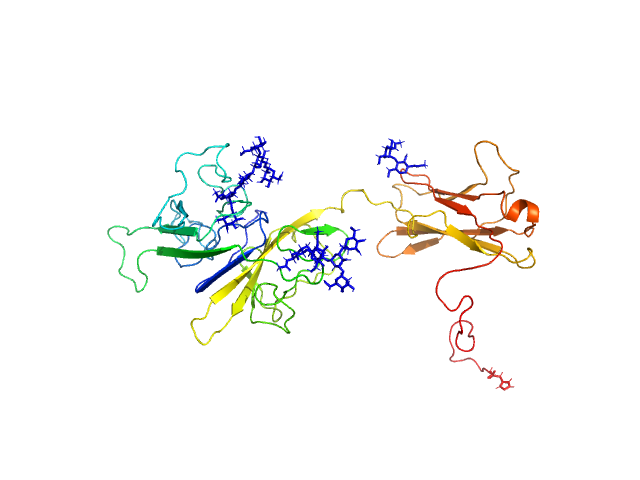
Interleukin-18 receptor accessory protein ectodomain with Rα linker monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 3% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2017 Jul 24
|
Functional Relevance of Interleukin-1 Receptor Inter-domain Flexibility for Cytokine Binding and Signaling.
Structure 27(8):1296-1307.e5 (2019)
Ge J, Remesh SG, Hammel M, Pan S, Mahan AD, Wang S, Wang X
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha
|
|
|
|
| Sample: |
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, 10 kDa Escherichia coli protein
|
| Buffer: |
10 mM sodium acetate, 100 mM NaCl, pH: 4.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jun 1
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
|
|