UniProt ID: P9WMU1 (None-None) Cell wall synthesis protein Wag31
|
|
|
|
| Sample: |
Cell wall synthesis protein Wag31, 30 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50mM Tris pH7.5, 300mM NaCl, 10% Glycerol, 1mM EDTA (ethylene diamine tetra acetic acid), 5mM β-mercaptoethanol (BME), pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2018 Jan 29
|
Higher order assembling of the mycobacterial polar growth factor DivIVA/Wag31.
J Struct Biol :107429 (2019)
Choukate K, Gupta A, Basu B, Virk K, Ganguli M, Chaudhuri B
|
|
|
UniProt ID: None (53-551) Alpha domain of Ag43a
UniProt ID: None (None-None) Fragment antigen-binding region Fab10C12
|
|
|

|
| Sample: |
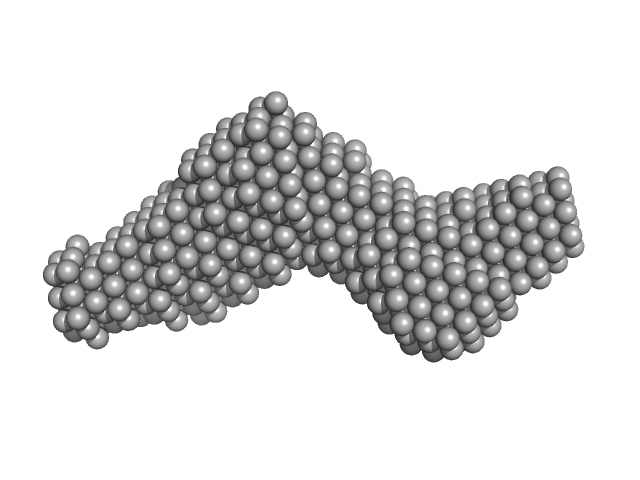
Alpha domain of Ag43a monomer, 49 kDa Escherichia coli protein
Fragment antigen-binding region Fab10C12 monomer, 47 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 Nov 3
|
Inhibition of aggregation and biofilm formation by Uropathogenic Escherichia coli
Andrew Whitten
|
| RgGuinier |
4.3 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
103 |
nm3 |
|
|
UniProt ID: Q06187 (391-659) Bruton's tyrosine kinase, kinase domain
|
|
|

|
| Sample: |
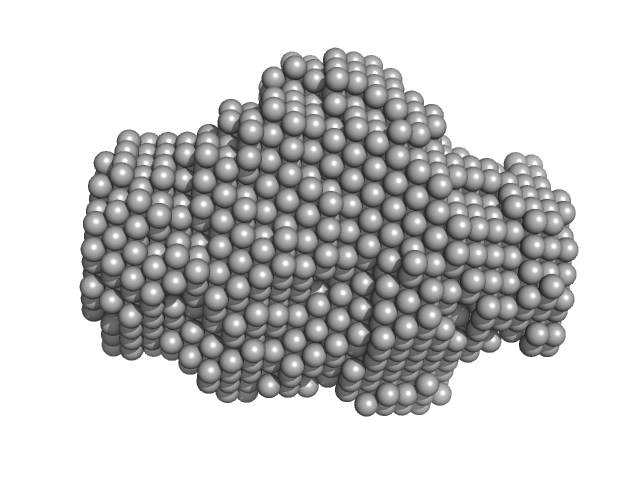
Bruton's tyrosine kinase, kinase domain monomer, 32 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 11
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.8 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
UniProt ID: Q06187 (214-659) Bruton's tyrosine kinase - Src homology 3-2 kinase domain
|
|
|
|
| Sample: |
Bruton's tyrosine kinase - Src homology 3-2 kinase domain monomer, 52 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Nov 2
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
UniProt ID: Q06187 (1-659) Bruton's tyrosine kinase - full length
|
|
|
|
| Sample: |
Bruton's tyrosine kinase - full length monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Nov 3
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
4.0 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
114 |
nm3 |
|
|
UniProt ID: A0A0U3FQH7 (None-None) Malate dehydrogenase
|
|
|
|
| Sample: |
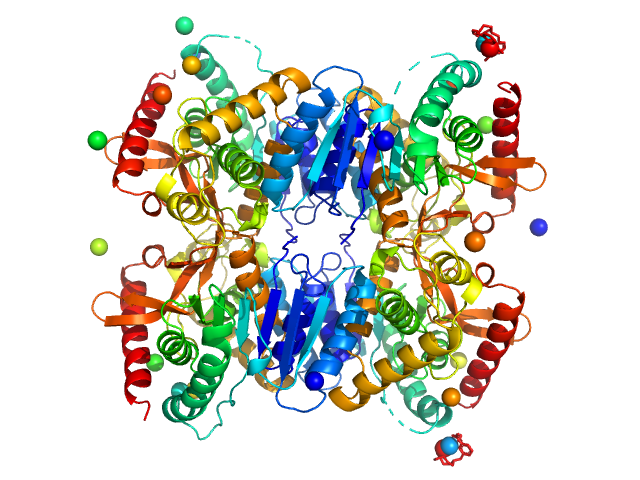
Malate dehydrogenase tetramer, 134 kDa Ignicoccus islandicus DSM … protein
|
| Buffer: |
50 mM Tris-HCl 50 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Sep 5
|
The archaeal LDH-like malate dehydrogenase from Ignicoccus islandicus displays dual substrate recognition, hidden allostery and a non-canonical tetrameric oligomeric organization
Journal of Structural Biology (2019)
Roche J, Girard E, Mas C, Madern D
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
198 |
nm3 |
|
|
UniProt ID: P53396-1 (None-None) ATP-citrate synthase
|
|
|
|
| Sample: |
ATP-citrate synthase tetramer, 458 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 4
|
Structure of ATP citrate lyase and the origin of citrate synthase in the Krebs cycle.
Nature 568(7753):571-575 (2019)
Verschueren KHG, Blanchet C, Felix J, Dansercoer A, De Vos D, Bloch Y, Van Beeumen J, Svergun D, Gutsche I, Savvides SN, Verstraete K
|
| RgGuinier |
6.1 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
765 |
nm3 |
|
|
UniProt ID: P53396-1 (None-None) ATP-citrate synthase
|
|
|
|
| Sample: |
ATP-citrate synthase tetramer, 458 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 50mM Tris, 20mM citrate, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 4
|
Structure of ATP citrate lyase and the origin of citrate synthase in the Krebs cycle.
Nature 568(7753):571-575 (2019)
Verschueren KHG, Blanchet C, Felix J, Dansercoer A, De Vos D, Bloch Y, Van Beeumen J, Svergun D, Gutsche I, Savvides SN, Verstraete K
|
| RgGuinier |
6.2 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
787 |
nm3 |
|
|
UniProt ID: P53396-1 (None-None) ATP-citrate synthase
|
|
|
|
| Sample: |
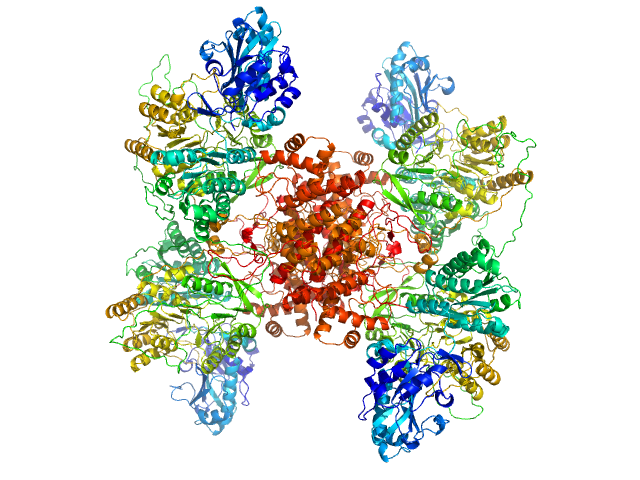
ATP-citrate synthase tetramer, 458 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 50mM Tris, 20mM citrate, 2mM CoA, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 4
|
Structure of ATP citrate lyase and the origin of citrate synthase in the Krebs cycle.
Nature 568(7753):571-575 (2019)
Verschueren KHG, Blanchet C, Felix J, Dansercoer A, De Vos D, Bloch Y, Van Beeumen J, Svergun D, Gutsche I, Savvides SN, Verstraete K
|
| RgGuinier |
5.9 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
775 |
nm3 |
|
|
UniProt ID: P21333 (574-869) Filamin A Ig-like domains 4-6
|
|
|

|
| Sample: |
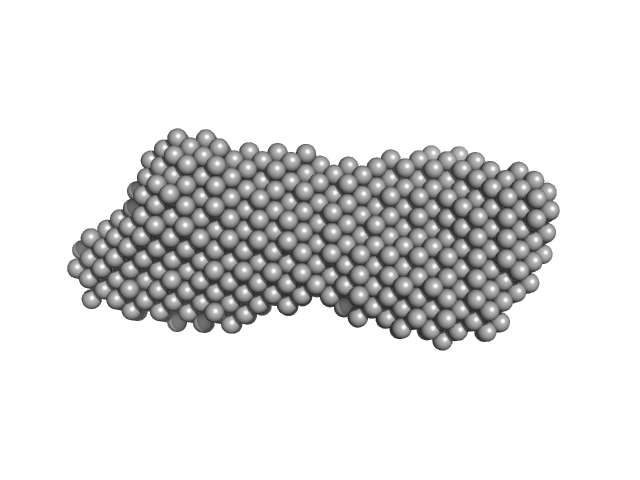
Filamin A Ig-like domains 4-6 monomer, 32 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Critical Structural Defects Explain Filamin A Mutations Causing Mitral Valve Dysplasia.
Biophys J 117(8):1467-1475 (2019)
Haataja TJK, Capoulade R, Lecointe S, Hellman M, Merot J, Permi P, Pentikäinen U
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
41 |
nm3 |
|
|