UniProt ID: P21333 (574-869) Filamin A Ig-like domains 4-6, V711D mutation
|
|
|
|
| Sample: |
Filamin A Ig-like domains 4-6, V711D mutation monomer, 32 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Critical Structural Defects Explain Filamin A Mutations Causing Mitral Valve Dysplasia.
Biophys J 117(8):1467-1475 (2019)
Haataja TJK, Capoulade R, Lecointe S, Hellman M, Merot J, Permi P, Pentikäinen U
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
64 |
nm3 |
|
|
UniProt ID: P21333 (574-869) Filamin A Ig-like domains 4-6, V711D mutation
|
|
|
|
| Sample: |
Filamin A Ig-like domains 4-6, V711D mutation monomer, 32 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Critical Structural Defects Explain Filamin A Mutations Causing Mitral Valve Dysplasia.
Biophys J 117(8):1467-1475 (2019)
Haataja TJK, Capoulade R, Lecointe S, Hellman M, Merot J, Permi P, Pentikäinen U
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
UniProt ID: P21333 (574-869) Filamin A Ig-like domains 4-6, H743P mutation
|
|
|
|
| Sample: |
Filamin A Ig-like domains 4-6, H743P mutation monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Critical Structural Defects Explain Filamin A Mutations Causing Mitral Valve Dysplasia.
Biophys J 117(8):1467-1475 (2019)
Haataja TJK, Capoulade R, Lecointe S, Hellman M, Merot J, Permi P, Pentikäinen U
|
| RgGuinier |
4.3 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
UniProt ID: P21333 (574-869) Filamin A Ig-like domains 4-6, H743P mutation
|
|
|
|
| Sample: |
Filamin A Ig-like domains 4-6, H743P mutation monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 100 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Feb 10
|
Critical Structural Defects Explain Filamin A Mutations Causing Mitral Valve Dysplasia.
Biophys J 117(8):1467-1475 (2019)
Haataja TJK, Capoulade R, Lecointe S, Hellman M, Merot J, Permi P, Pentikäinen U
|
| RgGuinier |
4.0 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
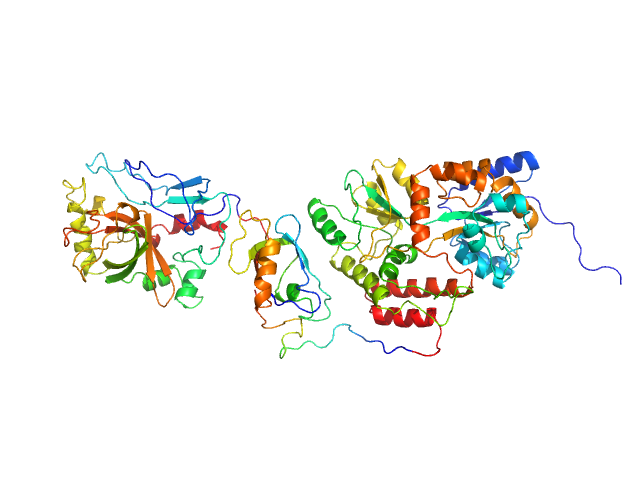
UniProt ID: P17896 (None-None) SpoIVB peptidase (MBP fusion)
|
|
|
|
| Sample: |
SpoIVB peptidase (MBP fusion) monomer, 80 kDa Bacillus subtilis protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Jul 12
|
Solution Structure of SpoIVB Reveals Mechanism of PDZ Domain-Regulated Protease Activity.
Front Microbiol 10:1232 (2019)
Xie X, Guo N, Xue G, Xie D, Yuan C, Harrison J, Li J, Jiang L, Huang M
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
96 |
nm3 |
|
|
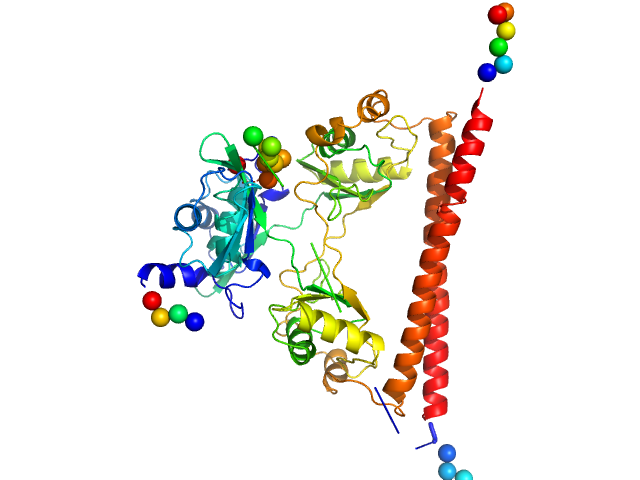
UniProt ID: P23246 (276-535) Splicing factor, proline- and glutamine-rich
|
|
|
|
| Sample: |
Splicing factor, proline- and glutamine-rich dimer, 60 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl, 250 mM NaCl, 5% (v/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 19
|
A new crystal structure and small-angle X-ray scattering analysis of the homodimer of human SFPQ.
Acta Crystallogr F Struct Biol Commun 75(Pt 6):439-449 (2019)
Hewage TW, Caria S, Lee M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
91 |
nm3 |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
| Sample: |
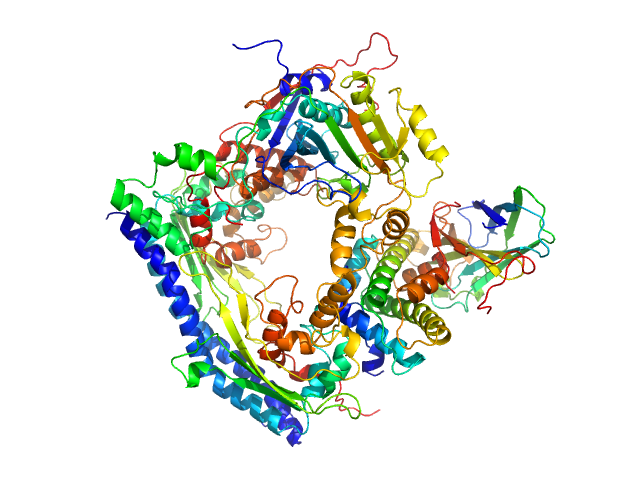
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Mar 3
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.8 |
nm |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
| Sample: |
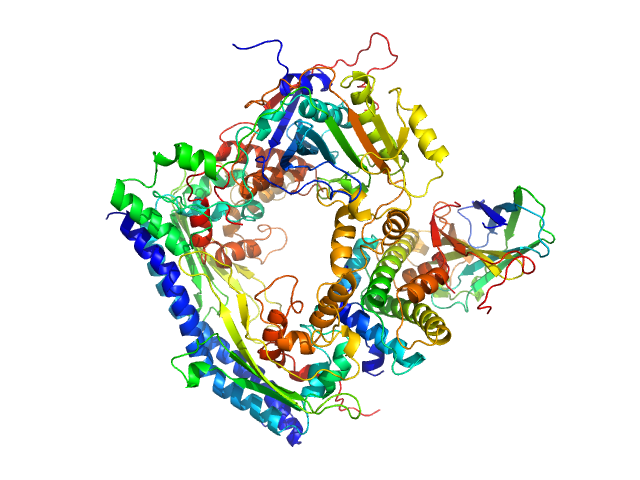
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Mar 4
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
|
|
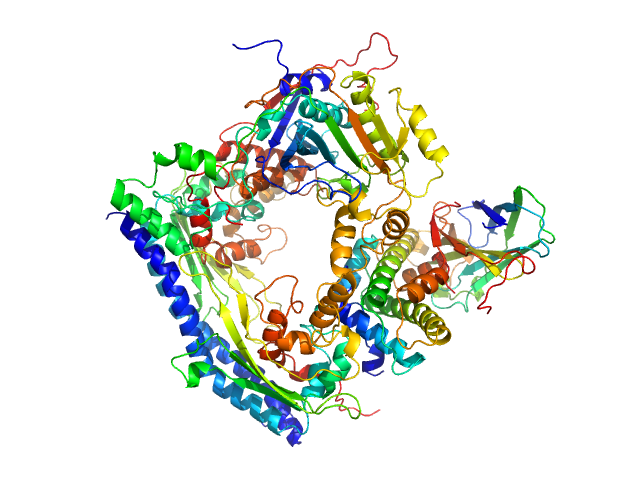
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Mar 4
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.5 |
nm |
|
|
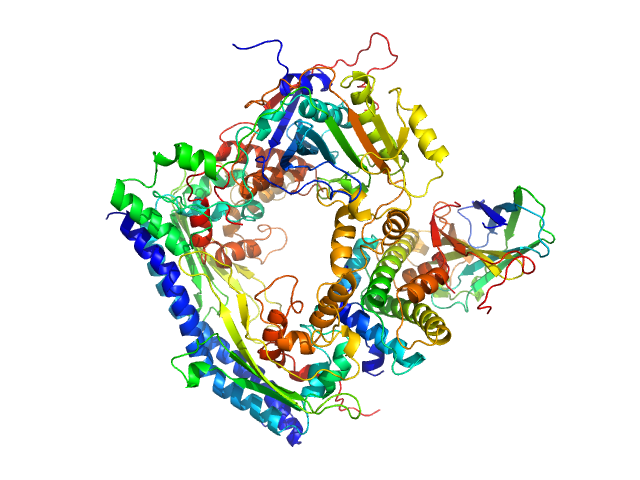
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2018 May 29
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.5 |
nm |
|
|