UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P62799 (1-103) Histone H4
UniProt ID: Q6PI79 (1-136) Histone H3 full-length
|
|
|
|
| Sample: |
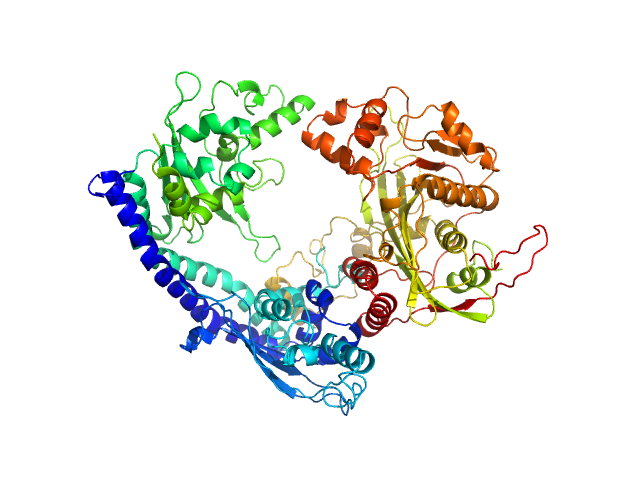
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
Histone H3 full-length monomer, 15 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 42% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Nov 14
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.5 |
nm |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P62799 (1-103) Histone H4
UniProt ID: Q6PI79 (1-136) Histone H3 full-length
|
|
|
|
| Sample: |
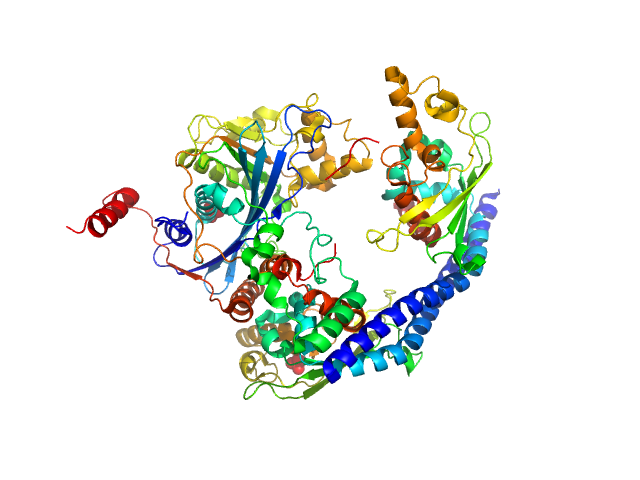
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
Histone H3 full-length monomer, 15 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 42% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Nov 14
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.0 |
nm |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P62799 (1-103) Histone H4
UniProt ID: Q6PI79 (1-136) Histone H3 full-length
|
|
|
|
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
Histone H3 full-length monomer, 15 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2018 May 30
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.0 |
nm |
|
|
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P62799 (1-103) Histone H4
UniProt ID: Q6PI79 (1-136) Histone H3 full-length
UniProt ID: P53853 (1-264) Vacuolar protein sorting-associated protein 75 full-length
|
|
|
|
| Sample: |
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
Histone H3 full-length monomer, 15 kDa Xenopus laevis protein
Vacuolar protein sorting-associated protein 75 full-length dimer, 61 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Jun 9
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.5 |
nm |
|
|
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P62799 (1-103) Histone H4
UniProt ID: Q6PI79 (1-136) Histone H3 full-length
UniProt ID: P53853 (1-264) Vacuolar protein sorting-associated protein 75 full-length
|
|
|
|
| Sample: |
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
Histone H3 full-length monomer, 15 kDa Xenopus laevis protein
Vacuolar protein sorting-associated protein 75 full-length dimer, 61 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 42% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Jun 9
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.5 |
nm |
|
|
UniProt ID: P00883 (2-364) Fructose-bisphosphate aldolase A
|
|
|
|
| Sample: |
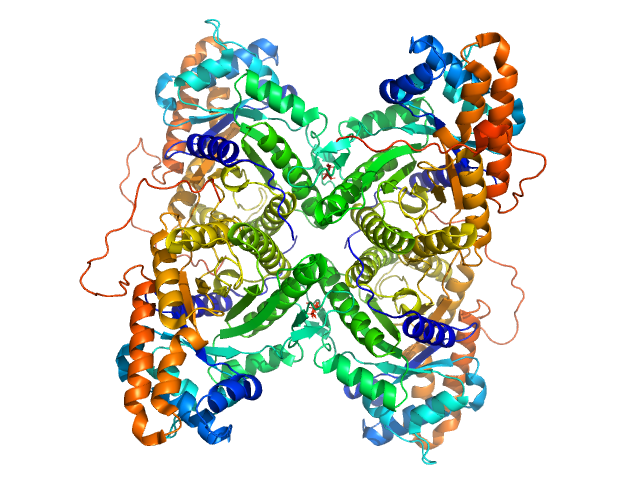
Fructose-bisphosphate aldolase A tetramer, 157 kDa Oryctolagus cuniculus protein
|
| Buffer: |
20 mM HEPES, pH: 7 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, Département de Biochimie, Université de Montréal on 2019 Jun 14
|
Rabbit muscle fructose-1,6-bisphosphate aldolase K229M mutant
Normand Cyr
|
| RgGuinier |
3.6 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
213 |
nm3 |
|
|
UniProt ID: Q9Y490 (1-405) Talin-1 (Δ139-168), human
|
|
|
|
| Sample: |
Talin-1 (Δ139-168), human monomer, 48 kDa Homo sapiens protein
|
| Buffer: |
50 mM sodium phosphate, 150mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Sep 24
|
The F1 loop of the talin head domain acts as a gatekeeper in integrin activation and clustering.
J Cell Sci 133(19) (2020)
Kukkurainen S, Azizi L, Zhang P, Jacquier MC, Baikoghli M, von Essen M, Tuukkanen A, Laitaoja M, Liu X, Rahikainen R, Orłowski A, Jänis J, Määttä JAE, Varjosalo M, Vattulainen I, Róg T, Svergun D, Cheng RH, Wu J, Hytönen VP, Wehrle-Haller B
|
| RgGuinier |
3.3 |
nm |
| Dmax |
13.3 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: Q9Y490 (1-405) Talin-1, human
|
|
|
|
| Sample: |
Talin-1, human monomer, 51 kDa Homo sapiens protein
|
| Buffer: |
50 mM sodium phosphate, 150 mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Sep 24
|
The F1 loop of the talin head domain acts as a gatekeeper in integrin activation and clustering.
J Cell Sci 133(19) (2020)
Kukkurainen S, Azizi L, Zhang P, Jacquier MC, Baikoghli M, von Essen M, Tuukkanen A, Laitaoja M, Liu X, Rahikainen R, Orłowski A, Jänis J, Määttä JAE, Varjosalo M, Vattulainen I, Róg T, Svergun D, Cheng RH, Wu J, Hytönen VP, Wehrle-Haller B
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
94 |
nm3 |
|
|
UniProt ID: Q9Y490 (1-405) Talin-1 (Δ134-170/GAG insert), human
|
|
|
|
| Sample: |
Talin-1 (Δ134-170/GAG insert), human monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
50 mM sodium phosphate, 150mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Sep 24
|
The F1 loop of the talin head domain acts as a gatekeeper in integrin activation and clustering.
J Cell Sci 133(19) (2020)
Kukkurainen S, Azizi L, Zhang P, Jacquier MC, Baikoghli M, von Essen M, Tuukkanen A, Laitaoja M, Liu X, Rahikainen R, Orłowski A, Jänis J, Määttä JAE, Varjosalo M, Vattulainen I, Róg T, Svergun D, Cheng RH, Wu J, Hytönen VP, Wehrle-Haller B
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
71 |
nm3 |
|
|
UniProt ID: Q9UBF9 (2-498) his-trx-myotilin
|
|
|
|
| Sample: |
His-trx-myotilin monomer, 70 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 400 mM NaCl, 250 mM arginine, 5% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jun 11
|
Molecular basis of F-actin regulation and sarcomere assembly via myotilin
PLOS Biology 19(4):e3001148 (2021)
Kostan J, Pavšič M, Puž V, Schwarz T, Drepper F, Molt S, Graewert M, Schreiner C, Sajko S, van der Ven P, Onipe A, Svergun D, Warscheid B, Konrat R, Fürst D, Lenarčič B, Djinović-Carugo K, Machesky L
|
| RgGuinier |
5.1 |
nm |
| Dmax |
16.7 |
nm |
| VolumePorod |
131 |
nm3 |
|
|