UniProt ID: B1HQ59 (None-None) Lysinibacillus Mosquito-larvicidal receptor binding component
|
|
|
|
| Sample: |
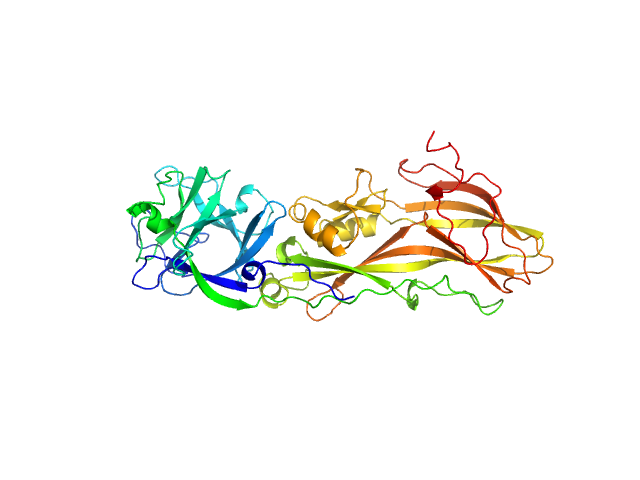
Lysinibacillus Mosquito-larvicidal receptor binding component monomer, 53 kDa synthetic construct protein
|
| Buffer: |
25 mM HEPES, pH 7.5, 25 mM NaCl, in 100% D2O, pH: 7.5 |
| Experiment: |
SANS
data collected at SANS-I facility, Dhruva Reactor, Bhabha Atomic Research Centre on 2019 Apr 24
|
Small-angle neutron scattering studies suggest the mechanism of BinAB protein internalization
IUCrJ 7(2) (2020)
Sharma M, Aswal V, Kumar V, Chidambaram R
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.9 |
nm |
|
|
UniProt ID: P00439 (1-452) Phenylalanine-4-hydroxylase
|
|
|
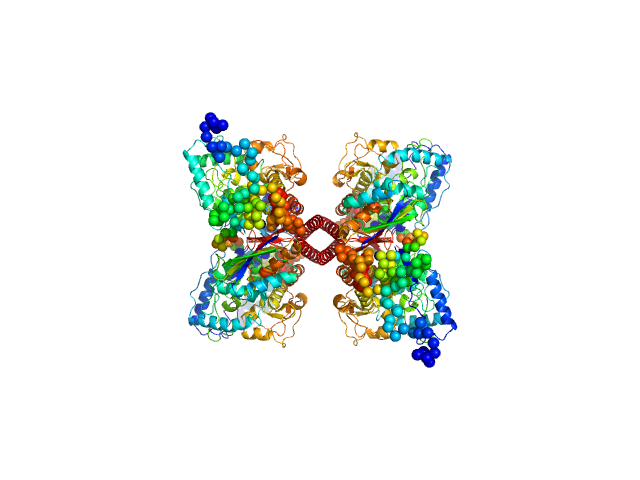
|
| Sample: |
Phenylalanine-4-hydroxylase tetramer, 223 kDa Homo sapiens protein
|
| Buffer: |
0.02 mM Hepes, 0.2 M NaCl,, pH: 7 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Feb 25
|
Modulation of Human Phenylalanine Hydroxylase by 3-Hydroxyquinolin-2(1H)-One Derivatives
Biomolecules 11(3):462 (2021)
Lopes R, Tomé C, Russo R, Paterna R, Leandro J, Candeias N, Gonçalves L, Teixeira M, Sousa P, Guedes R, Vicente J, Gois P, Leandro P
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
336 |
nm3 |
|
|
UniProt ID: P00439 (1-452) Phenylalanine-4-hydroxylase
|
|
|

|
| Sample: |
Phenylalanine-4-hydroxylase tetramer, 223 kDa Homo sapiens protein
|
| Buffer: |
0.02 mM Hepes, 0.2 M NaCl,, pH: 7 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Feb 25
|
Modulation of Human Phenylalanine Hydroxylase by 3-Hydroxyquinolin-2(1H)-One Derivatives
Biomolecules 11(3):462 (2021)
Lopes R, Tomé C, Russo R, Paterna R, Leandro J, Candeias N, Gonçalves L, Teixeira M, Sousa P, Guedes R, Vicente J, Gois P, Leandro P
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.3 |
nm |
| VolumePorod |
367 |
nm3 |
|
|
UniProt ID: H9N289 (None-None) Methylxanthine N1-demethylase NdmA
UniProt ID: H9N290 (None-None) Methylxanthine N3-demethylase NdmB
|
|
|
|
| Sample: |
Methylxanthine N1-demethylase NdmA trimer, 127 kDa Pseudomonas putida protein
Methylxanthine N3-demethylase NdmB trimer, 129 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP 10% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2018 Jul 27
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
4.5 |
nm |
| Dmax |
12.3 |
nm |
|
|
UniProt ID: H9N289 (None-None) Methylxanthine N1-demethylase NdmA
UniProt ID: H9N290 (None-None) Methylxanthine N3-demethylase NdmB
|
|
|

|
| Sample: |
Methylxanthine N1-demethylase NdmA trimer, 207 kDa Pseudomonas putida protein
Methylxanthine N3-demethylase NdmB trimer, 129 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP 10% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2018 Jul 27
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
5.4 |
nm |
| Dmax |
13.8 |
nm |
|
|
UniProt ID: H9N289 (None-None) Methylxanthine N1-demethylase NdmA
|
|
|

|
| Sample: |
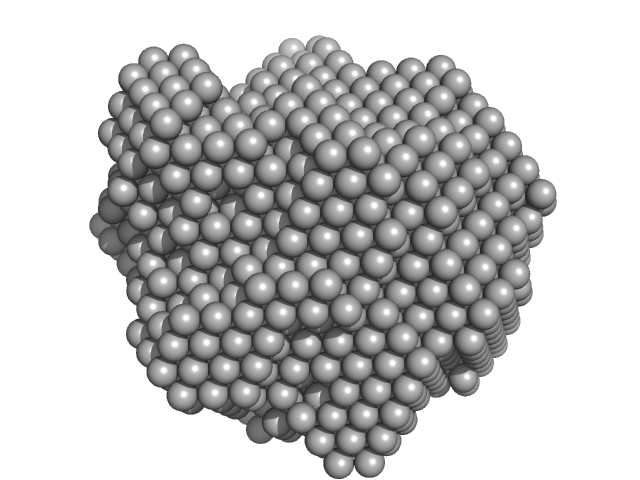
Methylxanthine N1-demethylase NdmA hexamer, 254 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2018 May 21
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
4.2 |
nm |
| Dmax |
11.0 |
nm |
|
|
UniProt ID: H9N290 (None-None) Methylxanthine N3-demethylase NdmB
|
|
|

|
| Sample: |
Methylxanthine N3-demethylase NdmB hexamer, 258 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2018 May 21
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
4.3 |
nm |
| Dmax |
12.2 |
nm |
|
|
UniProt ID: H9N289 (None-None) Methylxanthine N1-demethylase NdmA
UniProt ID: H9N290 (None-None) Methylxanthine N3-demethylase NdmB
|
|
|

|
| Sample: |
Methylxanthine N1-demethylase NdmA trimer, 127 kDa Pseudomonas putida protein
Methylxanthine N3-demethylase NdmB trimer, 129 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2018 May 21
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
4.2 |
nm |
| Dmax |
10.9 |
nm |
|
|
UniProt ID: H9N289 (None-None) Methylxanthine N1-demethylase NdmA
UniProt ID: H9N290 (None-None) Methylxanthine N3-demethylase NdmB
|
|
|
|
| Sample: |
Methylxanthine N1-demethylase NdmA trimer, 207 kDa Pseudomonas putida protein
Methylxanthine N3-demethylase NdmB trimer, 208 kDa Pseudomonas putida protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl 2 mM TCEP 10% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 4C, Pohang Accelerator Laboratory on 2018 Jul 27
|
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J Mol Biol 431(19):3647-3661 (2019)
Kim JH, Kim BH, Brooks S, Kang SY, Summers RM, Song HK
|
| RgGuinier |
5.6 |
nm |
| Dmax |
19.1 |
nm |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 100% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Nov 14
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity (Asf1-H3:H4-Rtt109-Vps75 protein complex, data for docking block selections)
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.5 |
nm |
|
|