|
|
|
|
|
| Sample: |
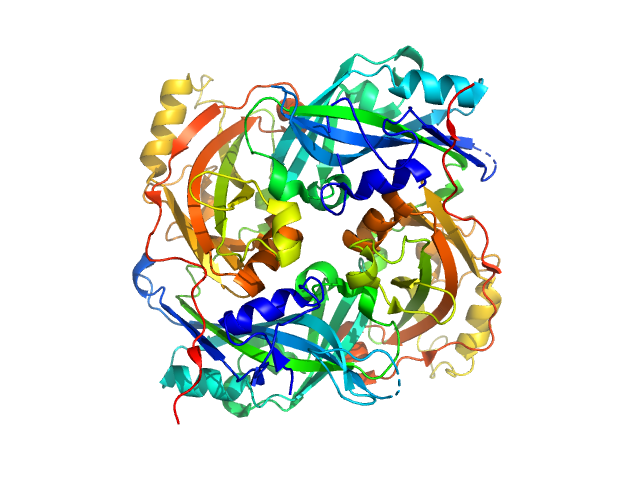
Matrix protein dimer, 79 kDa Newcastle disease virus … protein
|
| Buffer: |
STE buffer 100 mM NaCl, 10 mM Tris-HCl, and 1 mM EDTA, pH: 4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Dec 11
|
Solution Structure, Self-Assembly, and Membrane Interactions of the Matrix Protein from Newcastle Disease Virus at Neutral and Acidic pH
Journal of Virology 93(6) (2019)
Shtykova E, Petoukhov M, Dadinova L, Fedorova N, Tashkin V, Timofeeva T, Ksenofontov A, Loshkarev N, Baratova L, Jeffries C, Svergun D, Batishchev O, García-Sastre A
|
|
|
|
|
|
|

|
| Sample: |
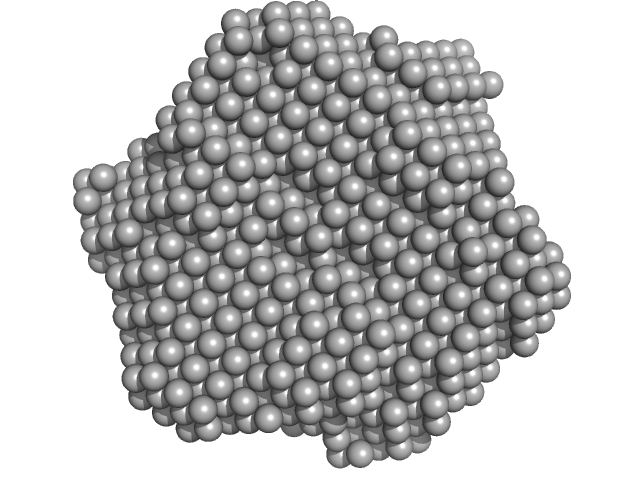
DsbA-like disulfide oxidoreductase (thiol-disulfide exchange protein) trimer, 81 kDa Wolbachia endosymbiont of … protein
|
| Buffer: |
25 mM TRIS, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2014 Mar 29
|
The atypical thiol-disulfide exchange protein α-DsbA2 from Wolbachia pipientis is a homotrimeric disulfide isomerase.
Acta Crystallogr D Struct Biol 75(Pt 3):283-295 (2019)
Walden PM, Whitten AE, Premkumar L, Halili MA, Heras B, King GJ, Martin JL
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.7 |
nm |
| VolumePorod |
913 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
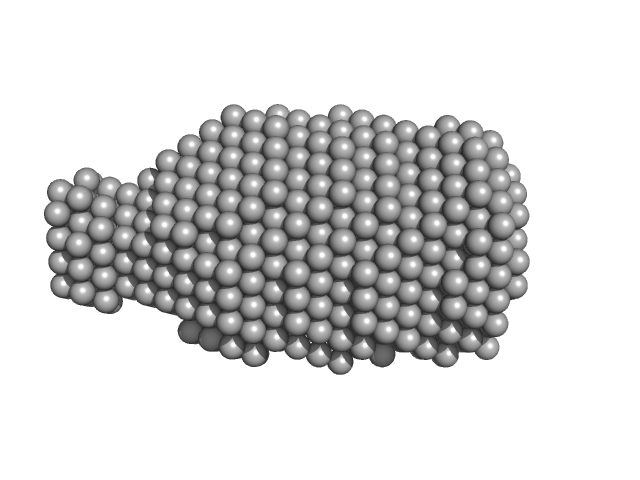
DsbA-like disulfide oxidoreductase (thiol-disulfide exchange protein) monomer, 21 kDa Wolbachia endosymbiont of … protein
|
| Buffer: |
25 mM TRIS, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2012 Feb 29
|
The atypical thiol-disulfide exchange protein α-DsbA2 from Wolbachia pipientis is a homotrimeric disulfide isomerase.
Acta Crystallogr D Struct Biol 75(Pt 3):283-295 (2019)
Walden PM, Whitten AE, Premkumar L, Halili MA, Heras B, King GJ, Martin JL
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.3 |
nm |
| VolumePorod |
275 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
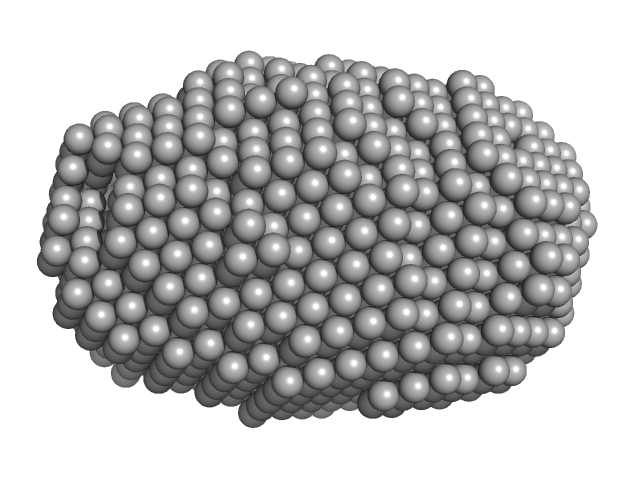
C-terminal catalytic domain of Suppressor of Copper Sensitivity C protein monomer, 20 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Acta Crystallogr D Struct Biol 75(Pt 3):296-307 (2019)
Furlong EJ, Kurth F, Premkumar L, Whitten AE, Martin JL
|
| RgGuinier |
1.7 |
nm |
| Dmax |
5.4 |
nm |
| VolumePorod |
22200 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
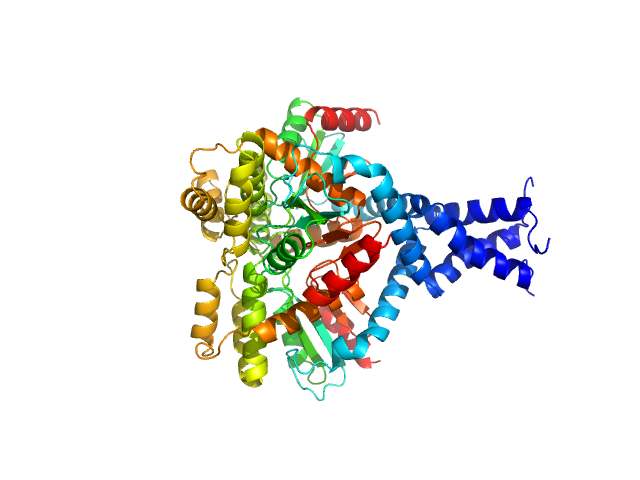
Deletion mutant of PmScsC, 23 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Acta Crystallogr D Struct Biol 75(Pt 3):296-307 (2019)
Furlong EJ, Kurth F, Premkumar L, Whitten AE, Martin JL
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
54 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
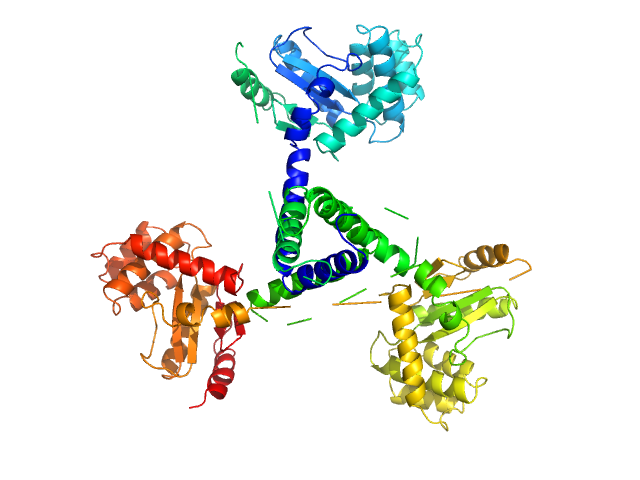
Suppressor of Copper Sensitivity C protein trimer, 74 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Acta Crystallogr D Struct Biol 75(Pt 3):296-307 (2019)
Furlong EJ, Kurth F, Premkumar L, Whitten AE, Martin JL
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
|
|
|
|
|
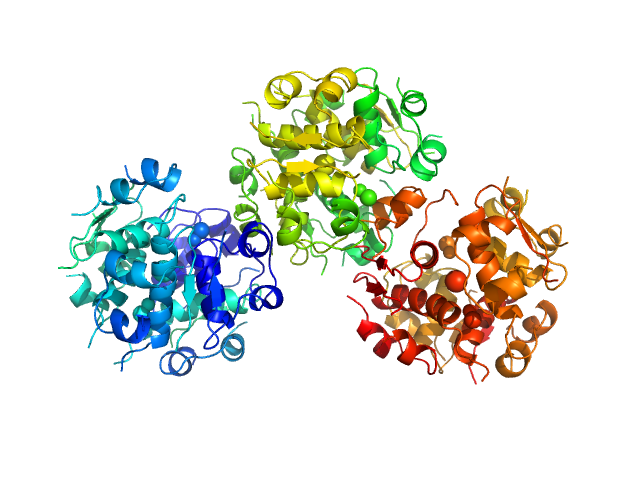
| Sample: |
Insulin detemir 18-mer, 106 kDa protein
|
| Buffer: |
5.0 mM Na2HPO4, 13.1 mM m-cresol, 15.1 mM phenol, 173.7 mM glycerol, 20.0 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.3 |
nm |
| VolumePorod |
137 |
nm3 |
|
|
|
|
|
|
|
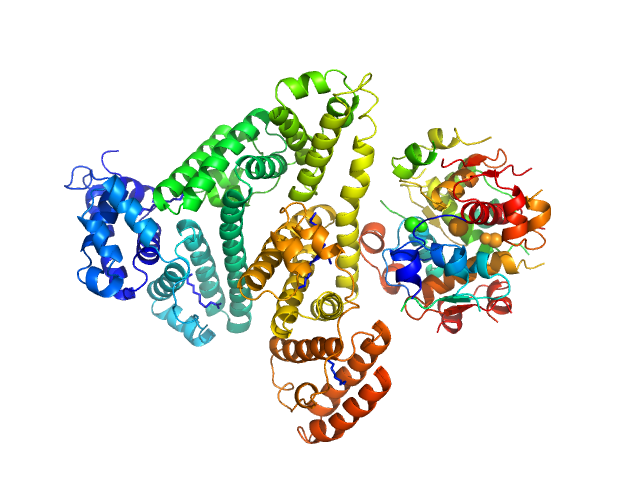
| Sample: |
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
|
|
|
|
|
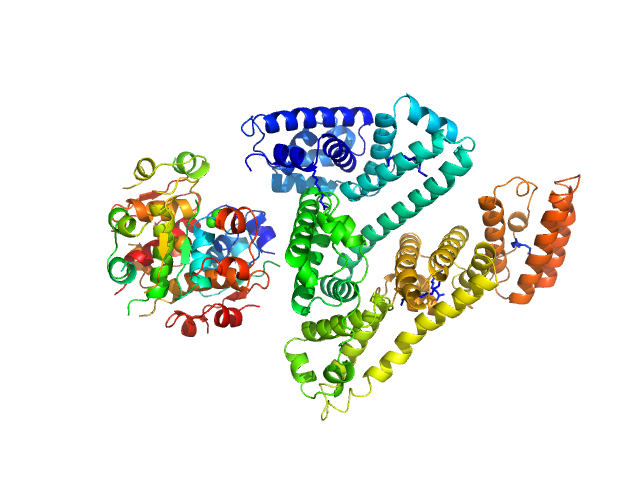
| Sample: |
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
|
|
|
|
|
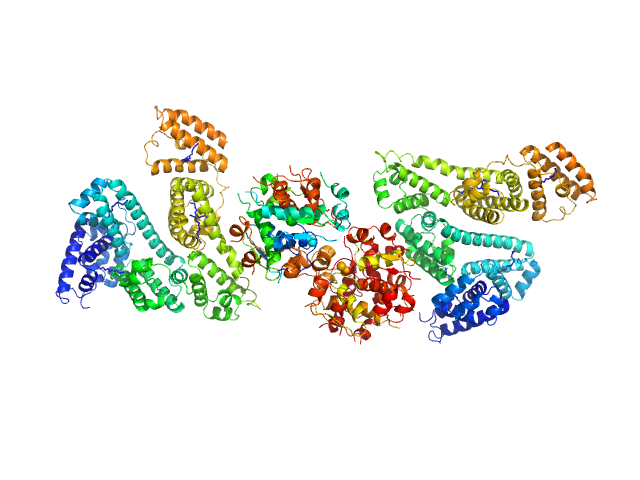
| Sample: |
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
Insulin detemir (Levemir(R), Novo Nordisk A/S) dodecamer, 71 kDa protein
|
| Buffer: |
8.8 mM Na2HPO4, 10.6 mM m-cresol, 12.2 mM phenol, 140.9 mM glycerol, 56.9 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
309 |
nm3 |
|
|