|
|
|
|

|
| Sample: |
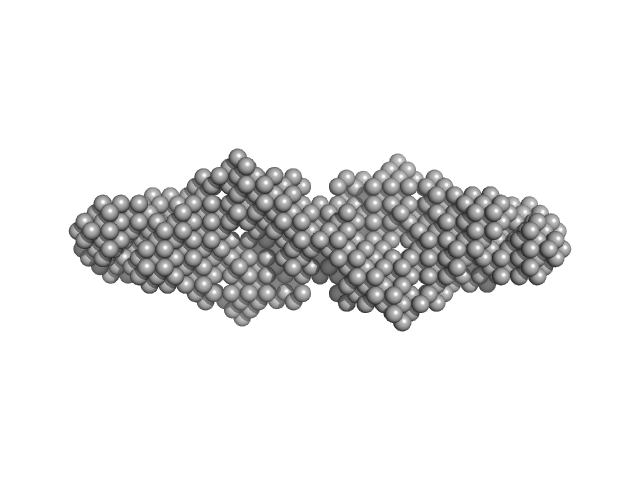
Hypothetical protein CTHT_0072540 tetramer, 62 kDa Chaetomium thermophilum protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 2 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Jul 30
|
Prp19/Pso4 Is an Autoinhibited Ubiquitin Ligase Activated by Stepwise Assembly of Three Splicing Factors.
Mol Cell 69(6):979-992.e6 (2018)
de Moura TR, Mozaffari-Jovin S, Szabó CZK, Schmitzová J, Dybkov O, Cretu C, Kachala M, Svergun D, Urlaub H, Lührmann R, Pena V
|
| RgGuinier |
4.1 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
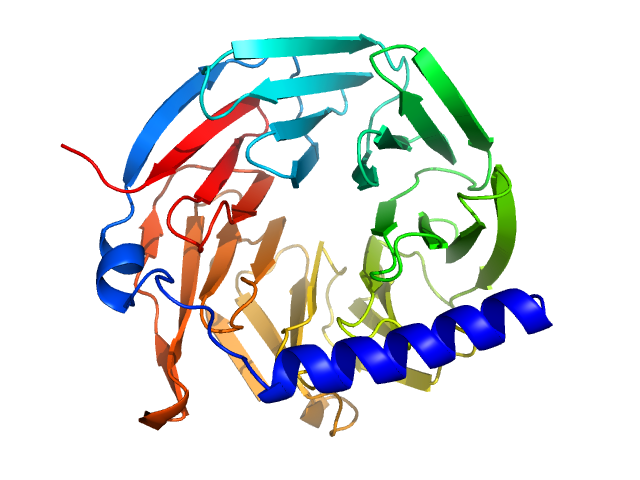
Hypothetical protein CTHT_0072540 monomer, 35 kDa Chaetomium thermophilum protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 2 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Jul 19
|
Prp19/Pso4 Is an Autoinhibited Ubiquitin Ligase Activated by Stepwise Assembly of Three Splicing Factors.
Mol Cell 69(6):979-992.e6 (2018)
de Moura TR, Mozaffari-Jovin S, Szabó CZK, Schmitzová J, Dybkov O, Cretu C, Kachala M, Svergun D, Urlaub H, Lührmann R, Pena V
|
| RgGuinier |
2.3 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
68 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
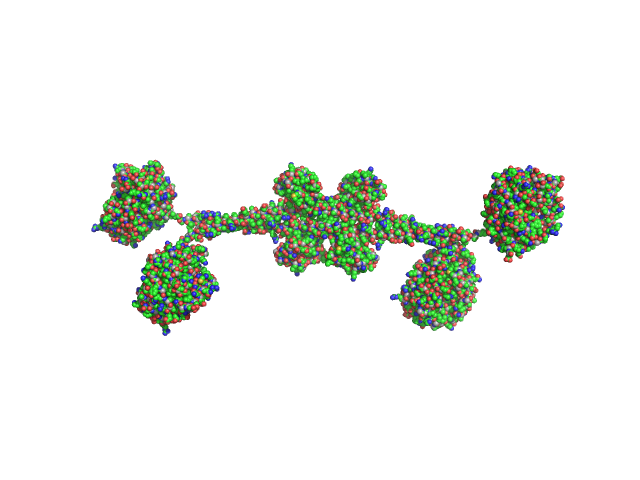
Full-length hypothetical protein CTHT_0072540 tetramer, 207 kDa Chaetomium thermophilum protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 2 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2013 Oct 15
|
Prp19/Pso4 Is an Autoinhibited Ubiquitin Ligase Activated by Stepwise Assembly of Three Splicing Factors.
Mol Cell 69(6):979-992.e6 (2018)
de Moura TR, Mozaffari-Jovin S, Szabó CZK, Schmitzová J, Dybkov O, Cretu C, Kachala M, Svergun D, Urlaub H, Lührmann R, Pena V
|
| RgGuinier |
6.2 |
nm |
| Dmax |
23.0 |
nm |
| VolumePorod |
280 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
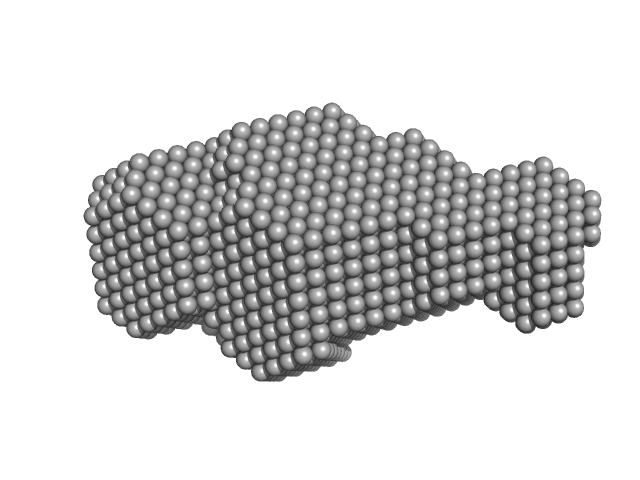
Deoxyuridine 5'-triphosphate nucleotidohydrolase trimer, 54 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES 300 mM NaCl 5 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 9
|
Structural model of human dUTPase in complex with a novel proteinaceous inhibitor.
Sci Rep 8(1):4326 (2018)
Nyíri K, Mertens HDT, Tihanyi B, Nagy GN, Kőhegyi B, Matejka J, Harris MJ, Szabó JE, Papp-Kádár V, Németh-Pongrácz V, Ozohanics O, Vékey K, Svergun DI, Borysik AJ, Vértessy BG
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
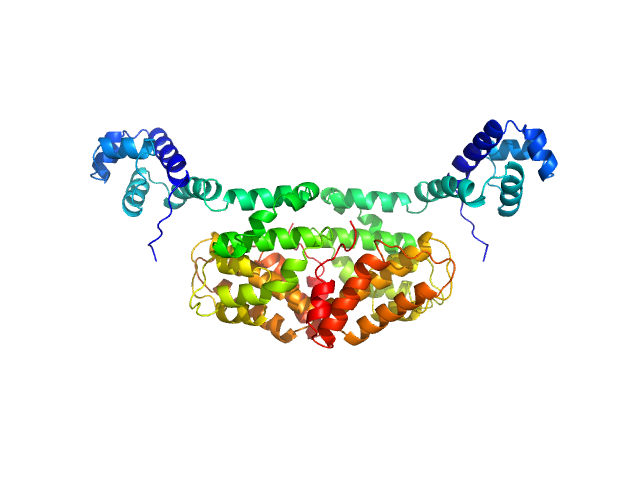
SaPIbov1 pathogenicity island repressor dimer, 64 kDa Staphylococcus aureus protein
|
| Buffer: |
50 mM HEPES 300 mM NaCl 5 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 9
|
Structural model of human dUTPase in complex with a novel proteinaceous inhibitor.
Sci Rep 8(1):4326 (2018)
Nyíri K, Mertens HDT, Tihanyi B, Nagy GN, Kőhegyi B, Matejka J, Harris MJ, Szabó JE, Papp-Kádár V, Németh-Pongrácz V, Ozohanics O, Vékey K, Svergun DI, Borysik AJ, Vértessy BG
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
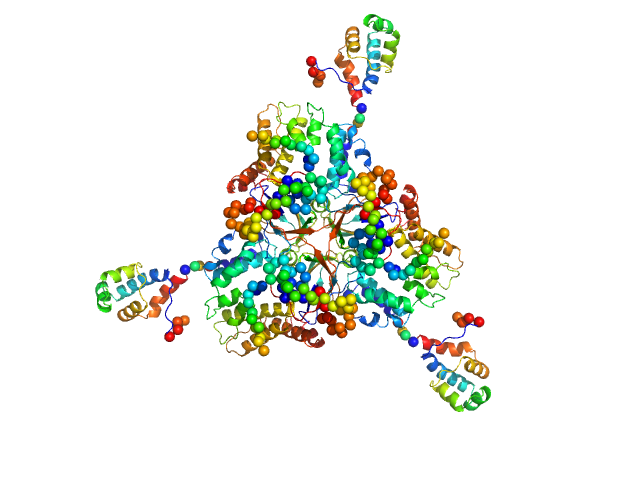
Deoxyuridine 5'-triphosphate nucleotidohydrolase trimer, 54 kDa Homo sapiens protein
SaPIbov1 pathogenicity island repressor dimer, 64 kDa Staphylococcus aureus protein
|
| Buffer: |
50 mM HEPES 300 mM NaCl 5 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 10
|
Structural model of human dUTPase in complex with a novel proteinaceous inhibitor.
Sci Rep 8(1):4326 (2018)
Nyíri K, Mertens HDT, Tihanyi B, Nagy GN, Kőhegyi B, Matejka J, Harris MJ, Szabó JE, Papp-Kádár V, Németh-Pongrácz V, Ozohanics O, Vékey K, Svergun DI, Borysik AJ, Vértessy BG
|
| RgGuinier |
4.4 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
227 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
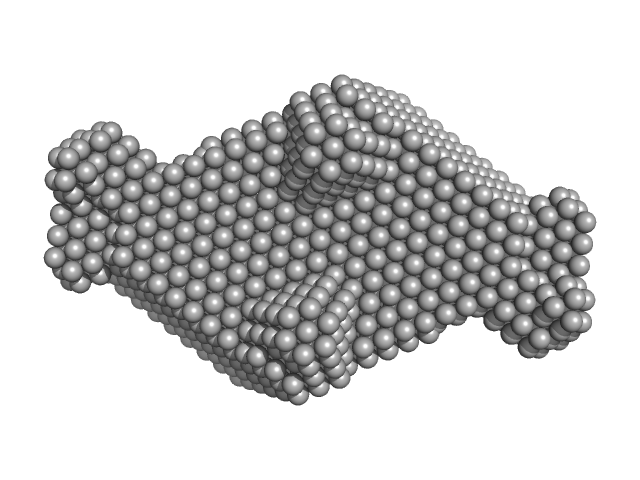
Deoxyuridine 5'-triphosphate nucleotidohydrolase trimer, 54 kDa Homo sapiens protein
SaPIbov1 pathogenicity island repressor dimer, 64 kDa Staphylococcus aureus protein
|
| Buffer: |
50 mM HEPES 300 mM NaCl 5 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 10
|
Structural model of human dUTPase in complex with a novel proteinaceous inhibitor.
Sci Rep 8(1):4326 (2018)
Nyíri K, Mertens HDT, Tihanyi B, Nagy GN, Kőhegyi B, Matejka J, Harris MJ, Szabó JE, Papp-Kádár V, Németh-Pongrácz V, Ozohanics O, Vékey K, Svergun DI, Borysik AJ, Vértessy BG
|
| RgGuinier |
3.8 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
|
|
|
|
|
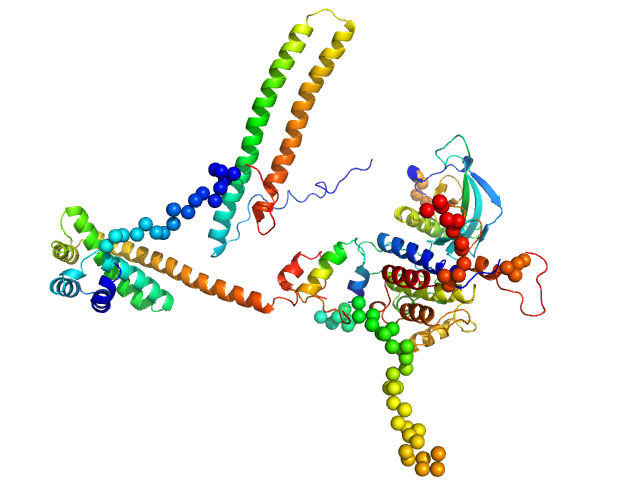
| Sample: |
Hsp90 co-chaperone Cdc37 monomer, 44 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris.Cl, 150 mM NaCl, 5% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 10
|
Disease Variants of FGFR3 Reveal Molecular Basis for the Recognition and Additional Roles for Cdc37 in Hsp90 Chaperone System.
Structure 26(3):446-458.e8 (2018)
Bunney TD, Inglis AJ, Sanfelice D, Farrell B, Kerr CJ, Thompson GS, Masson GR, Thiyagarajan N, Svergun DI, Williams RL, Breeze AL, Katan M
|
| RgGuinier |
4.0 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|
|
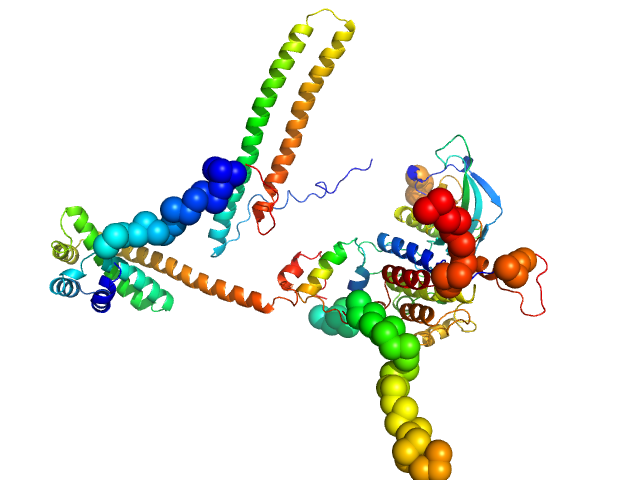
| Sample: |
Hsp90 co-chaperone Cdc37 monomer, 44 kDa Homo sapiens protein
Fibroblast growth factor receptor 3 monomer, 35 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris.Cl, 150 mM NaCl, 5% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 10
|
Disease Variants of FGFR3 Reveal Molecular Basis for the Recognition and Additional Roles for Cdc37 in Hsp90 Chaperone System.
Structure 26(3):446-458.e8 (2018)
Bunney TD, Inglis AJ, Sanfelice D, Farrell B, Kerr CJ, Thompson GS, Masson GR, Thiyagarajan N, Svergun DI, Williams RL, Breeze AL, Katan M
|
| RgGuinier |
4.7 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
161 |
nm3 |
|
|
|
|
|
|
|
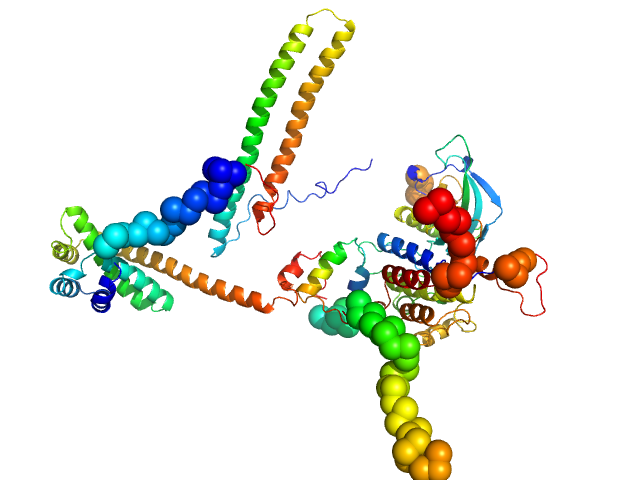
| Sample: |
Fibroblast growth factor receptor 3 monomer, 35 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris.Cl, 150 mM NaCl, 5% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 10
|
Disease Variants of FGFR3 Reveal Molecular Basis for the Recognition and Additional Roles for Cdc37 in Hsp90 Chaperone System.
Structure 26(3):446-458.e8 (2018)
Bunney TD, Inglis AJ, Sanfelice D, Farrell B, Kerr CJ, Thompson GS, Masson GR, Thiyagarajan N, Svergun DI, Williams RL, Breeze AL, Katan M
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
59 |
nm3 |
|
|