|
|
|
|
|
| Sample: |
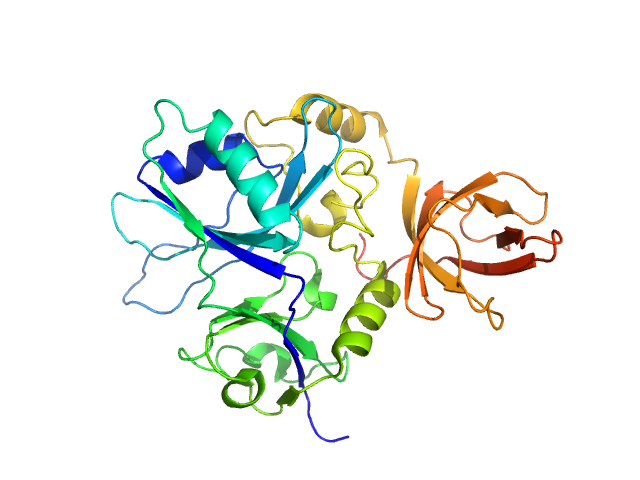
Putative transferase CAF17, mitochondrial monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
50 mM phosphate, 150 mM NaCl, 5 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 10
|
Structural properties of [2Fe-2S] ISCA2-IBA57: a complex of the mitochondrial iron-sulfur cluster assembly machinery
Scientific Reports 9(1) (2019)
Nasta V, Da Vela S, Gourdoupis S, Ciofi-Baffoni S, Svergun D, Banci L
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
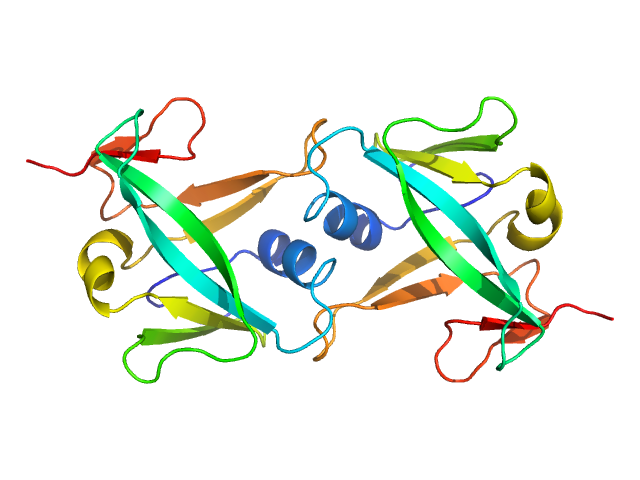
Iron-sulfur cluster assembly 2 homolog, mitochondrial dimer, 24 kDa Homo sapiens protein
|
| Buffer: |
50 mM phosphate, 150 mM NaCl, 5 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Sep 16
|
Structural properties of [2Fe-2S] ISCA2-IBA57: a complex of the mitochondrial iron-sulfur cluster assembly machinery
Scientific Reports 9(1) (2019)
Nasta V, Da Vela S, Gourdoupis S, Ciofi-Baffoni S, Svergun D, Banci L
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
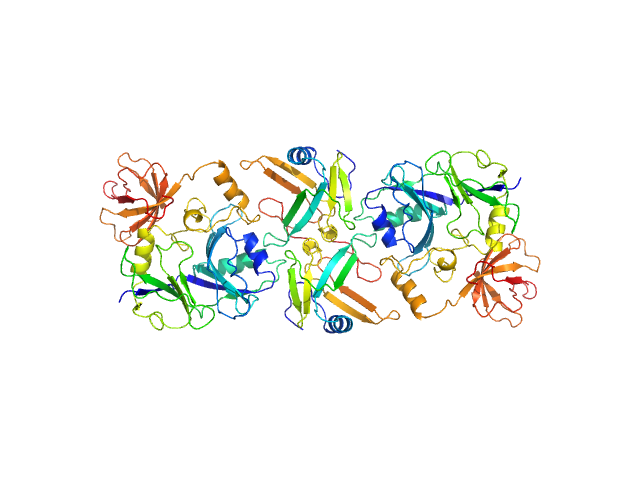
Putative transferase CAF17, mitochondrial monomer, 33 kDa Homo sapiens protein
Iron-sulfur cluster assembly 2 homolog, mitochondrial dimer, 24 kDa Homo sapiens protein
|
| Buffer: |
50 mM phosphate, 150 mM NaCl, 5 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Nov 22
|
Structural properties of [2Fe-2S] ISCA2-IBA57: a complex of the mitochondrial iron-sulfur cluster assembly machinery
Scientific Reports 9(1) (2019)
Nasta V, Da Vela S, Gourdoupis S, Ciofi-Baffoni S, Svergun D, Banci L
|
| RgGuinier |
4.0 |
nm |
| Dmax |
16.4 |
nm |
| VolumePorod |
114 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
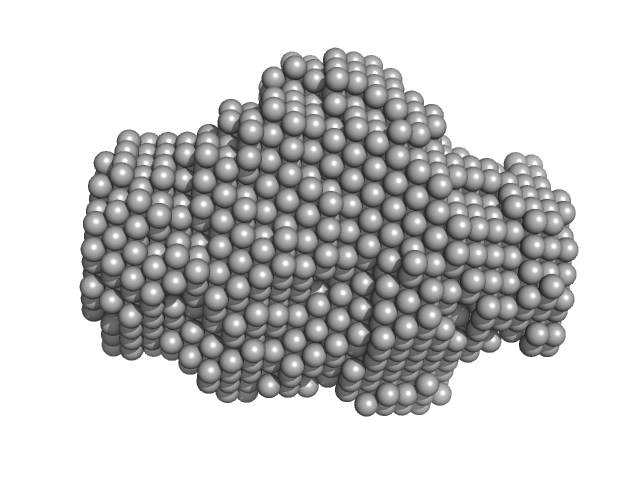
Bruton's tyrosine kinase, kinase domain monomer, 32 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 11
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.8 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bruton's tyrosine kinase - Src homolgy domain 2 - kinase domain monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Nov 3
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
64 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bruton's tyrosine kinase - Src homology 3-2 kinase domain monomer, 52 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Nov 2
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bruton's tyrosine kinase - full length monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris, 150mM NaCl, 1mM TCEP, 5% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Nov 3
|
Btk SH2-kinase interface is critical for allosteric kinase activation and its targeting inhibits B-cell neoplasms
Nature Communications 11(1) (2020)
Duarte D, Lamontanara A, La Sala G, Jeong S, Sohn Y, Panjkovich A, Georgeon S, Kükenshöner T, Marcaida M, Pojer F, De Vivo M, Svergun D, Kim H, Dal Peraro M, Hantschel O
|
| RgGuinier |
4.0 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
114 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
EKC/KEOPS complex subunit GON7 monomer, 13 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES, 200 mM NaCl, 5 mM β-mercaptoethanol, pH: 6.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Mar 26
|
Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome.
Nat Commun 10(1):3967 (2019)
Arrondel C, Missoury S, Snoek R, Patat J, Menara G, Collinet B, Liger D, Durand D, Gribouval O, Boyer O, Buscara L, Martin G, Machuca E, Nevo F, Lescop E, Braun DA, Boschat AC, Sanquer S, Guerrera IC, Revy P, Parisot M, Masson C, Boddaert N, Charbit M, Decramer S, Novo R, Macher MA, Ranchin B, Bacchetta J, Laurent A, Collardeau-Frachon S, van Eerde AM, Hildebrandt F, Magen D, Antignac C, van Tilbeurgh H, Mollet G
|
| RgGuinier |
3.1 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
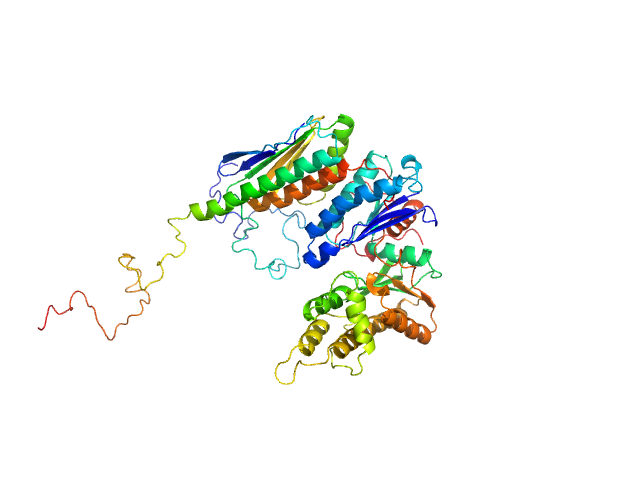
EKC/KEOPS complex subunit GON7 monomer, 12 kDa Homo sapiens protein
EKC/KEOPS complex subunit LAGE3 monomer, 15 kDa Homo sapiens protein
Probable tRNA N6-adenosine threonylcarbamoyltransferase monomer, 36 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES, 200 mM NaCl, 5 mM β-mercaptoethanol, pH: 6.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 May 12
|
Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome.
Nat Commun 10(1):3967 (2019)
Arrondel C, Missoury S, Snoek R, Patat J, Menara G, Collinet B, Liger D, Durand D, Gribouval O, Boyer O, Buscara L, Martin G, Machuca E, Nevo F, Lescop E, Braun DA, Boschat AC, Sanquer S, Guerrera IC, Revy P, Parisot M, Masson C, Boddaert N, Charbit M, Decramer S, Novo R, Macher MA, Ranchin B, Bacchetta J, Laurent A, Collardeau-Frachon S, van Eerde AM, Hildebrandt F, Magen D, Antignac C, van Tilbeurgh H, Mollet G
|
| RgGuinier |
3.1 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
91 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
EKC/KEOPS complex subunit GON7 dimer, 25 kDa Homo sapiens protein
EKC/KEOPS complex subunit LAGE3 dimer, 33 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES, 200 mM NaCl, 5 mM β-mercaptoethanol, pH: 6.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Mar 19
|
Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome.
Nat Commun 10(1):3967 (2019)
Arrondel C, Missoury S, Snoek R, Patat J, Menara G, Collinet B, Liger D, Durand D, Gribouval O, Boyer O, Buscara L, Martin G, Machuca E, Nevo F, Lescop E, Braun DA, Boschat AC, Sanquer S, Guerrera IC, Revy P, Parisot M, Masson C, Boddaert N, Charbit M, Decramer S, Novo R, Macher MA, Ranchin B, Bacchetta J, Laurent A, Collardeau-Frachon S, van Eerde AM, Hildebrandt F, Magen D, Antignac C, van Tilbeurgh H, Mollet G
|
| RgGuinier |
3.6 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
126 |
nm3 |
|
|