|
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM Acetate, 72% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2019 Sep 21
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
|
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2019 Sep 21
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
|
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
|
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
GlcNAc-binding protein A monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
|
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
GlcNAc-binding protein A monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
|
|
|
|
|
| Sample: |
Endo-β-1,4-xylanase (AcXyn30B_12) monomer, 74 kDa Acetivibrio clariflavus protein
|
| Buffer: |
50 mM Sodium Phosphate, pH: 7.5 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, Medical University of Graz on 2024 Jan 4
|
Deciphering the structural insights and concentration-dependent dimerisation of endo-β-1,4-xylanase (AcXyn30B_12) from Acetivibrio clariflavus using SAXS and computational methods.
FEBS J (2026)
Choudhury B, Goyal A
|
|
|
|
|
|
|

|
| Sample: |
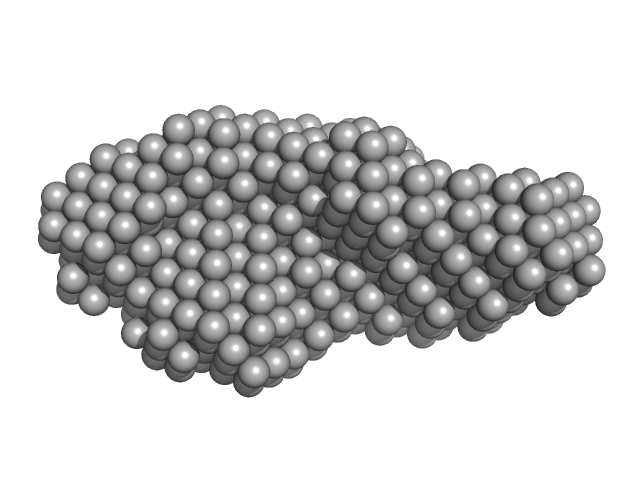
15 nucleotide RNA duplex (ATP-dependent RNA helicase DDX3X binding target) dimer, 10 kDa RNA
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10% (v/v) glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Jun 26
|
Solution structures of DEAD-box helicase DDX3X reveal the N-terminal extension binds RNA to modulate catalysis and influence conformation
Sarah Atkinson
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.2 |
nm |
| VolumePorod |
14 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
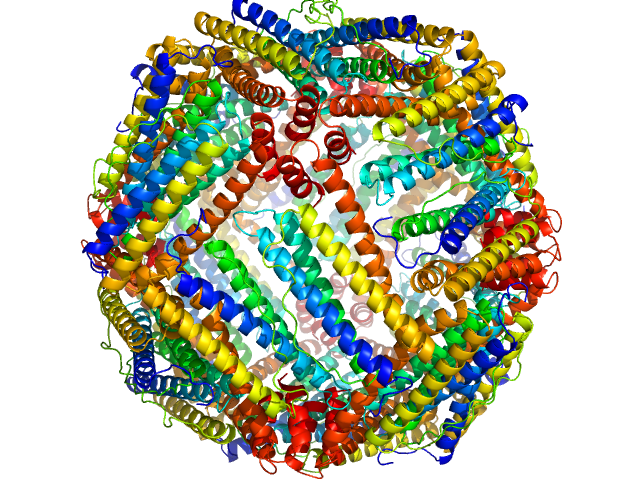
Ferritin light chain 24-mer, 476 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
5.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
651 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
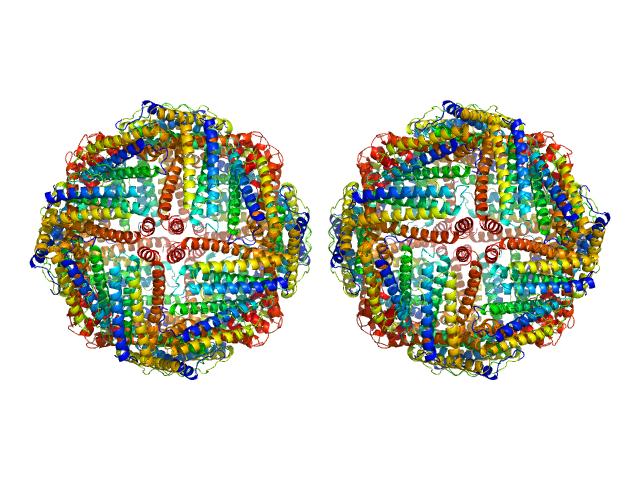
Ferritin light chain, 953 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
7.7 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
1751 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
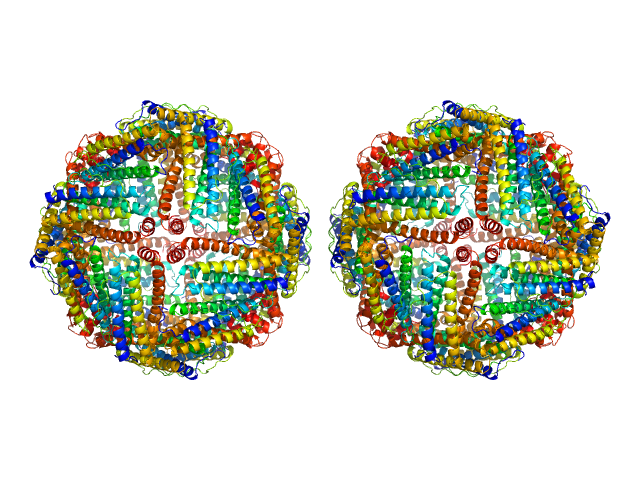
Ferritin light chain, 953 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
8.0 |
nm |
| Dmax |
26.8 |
nm |
| VolumePorod |
1369 |
nm3 |
|
|