|
|
|
|
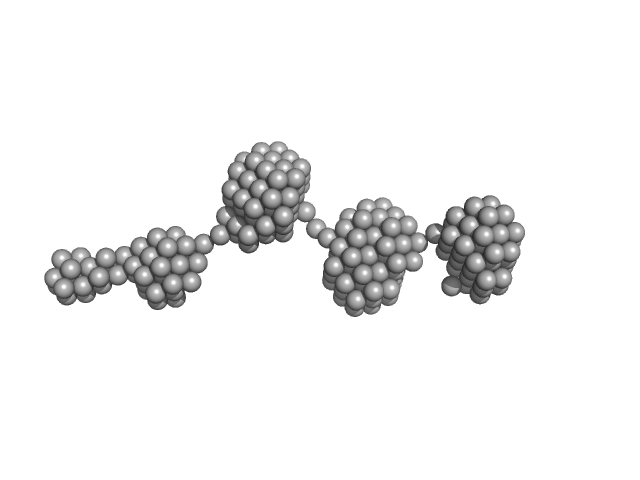
|
| Sample: |
Epstein-Barr nuclear antigen 2 dimer, 16 kDa Human gammaherpesvirus 4 protein
|
| Buffer: |
20mM Tris-HCl, 100mM NaCl, 2% Sucrose and 1mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Sep 23
|
Increased association between Epstein-Barr virus EBNA2 from type 2 strains and the transcriptional repressor BS69 restricts B cell growth
(2018)
Ponnusamy R, Khatri R, Correia P, Mancini E, Farrell P, West M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
20 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
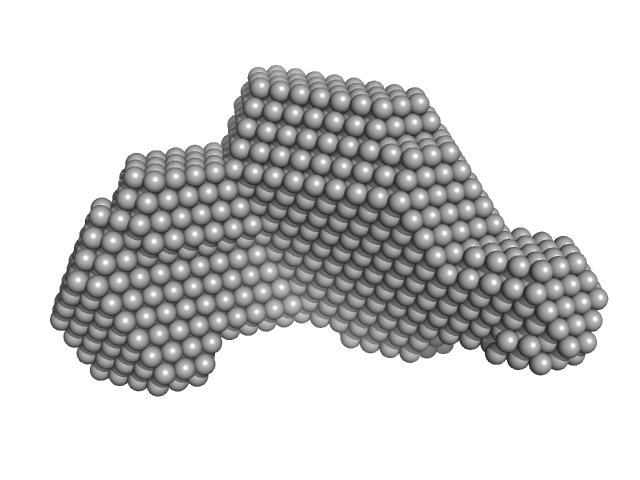
Transcriptional regulator Lrs14-like protein dimer, 33 kDa Sulfolobus acidocaldarius protein
|
| Buffer: |
300 mM NaCl, 20 mM HEPES, pH 7.5, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 May 5
|
Crystal structure of an Lrs14-like archaeal biofilm regulator from Sulfolobus acidocaldarius.
Acta Crystallogr D Struct Biol 74(Pt 11):1105-1114 (2018)
Vogt MS, Völpel SL, Albers SV, Essen LO, Banerjee A
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Glutamate receptor 2 monomer, 368 kDa Rattus norvegicus protein
|
| Buffer: |
D2O based buffer. 20 mM Tris/DCl, 100 mM NaCl, 0.5 mM deuterated n-dodecyl-β-D-maltopyranoside (synthesized to match out at 100% D2O), pH: 7.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Sep 19
|
Small-angle neutron scattering studies on the AMPA receptor GluA2 in the resting, AMPA-bound and GYKI-53655-bound states.
IUCrJ 5(Pt 6):780-793 (2018)
Larsen AH, Dorosz J, Thorsen TS, Johansen NT, Darwish T, Midtgaard SR, Arleth L, Kastrup JS
|
| RgGuinier |
6.0 |
nm |
| Dmax |
17.9 |
nm |
| VolumePorod |
396 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Glutamate receptor 2 monomer, 368 kDa Rattus norvegicus protein
|
| Buffer: |
D2O based buffer. 1 mM AMPA, 20 mM Tris/DCl, 100 mM NaCl, 0.5 mM deuterated n-dodecyl-β-D-maltopyranoside (synthesized to match out at 100% D2O), pH: 7.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2016 Oct 19
|
Small-angle neutron scattering studies on the AMPA receptor GluA2 in the resting, AMPA-bound and GYKI-53655-bound states.
IUCrJ 5(Pt 6):780-793 (2018)
Larsen AH, Dorosz J, Thorsen TS, Johansen NT, Darwish T, Midtgaard SR, Arleth L, Kastrup JS
|
| RgGuinier |
6.3 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
407 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Glutamate receptor 2 monomer, 368 kDa Rattus norvegicus protein
|
| Buffer: |
D2O based buffer. 10 mM AMPA, 20 mM Tris/DCl, 100 mM NaCl, 0.5 mM deuterated n-dodecyl-β-D-maltopyranoside (synthesized to match out at 100% D2O), pH: 5.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Sep 19
|
Small-angle neutron scattering studies on the AMPA receptor GluA2 in the resting, AMPA-bound and GYKI-53655-bound states.
IUCrJ 5(Pt 6):780-793 (2018)
Larsen AH, Dorosz J, Thorsen TS, Johansen NT, Darwish T, Midtgaard SR, Arleth L, Kastrup JS
|
| RgGuinier |
6.5 |
nm |
| Dmax |
18.9 |
nm |
| VolumePorod |
899 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Glutamate receptor 2 monomer, 368 kDa Rattus norvegicus protein
|
| Buffer: |
D2O based buffer. 1 mM GYKI-53655, 20 mM Tris/DCl, 100 mM NaCl, 0.5 mM deuterated n-dodecyl-β-D-maltopyranoside (synthesized to match out at 100% D2O), pH: 7.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2016 Oct 19
|
Small-angle neutron scattering studies on the AMPA receptor GluA2 in the resting, AMPA-bound and GYKI-53655-bound states.
IUCrJ 5(Pt 6):780-793 (2018)
Larsen AH, Dorosz J, Thorsen TS, Johansen NT, Darwish T, Midtgaard SR, Arleth L, Kastrup JS
|
| RgGuinier |
6.3 |
nm |
| Dmax |
18.6 |
nm |
| VolumePorod |
384 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Paenibacillus xanthan lyase monomer, 113 kDa Paenibacillus sp-62047 protein
|
| Buffer: |
20 mM Tris,, pH: 8.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Dec 15
|
Structure and Dynamics of a Promiscuous Xanthan Lyase from Paenibacillus nanensis and the Design of Variants with Increased Stability and Activity.
Cell Chem Biol 26(2):191-202.e6 (2019)
Jensen PF, Kadziola A, Comamala G, Segura DR, Anderson L, Poulsen JN, Rasmussen KK, Agarwal S, Sainathan RK, Monrad RN, Svendsen A, Nielsen JE, Lo Leggio L, Rand KD
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
137 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Paenibacillus xanthan lyase monomer, 113 kDa Paenibacillus sp-62047 protein
|
| Buffer: |
20 mM Tris,, pH: 8.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Dec 15
|
Structure and Dynamics of a Promiscuous Xanthan Lyase from Paenibacillus nanensis and the Design of Variants with Increased Stability and Activity.
Cell Chem Biol 26(2):191-202.e6 (2019)
Jensen PF, Kadziola A, Comamala G, Segura DR, Anderson L, Poulsen JN, Rasmussen KK, Agarwal S, Sainathan RK, Monrad RN, Svendsen A, Nielsen JE, Lo Leggio L, Rand KD
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.8 |
nm |
| VolumePorod |
134 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
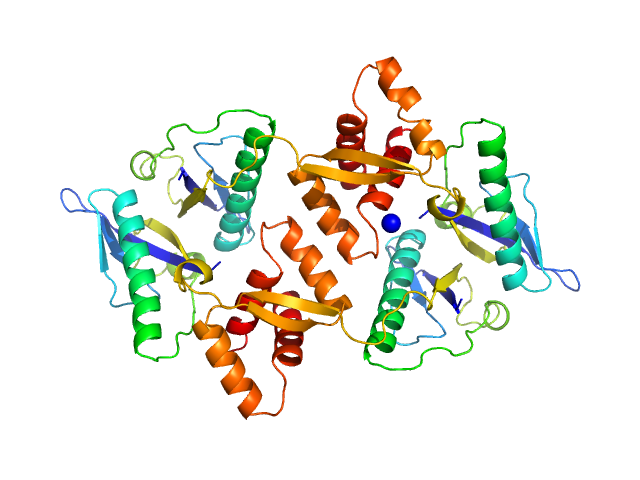
Type II toxin-antitoxin system HicB family antitoxin tetramer, 63 kDa Burkholderia pseudomallei protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Jul 28
|
The molecular basis of protein toxin HicA-dependent binding of the protein antitoxin HicB to DNA.
J Biol Chem (2018)
Winter AJ, Williams C, Isupov MN, Crocker H, Gromova M, Marsh P, Wilkinson OJ, Dillingham MS, Harmer NJ, Titball RW, Crump MP
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
112 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
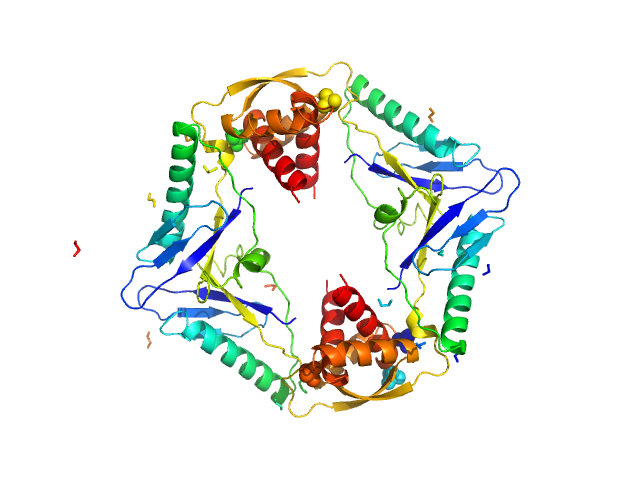
Type II toxin-antitoxin system HicB family antitoxin tetramer, 63 kDa Burkholderia pseudomallei protein
Addiction module toxin, HicA monomer, 7 kDa Burkholderia pseudomallei protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Jul 27
|
The molecular basis of protein toxin HicA-dependent binding of the protein antitoxin HicB to DNA.
J Biol Chem (2018)
Winter AJ, Williams C, Isupov MN, Crocker H, Gromova M, Marsh P, Wilkinson OJ, Dillingham MS, Harmer NJ, Titball RW, Crump MP
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
110 |
nm3 |
|
|