|
|
|
|

|
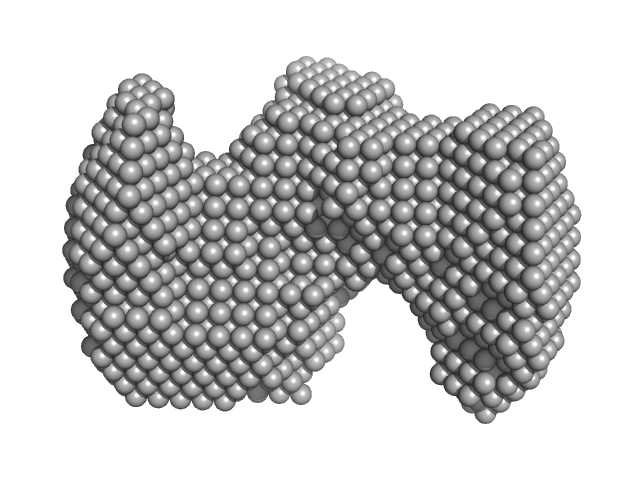
| Sample: |
Monekypox DNA sequence 1 monomer, 6 kDa Monkeypox virus DNA
|
| Buffer: |
20 mM HEPES, 100 mM KCl, and 0.2 mM EDTA, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Sep 23
|
Mapping and characterization of G‐quadruplexes in monkeypox genomes
Journal of Medical Virology 95(5) (2023)
Pereira H, Gemmill D, Siddiqui M, Vasudeva G, Patel T
|
| RgGuinier |
1.6 |
nm |
| Dmax |
4.5 |
nm |
| VolumePorod |
11 |
nm3 |
|
|
|
|
|
|

|
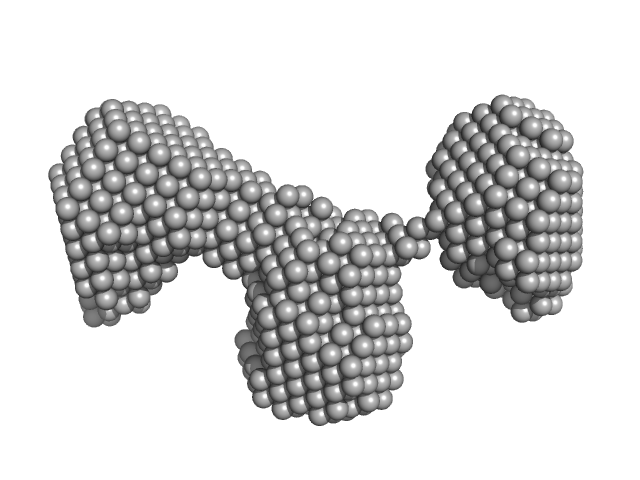
| Sample: |
Monekypox DNA sequence 1 mutant monomer, 6 kDa Monkeypox virus DNA
|
| Buffer: |
20 mM HEPES, 100 mM KCl, and 0.2 mM EDTA, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Sep 23
|
Mapping and characterization of G‐quadruplexes in monkeypox genomes
Journal of Medical Virology 95(5) (2023)
Pereira H, Gemmill D, Siddiqui M, Vasudeva G, Patel T
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.5 |
nm |
| VolumePorod |
16 |
nm3 |
|
|
|
|
|
|
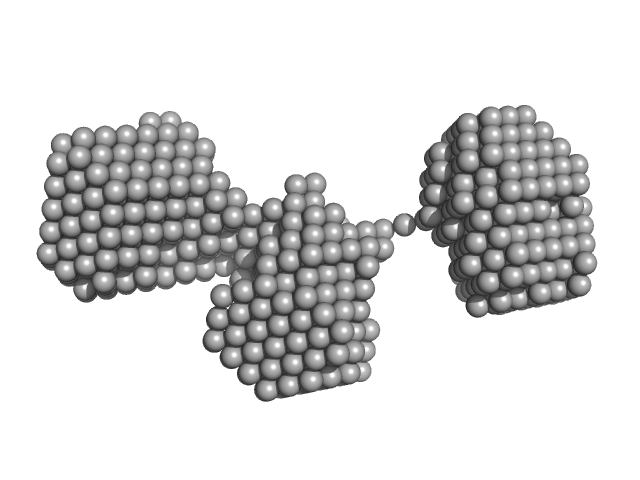
|
| Sample: |
Monekypox DNA sequence 2 mutant monomer, 6 kDa Monkeypox virus DNA
|
| Buffer: |
20 mM HEPES, 100 mM KCl, and 0.2 mM EDTA, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Sep 23
|
Mapping and characterization of G‐quadruplexes in monkeypox genomes
Journal of Medical Virology 95(5) (2023)
Pereira H, Gemmill D, Siddiqui M, Vasudeva G, Patel T
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.0 |
nm |
| VolumePorod |
8 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
Chaotic advection mixer for capturing transient states of diverse biological macromolecular systems with time-resolved small-angle X-ray scattering
IUCrJ 10(3):363-375 (2023)
Zielinski K, Katz A, Calvey G, Pabit S, Milano S, Aplin C, San Emeterio J, Cerione R, Pollack L
|
| RgGuinier |
2.2 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
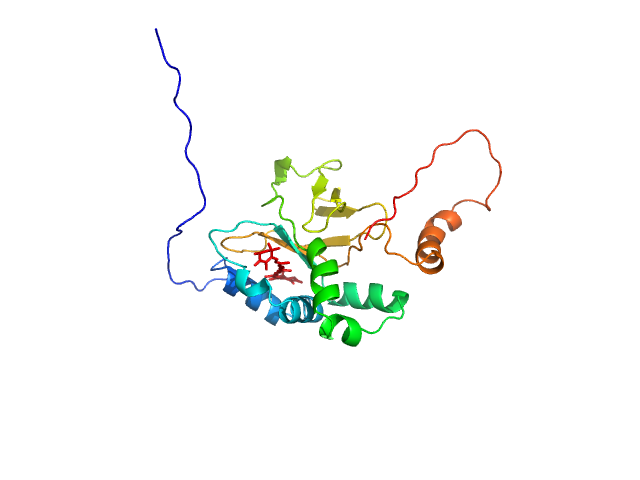
Astaxanthin binding fasciclin family protein monomer, 22 kDa Coelastrella astaxanthina protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 7.6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Sep 24
|
Structural basis for the ligand promiscuity of the neofunctionalized, carotenoid-binding fasciclin domain protein AstaP.
Commun Biol 6(1):471 (2023)
Kornilov FD, Slonimskiy YB, Lunegova DA, Egorkin NA, Savitskaya AG, Kleymenov SY, Maksimov EG, Goncharuk SA, Mineev KS, Sluchanko NN
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
43 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
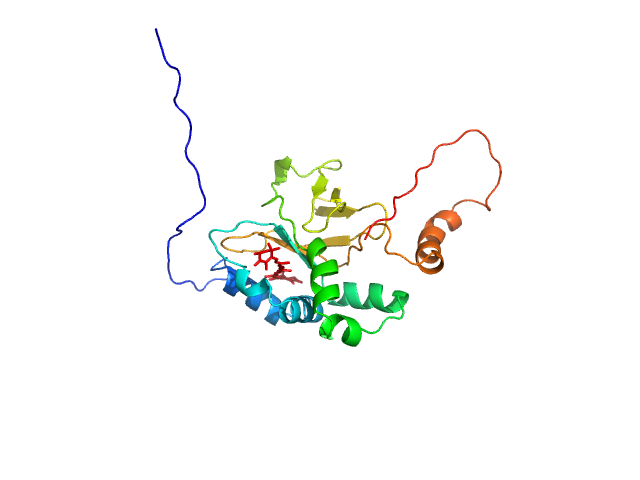
Astaxanthin binding fasciclin family protein monomer, 22 kDa Coelastrella astaxanthina protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 7.6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Sep 24
|
Structural basis for the ligand promiscuity of the neofunctionalized, carotenoid-binding fasciclin domain protein AstaP.
Commun Biol 6(1):471 (2023)
Kornilov FD, Slonimskiy YB, Lunegova DA, Egorkin NA, Savitskaya AG, Kleymenov SY, Maksimov EG, Goncharuk SA, Mineev KS, Sluchanko NN
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
65 kDa invariant surface glycoprotein, putative monomer, 41 kDa Trypanosoma brucei gambiense … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3% (v/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Nov 25
|
Cryo-EM structures of Trypanosoma brucei gambiense ISG65 with human complement C3 and C3b and their roles in alternative pathway restriction.
Nat Commun 14(1):2403 (2023)
Sülzen H, Began J, Dhillon A, Kereïche S, Pompach P, Votrubova J, Zahedifard F, Šubrtova A, Šafner M, Hubalek M, Thompson M, Zoltner M, Zoll S
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
53 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Humanized immunoglobulin G1 monoclonal antibody monomer, 148 kDa Mouse/human (chimera)
|
| Buffer: |
100 mM glycine-HCl, pH: 2 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2016 Mar 2
|
Getting Smaller by Denaturation: Acid-Induced Compaction of Antibodies
The Journal of Physical Chemistry Letters :3898-3906 (2023)
Imamura H, Ooishi A, Honda S
|
| RgGuinier |
5.1 |
nm |
| Dmax |
17.6 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Humanized immunoglobulin G1 monoclonal antibody monomer, 148 kDa Mouse/human (chimera)
|
| Buffer: |
100 mM glycine-HCl, 200 mM NaCl, pH: 2 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2017 Mar 5
|
Getting Smaller by Denaturation: Acid-Induced Compaction of Antibodies
The Journal of Physical Chemistry Letters :3898-3906 (2023)
Imamura H, Ooishi A, Honda S
|
| RgGuinier |
4.1 |
nm |
| Dmax |
14.2 |
nm |
| VolumePorod |
240 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
40 mM sodium acetate, 50 mM NaCl, pH: 4 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2021 Apr 2
|
NSRRC TPS13A standard protein archive
Orion Shih
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.6 |
nm |
| VolumePorod |
13 |
nm3 |
|
|