|
|
|
|
|
| Sample: |
Synthetic FRET biosensor Twitch 2B monomer, 62 kDa synthetic construct protein
|
| Buffer: |
20 mM MOPS 100 mM NaCl 10 mM EDTA, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Mar 17
|
Synthetic FRET biosensor Twitch 2B
Pablo Trigo Mourino
|
|
|
|
|
|
|
|
| Sample: |
Synthetic FRET biosensor Twitch2B N532F monomer, 62 kDa synthetic construct protein
|
| Buffer: |
20 mM MOPS 100 mM NaCl 5 mM CaCl2, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jun 30
|
Synthetic FRET biosensor Twitch 2B
Pablo Trigo Mourino
|
|
|
|
|
|
|
|
| Sample: |
Synthetic FRET biosensor Twitch2B N532F monomer, 62 kDa synthetic construct protein
|
| Buffer: |
20 mM MOPS 100 mM NaCl 10 mM EDTA, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jun 30
|
Synthetic FRET biosensor Twitch 2B
Pablo Trigo Mourino
|
|
|
|
|
|
|

|
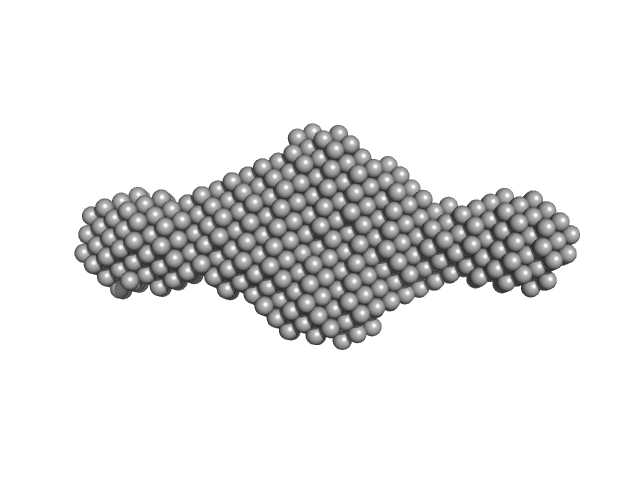
| Sample: |
5-10-5 gapmer phosphorothioate antisense oligonucleotide dimer, 14 kDa
|
| Buffer: |
20 mM Tris-Cl (pH 7.5), 250 mM KCl, 50 mM L-proline, 0.5 mM EDTA, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Apr 28
|
Structural basis of dimerization and nucleic acid binding of human DBHS proteins NONO and PSPC1.
Nucleic Acids Res (2021)
Knott GJ, Chong YS, Passon DM, Liang XH, Deplazes E, Conte MR, Marshall AC, Lee M, Fox AH, Bond CS
|
| RgGuinier |
1.8 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
15 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
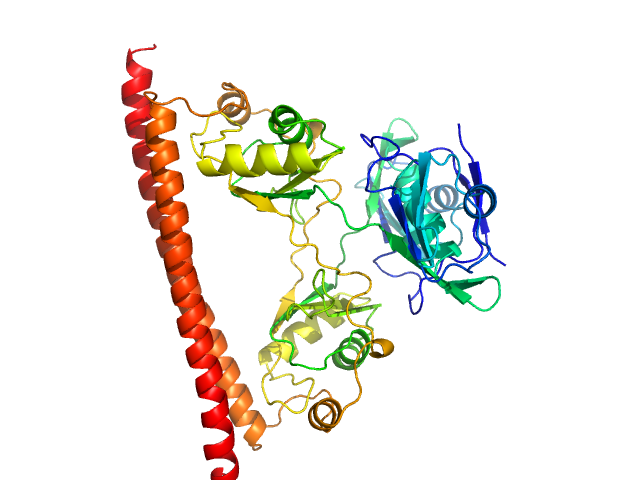
Non-POU domain-containing octamer-binding protein dimer, 60 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-Cl (pH 7.5), 250 mM KCl, 50 mM L-proline, 0.5 mM EDTA, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Apr 28
|
Structural basis of dimerization and nucleic acid binding of human DBHS proteins NONO and PSPC1.
Nucleic Acids Res (2021)
Knott GJ, Chong YS, Passon DM, Liang XH, Deplazes E, Conte MR, Marshall AC, Lee M, Fox AH, Bond CS
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
96 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
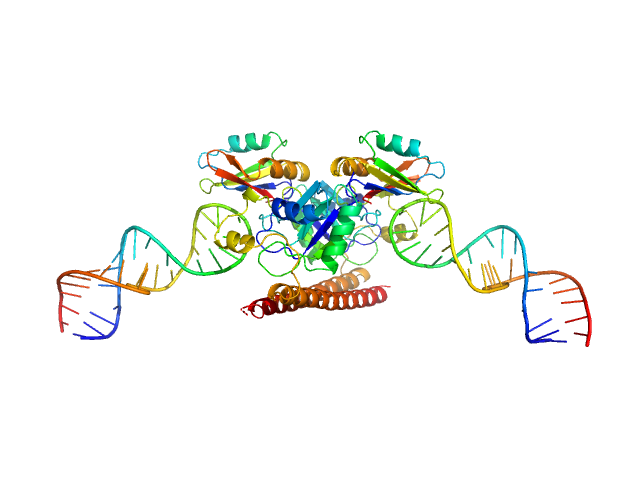
Non-POU domain-containing octamer-binding protein dimer, 60 kDa Homo sapiens protein
5-10-5 gapmer phosphorothioate antisense oligonucleotide tetramer tetramer, 28 kDa
|
| Buffer: |
20 mM Tris-Cl (pH 7.5), 250 mM KCl, 50 mM L-proline, 0.5 mM EDTA, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Apr 28
|
Structural basis of dimerization and nucleic acid binding of human DBHS proteins NONO and PSPC1.
Nucleic Acids Res (2021)
Knott GJ, Chong YS, Passon DM, Liang XH, Deplazes E, Conte MR, Marshall AC, Lee M, Fox AH, Bond CS
|
| RgGuinier |
3.9 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
153 |
nm3 |
|
|
|
|
|
|

|
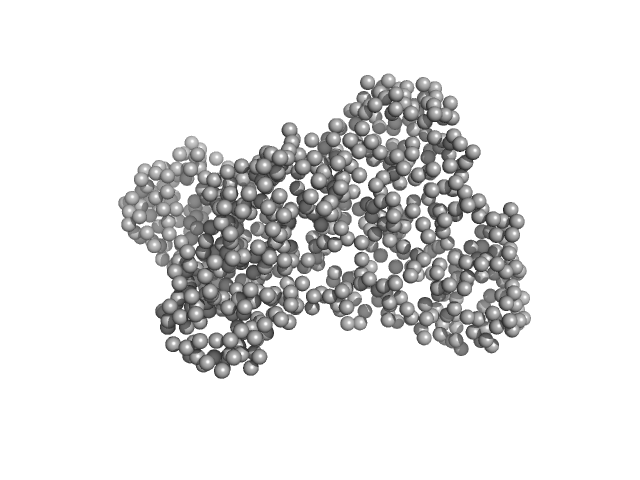
| Sample: |
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl, (APO Buffer), pH: 5.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
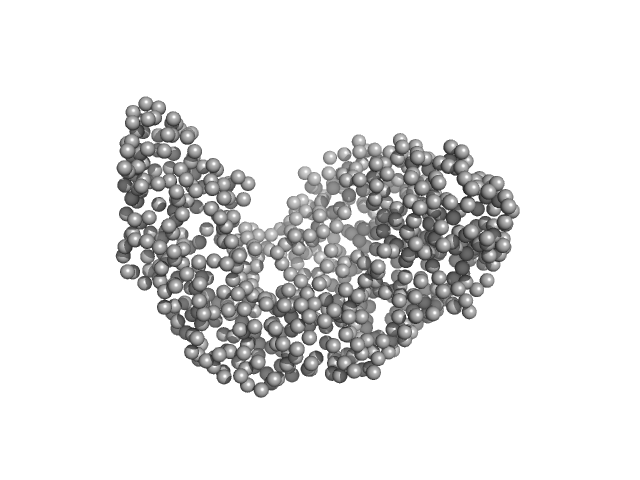
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (APO Buffer), pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.6 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
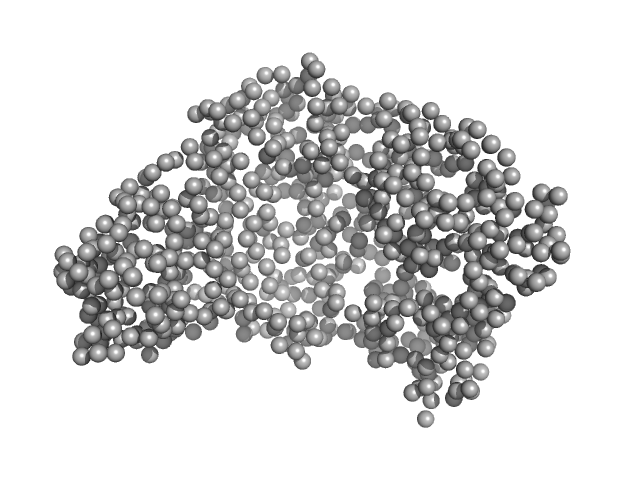
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (Buffer APO-Tf-1), pH: 8
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|

|
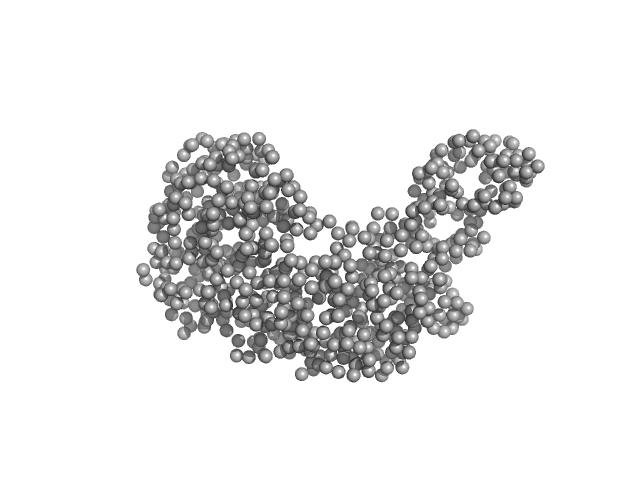
| Sample: |
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl, (APO Buffer), pH: 5.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.3 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
107 |
nm3 |
|
|