|
|
|
|

|
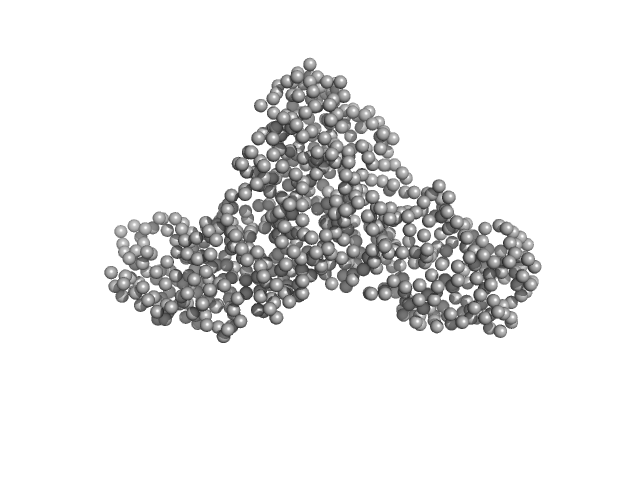
| Sample: |
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (APO Buffer), pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.2 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
103 |
nm3 |
|
|
|
|
|
|

|
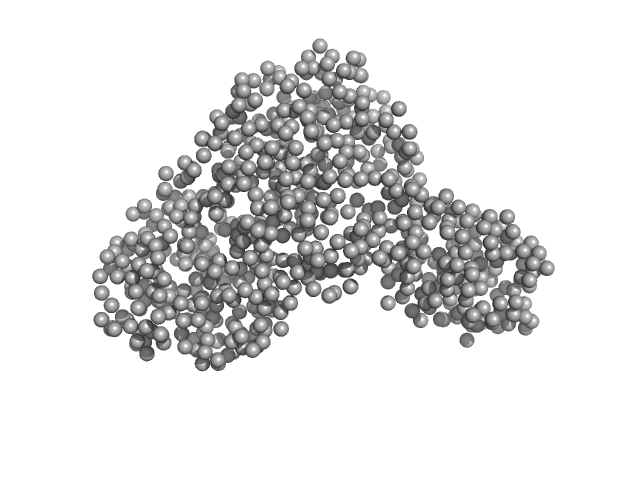
| Sample: |
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (Buffer APO-Tf-1), pH: 8
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.2 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
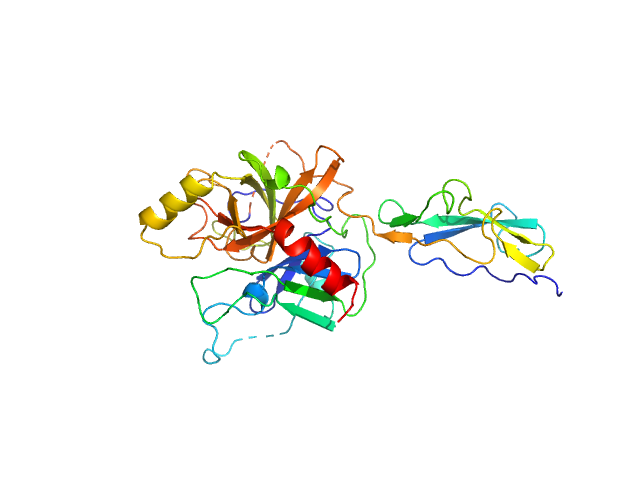
Complement C1r subcomponent monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES, 140 mM NaCl, pH: 7.3
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 24
|
A Structural Basis for Inhibition of the Complement Initiator Protease C1r by Lyme Disease Spirochetes.
J Immunol 207(11):2856-2867 (2021)
Garrigues RJ, Powell-Pierce AD, Hammel M, Skare JT, Garcia BL
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
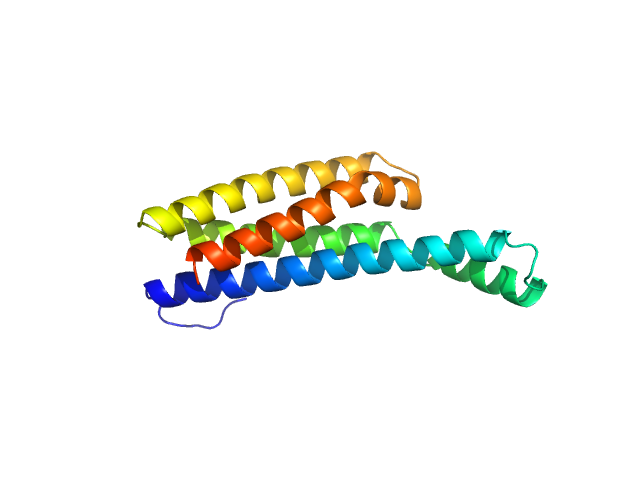
Fibronectin-binding protein BBK32 monomer, 17 kDa Borreliella burgdorferi B31 protein
|
| Buffer: |
10 mM HEPES, 140 mM NaCl, pH: 7.3
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Sep 24
|
A Structural Basis for Inhibition of the Complement Initiator Protease C1r by Lyme Disease Spirochetes.
J Immunol 207(11):2856-2867 (2021)
Garrigues RJ, Powell-Pierce AD, Hammel M, Skare JT, Garcia BL
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
34 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
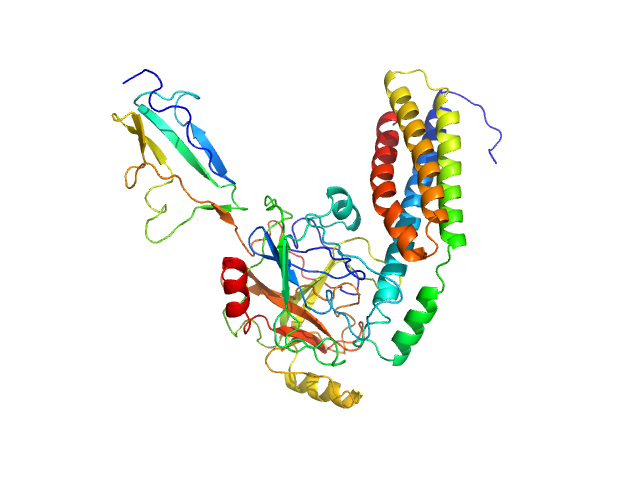
Fibronectin-binding protein BBK32 monomer, 17 kDa Borrelia burgdorferi (strain … protein
Complement C1r subcomponent monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES, 140 mM NaCl, pH: 7.3
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 24
|
A Structural Basis for Inhibition of the Complement Initiator Protease C1r by Lyme Disease Spirochetes.
J Immunol 207(11):2856-2867 (2021)
Garrigues RJ, Powell-Pierce AD, Hammel M, Skare JT, Garcia BL
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Guanidin-II riboswitch 23mer monomer, 7 kDa Escherichia coli RNA
|
| Buffer: |
25 mM potassium phosphate, 50 mM KCl, pH: 6.2
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Feb 13
|
Structure and Dynamics of the Guanidine-II Riboswitch from Escherichia coli by NMR Spectroscopy and Small-angle X-ray Scattering (SAXS).
Chembiochem (2021)
Schamber T, Binas O, Schlundt A, Wacker A, Schwalbe H
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.8 |
nm |
| VolumePorod |
14 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Guanidin-II riboswitch 23mer monomer, 7 kDa Escherichia coli RNA
|
| Buffer: |
25 mM potassium phosphate, 50 mM KCl, 10 mM Mg2+, pH: 6.2
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Feb 13
|
Structure and Dynamics of the Guanidine-II Riboswitch from Escherichia coli by NMR Spectroscopy and Small-angle X-ray Scattering (SAXS).
Chembiochem (2021)
Schamber T, Binas O, Schlundt A, Wacker A, Schwalbe H
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
15 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Guanidin-II riboswitch 23mer monomer, 7 kDa Escherichia coli RNA
|
| Buffer: |
25 mM potassium phosphate, 50 mM KCl, 10 mM Mg2+, 5 mM GdnCl, pH: 6.2
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Feb 13
|
Structure and Dynamics of the Guanidine-II Riboswitch from Escherichia coli by NMR Spectroscopy and Small-angle X-ray Scattering (SAXS).
Chembiochem (2021)
Schamber T, Binas O, Schlundt A, Wacker A, Schwalbe H
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Guanidin-II riboswitch 49mer dimer, 32 kDa Escherichia coli RNA
|
| Buffer: |
25 mM potassium phosphate, 50 mM KCl, pH: 6.2
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Feb 13
|
Structure and Dynamics of the Guanidine-II Riboswitch from Escherichia coli by NMR Spectroscopy and Small-angle X-ray Scattering (SAXS).
Chembiochem (2021)
Schamber T, Binas O, Schlundt A, Wacker A, Schwalbe H
|
| RgGuinier |
4.1 |
nm |
| Dmax |
16.9 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Guanidin-II riboswitch 49mer dimer, 32 kDa Escherichia coli RNA
|
| Buffer: |
25 mM potassium phosphate, 50 mM KCl, 10 mM Mg2+, pH: 6.2
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Feb 13
|
Structure and Dynamics of the Guanidine-II Riboswitch from Escherichia coli by NMR Spectroscopy and Small-angle X-ray Scattering (SAXS).
Chembiochem (2021)
Schamber T, Binas O, Schlundt A, Wacker A, Schwalbe H
|
| RgGuinier |
4.2 |
nm |
| Dmax |
18.1 |
nm |
| VolumePorod |
57 |
nm3 |
|
|