|
|
|
|
|
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
synthetic peptide AE234 monomer, 1 kDa protein
|
| Buffer: |
25 mM Sodium Phosphate 5 % Glycerol 50 mM NaSO4, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Jun 10
|
A flexible multidomain structure drives the function of the urokinase-type plasminogen activator receptor (uPAR).
J Biol Chem 287(41):34304-15 (2012)
Mertens HD, Kjaergaard M, Mysling S, Gårdsvoll H, Jørgensen TJ, Svergun DI, Ploug M
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
Urokinase-type plasminogen activator monomer, 49 kDa Homo sapiens protein
|
| Buffer: |
25 mM Sodium Phosphate 5 % Glycerol 50 mM NaSO4, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Jun 10
|
A flexible multidomain structure drives the function of the urokinase-type plasminogen activator receptor (uPAR).
J Biol Chem 287(41):34304-15 (2012)
Mertens HD, Kjaergaard M, Mysling S, Gårdsvoll H, Jørgensen TJ, Svergun DI, Ploug M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
|
|
|
|
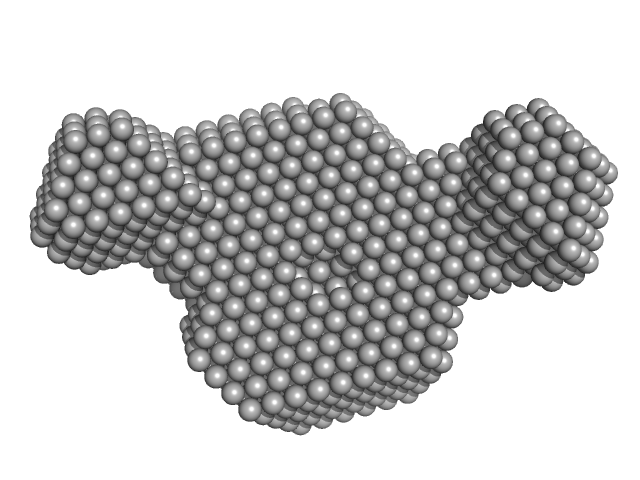

|
| Sample: |
Human Chromatin Remodeler CHD4 (363-682) monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES 50 mM KCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 13
|
The PHD and chromo domains regulate the ATPase activity of the human chromatin remodeler CHD4.
J Mol Biol 422(1):3-17 (2012)
Watson AA, Mahajan P, Mertens HD, Deery MJ, Zhang W, Pham P, Du X, Bartke T, Zhang W, Edlich C, Berridge G, Chen Y, Burgess-Brown NA, Kouzarides T, Wiechens N, Owen-Hughes T, Svergun DI, Gileadi O, La...
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.5 |
nm |
|
|
|
|
|
|
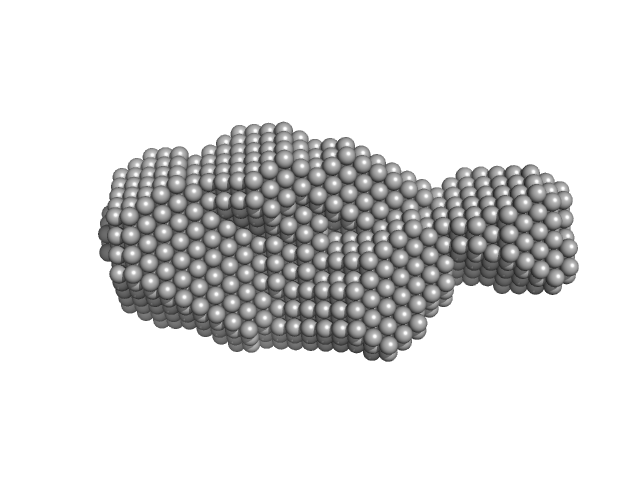

|
| Sample: |
Human Chromatin Remodeler CHD4 (363-1353) monomer, 117 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES 5% Glycerol 300 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 13
|
The PHD and chromo domains regulate the ATPase activity of the human chromatin remodeler CHD4.
J Mol Biol 422(1):3-17 (2012)
Watson AA, Mahajan P, Mertens HD, Deery MJ, Zhang W, Pham P, Du X, Bartke T, Zhang W, Edlich C, Berridge G, Chen Y, Burgess-Brown NA, Kouzarides T, Wiechens N, Owen-Hughes T, Svergun DI, Gileadi O, La...
|
| RgGuinier |
4.9 |
nm |
| Dmax |
17.4 |
nm |
|
|
|
|
|
|
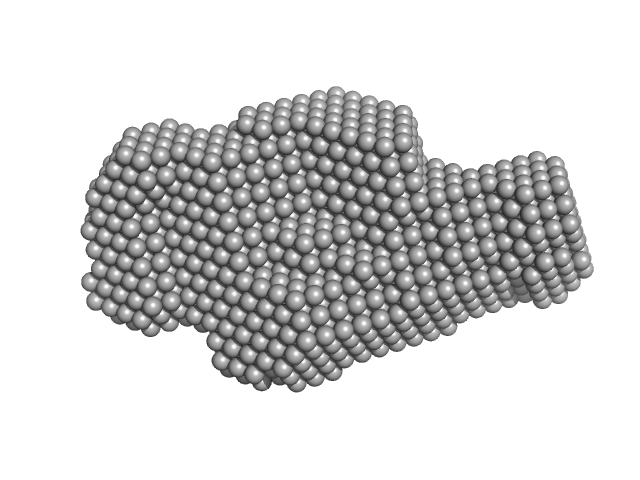

|
| Sample: |
Human Chromatin Remodeler CHD4 (494-1353) monomer, 101 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES 50 mM KCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 13
|
The PHD and chromo domains regulate the ATPase activity of the human chromatin remodeler CHD4.
J Mol Biol 422(1):3-17 (2012)
Watson AA, Mahajan P, Mertens HD, Deery MJ, Zhang W, Pham P, Du X, Bartke T, Zhang W, Edlich C, Berridge G, Chen Y, Burgess-Brown NA, Kouzarides T, Wiechens N, Owen-Hughes T, Svergun DI, Gileadi O, La...
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.5 |
nm |
|
|
|
|
|
|
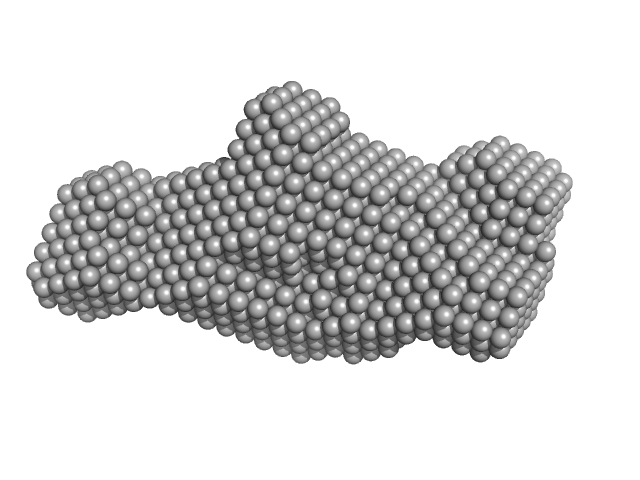

|
| Sample: |
Human Chromatin Remodeler CHD4 (685-1233) monomer, 63 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES 50 mM KCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 13
|
The PHD and chromo domains regulate the ATPase activity of the human chromatin remodeler CHD4.
J Mol Biol 422(1):3-17 (2012)
Watson AA, Mahajan P, Mertens HD, Deery MJ, Zhang W, Pham P, Du X, Bartke T, Zhang W, Edlich C, Berridge G, Chen Y, Burgess-Brown NA, Kouzarides T, Wiechens N, Owen-Hughes T, Svergun DI, Gileadi O, La...
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.5 |
nm |
|
|
|
|
|
|
|
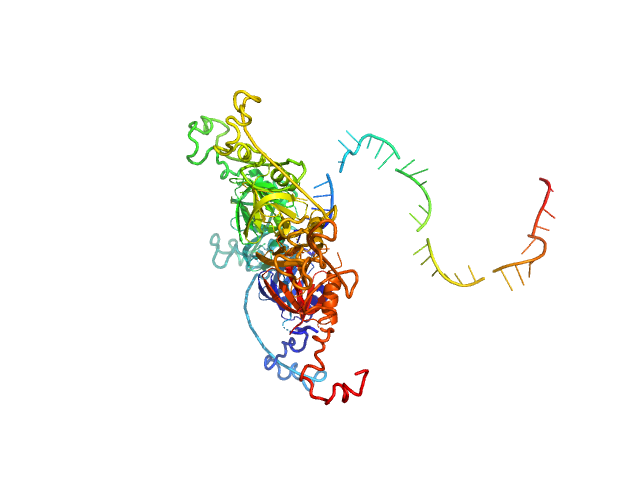
| Sample: |
RNA chaperone Hfq hexamer, 67 kDa Escherichia coli protein
RNA DsrA monomer, 12 kDa RNA
|
| Buffer: |
50 mM Tris-HCL 150 mM NaCl 1.0 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 16
|
Structural flexibility of RNA as molecular basis for Hfq chaperone function.
Nucleic Acids Res 40(16):8072-84 (2012)
Ribeiro Ede A Jr, Beich-Frandsen M, Konarev PV, Shang W, Vecerek B, Kontaxis G, Hämmerle H, Peterlik H, Svergun DI, Bläsi U, Djinović-Carugo K
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
210 |
nm3 |
|
|
|
|
|
|

|
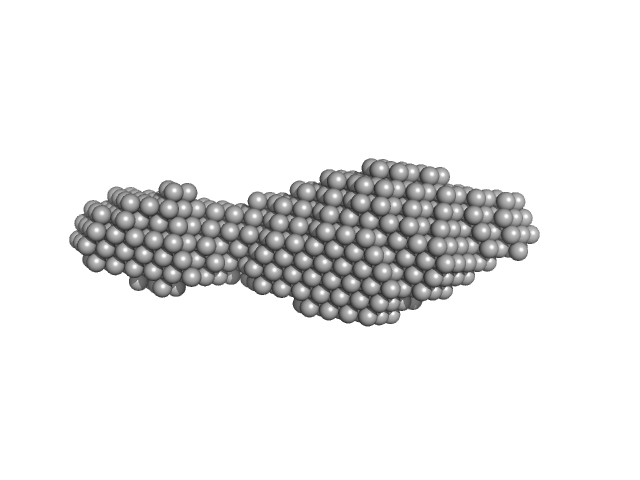
| Sample: |
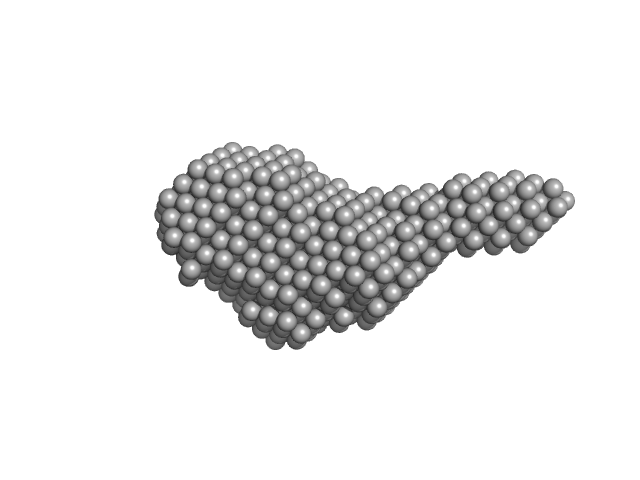
N-(2-hydroxypropyl)- 31 methacrylamide (HPMA) copolymers with cholesterol 1.4% 0, 16272 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Mar 27
|
Macromolecular HPMA-based nanoparticles with cholesterol for solid-tumor targeting: detailed study of the inner structure of a highly efficient drug delivery system.
Biomacromolecules 13(8):2594-604 (2012)
Filippov SK, Chytil P, Konarev PV, Dyakonova M, Papadakis C, Zhigunov A, Plestil J, Stepanek P, Etrych T, Ulbrich K, Svergun DI
|
| RgGuinier |
6.2 |
nm |
| Dmax |
22.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
N-(2-hydroxypropyl)- 31 methacrylamide (HPMA) copolymers with Cholesterol (2.7%) 0, 16740 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 7.2), pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Mar 27
|
Macromolecular HPMA-based nanoparticles with cholesterol for solid-tumor targeting: detailed study of the inner structure of a highly efficient drug delivery system.
Biomacromolecules 13(8):2594-604 (2012)
Filippov SK, Chytil P, Konarev PV, Dyakonova M, Papadakis C, Zhigunov A, Plestil J, Stepanek P, Etrych T, Ulbrich K, Svergun DI
|
| RgGuinier |
5.2 |
nm |
| Dmax |
28.1 |
nm |
|
|
|
|
|
|

|
| Sample: |
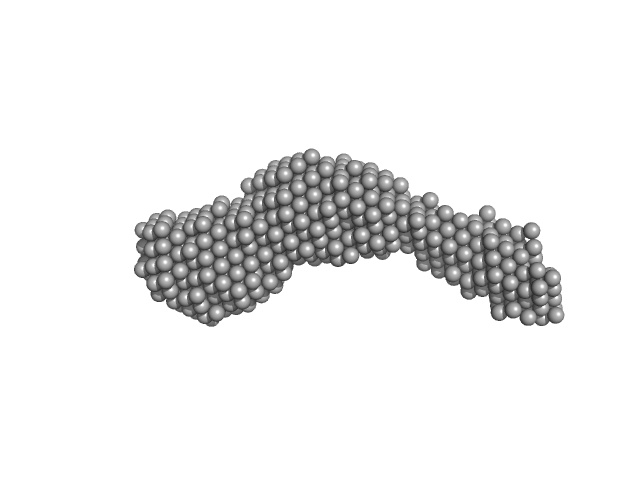
N-(2-hydroxypropyl)- 31 methacrylamide (HPMA) copolymers with Cholesterol (3.0%) 0, 29520 kDa
|
| Buffer: |
phosphate buffer saline (PBS) (pH 5.0), pH: 5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Mar 27
|
Macromolecular HPMA-based nanoparticles with cholesterol for solid-tumor targeting: detailed study of the inner structure of a highly efficient drug delivery system.
Biomacromolecules 13(8):2594-604 (2012)
Filippov SK, Chytil P, Konarev PV, Dyakonova M, Papadakis C, Zhigunov A, Plestil J, Stepanek P, Etrych T, Ulbrich K, Svergun DI
|
| RgGuinier |
9.4 |
nm |
| Dmax |
43.2 |
nm |
|
|