UniProt ID: Q8EG35 (35-333) Extracelllular iron oxide respiratory system periplasmic decaheme cytochrome c component MtrA
UniProt ID: Q8CVD4 (22-697) Extracellular iron oxide respiratory system outer membrane component MtrB
UniProt ID: Q8EG34 (22-671) Extracellular iron oxide respiratory system surface decaheme cytochrome c component MtrC
|
|
|

|
| Sample: |
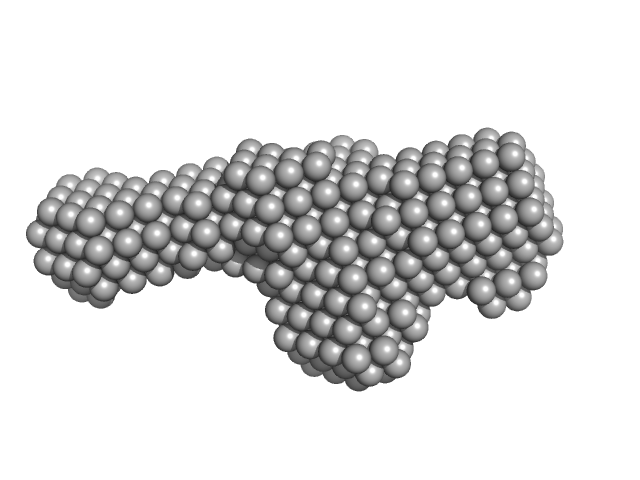
Extracelllular iron oxide respiratory system periplasmic decaheme cytochrome c component MtrA monomer, 39 kDa Shewanella oneidensis (strain … protein
Extracellular iron oxide respiratory system outer membrane component MtrB monomer, 75 kDa Shewanella oneidensis (strain … protein
Extracellular iron oxide respiratory system surface decaheme cytochrome c component MtrC monomer, 75 kDa Shewanella oneidensis (strain … protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 2.8 mM Fos-choline 12, 13% D2O, pH: 7.8 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Sep 12
|
Bespoke Biomolecular Wires for Transmembrane Electron Transfer: Spontaneous Assembly of a Functionalized Multiheme Electron Conduit
Frontiers in Microbiology 12 (2021)
Piper S, Edwards M, van Wonderen J, Casadevall C, Martel A, Jeuken L, Reisner E, Clarke T, Butt J
|
| RgGuinier |
5.2 |
nm |
| Dmax |
17.1 |
nm |
| VolumePorod |
234 |
nm3 |
|
|
UniProt ID: Q15113 (26-449) Procollagen C-endopeptidase enhancer 1
|
|
|

|
| Sample: |
Procollagen C-endopeptidase enhancer 1 monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
20 mM Hepes, 500 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2002 Jul 9
|
Low Resolution Structure Determination Shows Procollagen C-Proteinase Enhancer to be an Elongated Multidomain Glycoprotein
Journal of Biological Chemistry 278(9):7199-7205 (2003)
Bernocco S, Steiglitz B, Svergun D, Petoukhov M, Ruggiero F, Ricard-Blum S, Ebel C, Geourjon C, Deléage G, Font B, Eichenberger D, Greenspan D, Hulmes D
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
UniProt ID: A8FMV9 (1-718) Polyribonucleotide nucleotidyltransferase
|
|
|

|
| Sample: |
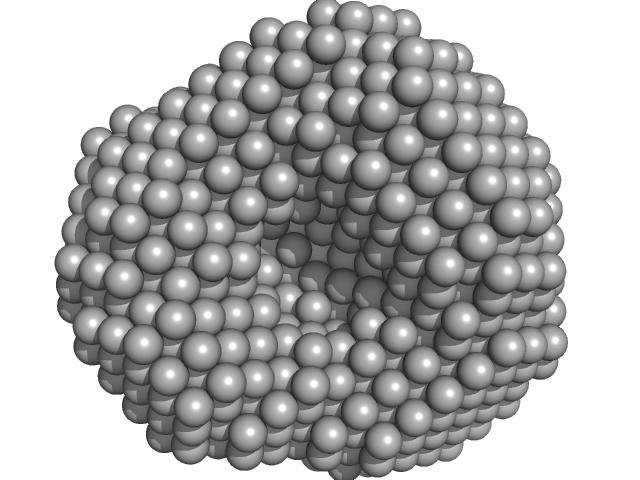
Polyribonucleotide nucleotidyltransferase trimer, 246 kDa Campylobacter jejuni subsp. … protein
|
| Buffer: |
20 mM Tris-HCl, 10 mM NAH2PO4, 60 mM KCl, 1 mM MgCl2, 2 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Mar 17
|
Structure and function of Campylobacter jejuni polynucleotide phosphorylase (PNPase): Insights into the role of this RNase in pathogenicity.
Biochimie (2023)
Bárria C, Athayde D, Hernandez G, Fonseca L, Casinhas J, Cordeiro TN, Archer M, Arraiano CM, Brito JA, Matos RG
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
310 |
nm3 |
|
|
UniProt ID: A8FMV9 (1-718) Polyribonucleotide nucleotidyltransferase
|
|
|

|
| Sample: |
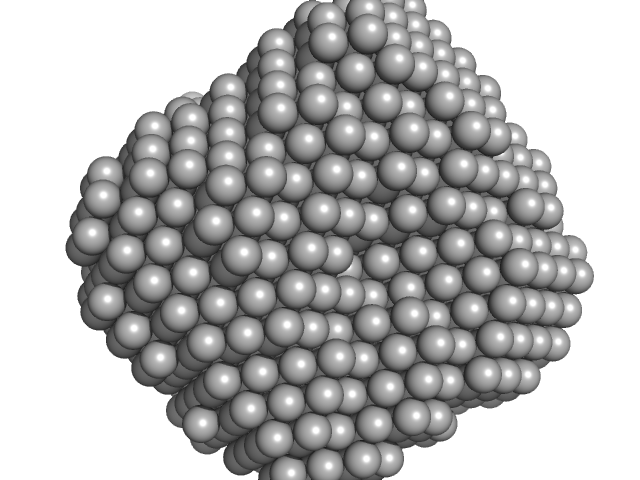
Polyribonucleotide nucleotidyltransferase trimer, 237 kDa Campylobacter jejuni subsp. … protein
|
| Buffer: |
20 mM Tris.HCl, 10 mM NAH2PO4, 60 mM KCl, 1 mM MgCl2, 2 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Mar 17
|
Structure and function of Campylobacter jejuni polynucleotide phosphorylase (PNPase): Insights into the role of this RNase in pathogenicity.
Biochimie (2023)
Bárria C, Athayde D, Hernandez G, Fonseca L, Casinhas J, Cordeiro TN, Archer M, Arraiano CM, Brito JA, Matos RG
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
315 |
nm3 |
|
|
UniProt ID: A8FMV9 (1-718) Polyribonucleotide nucleotidyltransferase
|
|
|

|
| Sample: |
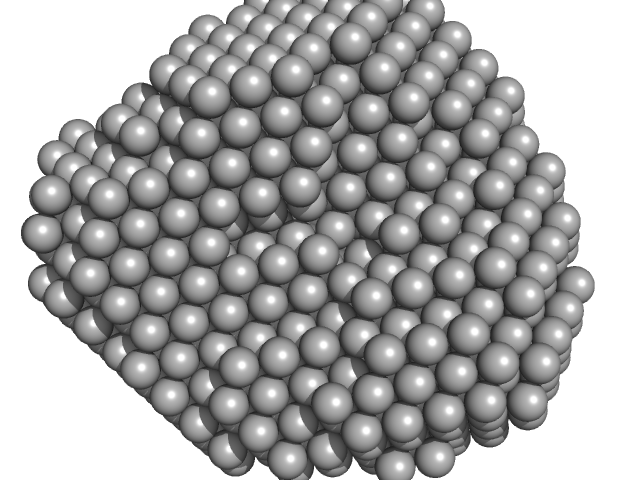
Polyribonucleotide nucleotidyltransferase trimer, 237 kDa Campylobacter jejuni subsp. … protein
|
| Buffer: |
20 mM Tris.HCl, 10 mM NAH2PO4, 60 mM KCl, 1mM MgCl2, 2 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Mar 17
|
Structure and function of Campylobacter jejuni polynucleotide phosphorylase (PNPase): Insights into the role of this RNase in pathogenicity.
Biochimie (2023)
Bárria C, Athayde D, Hernandez G, Fonseca L, Casinhas J, Cordeiro TN, Archer M, Arraiano CM, Brito JA, Matos RG
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
312 |
nm3 |
|
|
UniProt ID: P0DN75 (2-132) Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Nov 30
|
Pigeon iron-sulfur cluster assembly 1 homolog (clISCA1) 29.3 mg/ml
Shigeki Arai
|
| RgGuinier |
4.5 |
nm |
| Dmax |
15.8 |
nm |
| VolumePorod |
130 |
nm3 |
|
|
UniProt ID: Q9HBX9 (23-94) Relaxin receptor 1
|
|
|
|
| Sample: |
Relaxin receptor 1 monomer, 8 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10 mM CaCl2, 0.1%NaN3, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 6
|
Structural Insights into the Unique Modes of Relaxin-Binding and Tethered-Agonist Mediated Activation of RXFP1 and RXFP2.
J Mol Biol 433(21):167217 (2021)
Sethi A, Bruell S, Ryan T, Yan F, Tanipour MH, Mok YF, Draper-Joyce C, Khandokar Y, Metcalfe RD, Griffin MDW, Scott DJ, Hossain MA, Petrie EJ, Bathgate RAD, Gooley PR
|
| RgGuinier |
2.1 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
UniProt ID: Q8WXD0 (38-105) Relaxin receptor 2
|
|
|
|
| Sample: |
Relaxin receptor 2 monomer, 7 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 10 mM CaCl2, 0.1%NaN3, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 6
|
Structural Insights into the Unique Modes of Relaxin-Binding and Tethered-Agonist Mediated Activation of RXFP1 and RXFP2.
J Mol Biol 433(21):167217 (2021)
Sethi A, Bruell S, Ryan T, Yan F, Tanipour MH, Mok YF, Draper-Joyce C, Khandokar Y, Metcalfe RD, Griffin MDW, Scott DJ, Hossain MA, Petrie EJ, Bathgate RAD, Gooley PR
|
| RgGuinier |
1.7 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
12 |
nm3 |
|
|
UniProt ID: Q14721 (29-147) Potassium voltage-gated channel subfamily B member 1
|
|
|
|
| Sample: |
Potassium voltage-gated channel subfamily B member 1 pentamer, 73 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 5% v/v glycerol, 0.01% Triton X-100, 10 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Jul 13
|
Pentameric assembly of the Kv2.1 tetramerization domain
Acta Crystallographica Section D Structural Biology 78(6):792-802 (2022)
Xu Z, Khan S, Schnicker N, Baker S
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
140 |
nm3 |
|
|
UniProt ID: E6YFW2 (2-558) Bartonella effector protein (Bep) substrate of VirB T4SS
|
|
|
|
| Sample: |
Bartonella effector protein (Bep) substrate of VirB T4SS monomer, 64 kDa Bartonella clarridgeiae (strain … protein
|
| Buffer: |
25 mM Hepes, 300 mM NaCl, 1 mM TCEP, 5% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at Xenocs Xeuss 2.0 Q-Xoom, Center for Structural Studies, Heinrich-Heine-University on 2020 May 29
|
Structure and function of Bartonella effector protein 1: target and interdomain interactions
University of Basel PhD thesis 15051 (2023)
Markus Huber, Jens Reiners
|
| RgGuinier |
4.1 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
83 |
nm3 |
|
|