UniProt ID: None (None-None) Glyco_trans_2-like domain-containing protein
|
|
|
|
| Sample: |
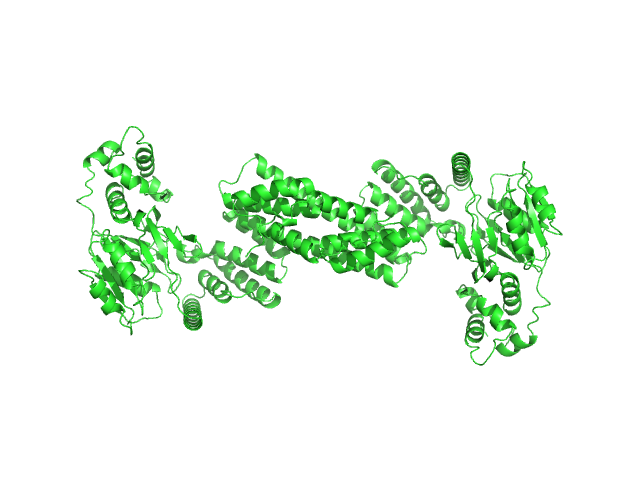
Glyco_trans_2-like domain-containing protein dimer, 96 kDa Streptomyces sp. M41(2017) protein
|
| Buffer: |
20 mM Tris, pH 7.5, 100 mM NaCl, 2 mM DTT, 1 mM EDTA, 2 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2019 Oct 18
|
Global shape of SvGT, a metal-dependent bacteriocin modifying S/O-HexNActransferase from actinobacteria: c -terminal dimerization modulates the function of this GT.
J Biomol Struct Dyn :1-15 (2023)
Sharma Y, Ahlawat S, Ashish, Rao A
|
| RgGuinier |
5.8 |
nm |
| Dmax |
17.9 |
nm |
| VolumePorod |
111 |
nm3 |
|
|
UniProt ID: P04156 (23-231) Major prion protein
|
|
|
|
| Sample: |
Major prion protein monomer, 23 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 100 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Dec 13
|
A tetracationic porphyrin with dual anti-prion activity.
iScience 26(9):107480 (2023)
Masone A, Zucchelli C, Caruso E, Lavigna G, Eraña H, Giachin G, Tapella L, Comerio L, Restelli E, Raimondi I, Elezgarai SR, De Leo F, Quilici G, Taiarol L, Oldrati M, Lorenzo NL, García-Martínez S, Cagnotto A, Lucchetti J, Gobbi M, Vanni I, Nonno R, Di Bari MA, Tully MD, Cecatiello V, Ciossani G, Pasqualato S, Van Anken E, Salmona M, Castilla J, Requena JR, Banfi S, Musco G, Chiesa R
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.2 |
nm |
|
|
UniProt ID: P04156 (23-231) Major prion protein
|
|
|
|
| Sample: |
Major prion protein monomer, 23 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 100 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Dec 13
|
A tetracationic porphyrin with dual anti-prion activity.
iScience 26(9):107480 (2023)
Masone A, Zucchelli C, Caruso E, Lavigna G, Eraña H, Giachin G, Tapella L, Comerio L, Restelli E, Raimondi I, Elezgarai SR, De Leo F, Quilici G, Taiarol L, Oldrati M, Lorenzo NL, García-Martínez S, Cagnotto A, Lucchetti J, Gobbi M, Vanni I, Nonno R, Di Bari MA, Tully MD, Cecatiello V, Ciossani G, Pasqualato S, Van Anken E, Salmona M, Castilla J, Requena JR, Banfi S, Musco G, Chiesa R
|
| RgGuinier |
3.1 |
nm |
| Dmax |
11.3 |
nm |
|
|
UniProt ID: P04156 (23-231) Major prion protein
|
|
|
|
| Sample: |
Major prion protein monomer, 23 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 100 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Dec 13
|
A tetracationic porphyrin with dual anti-prion activity.
iScience 26(9):107480 (2023)
Masone A, Zucchelli C, Caruso E, Lavigna G, Eraña H, Giachin G, Tapella L, Comerio L, Restelli E, Raimondi I, Elezgarai SR, De Leo F, Quilici G, Taiarol L, Oldrati M, Lorenzo NL, García-Martínez S, Cagnotto A, Lucchetti J, Gobbi M, Vanni I, Nonno R, Di Bari MA, Tully MD, Cecatiello V, Ciossani G, Pasqualato S, Van Anken E, Salmona M, Castilla J, Requena JR, Banfi S, Musco G, Chiesa R
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.3 |
nm |
|
|
UniProt ID: O95461 (34-756) Xylosyl- and glucuronyltransferase LARGE1
|
|
|

|
| Sample: |
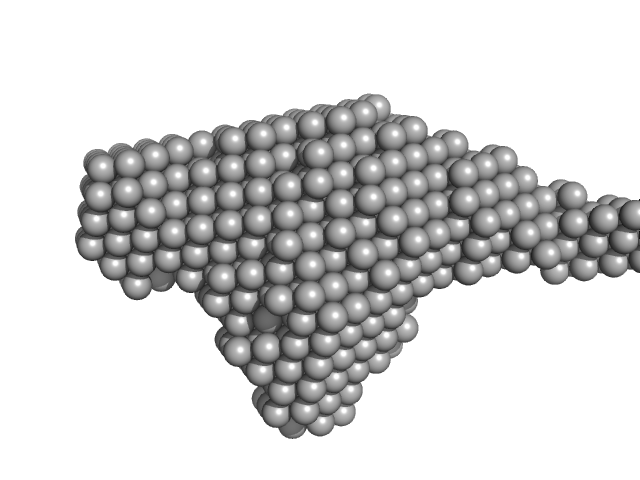
Xylosyl- and glucuronyltransferase LARGE1 dimer, 179 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Mar 18
|
LARGE1 processively polymerizes length-controlled matriglycan on prodystroglycan.
Nat Commun 16(1):9028 (2025)
Joseph S, Schnicker NJ, Spellmon N, Xu Z, Yan R, Yu Z, Davulcu O, Yang T, Hopkins J, Anderson ME, Venzke D, Campbell KP
|
| RgGuinier |
4.4 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
264 |
nm3 |
|
|
UniProt ID: O95461 (34-756) Xylosyl- and glucuronyltransferase LARGE1
|
|
|

|
| Sample: |
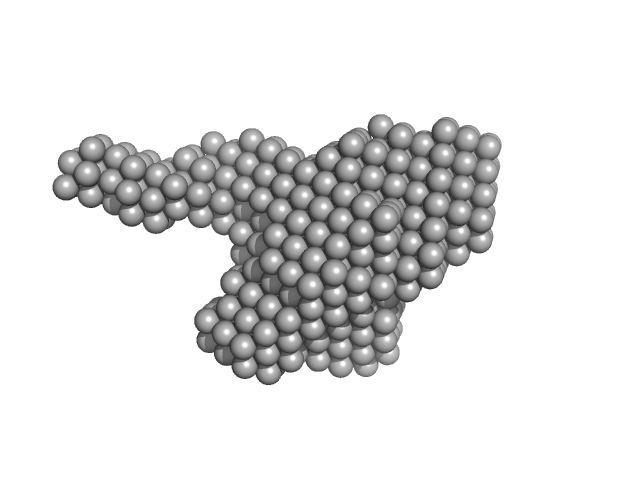
Xylosyl- and glucuronyltransferase LARGE1 dimer, 179 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Mar 18
|
LARGE1 processively polymerizes length-controlled matriglycan on prodystroglycan.
Nat Commun 16(1):9028 (2025)
Joseph S, Schnicker NJ, Spellmon N, Xu Z, Yan R, Yu Z, Davulcu O, Yang T, Hopkins J, Anderson ME, Venzke D, Campbell KP
|
| RgGuinier |
4.3 |
nm |
| Dmax |
18.0 |
nm |
|
|
UniProt ID: Q5XPT3 (30-690) Xylosyl- and glucuronyltransferase LARGE2
|
|
|

|
| Sample: |
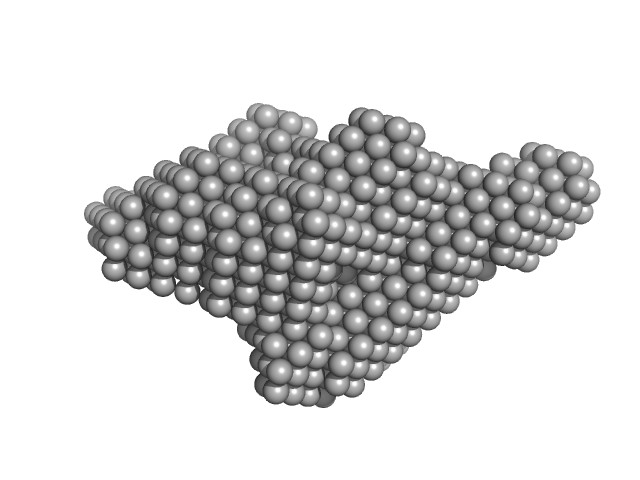
Xylosyl- and glucuronyltransferase LARGE2 dimer, 168 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Mar 18
|
LARGE1 processively polymerizes length-controlled matriglycan on prodystroglycan.
Nat Commun 16(1):9028 (2025)
Joseph S, Schnicker NJ, Spellmon N, Xu Z, Yan R, Yu Z, Davulcu O, Yang T, Hopkins J, Anderson ME, Venzke D, Campbell KP
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.5 |
nm |
|
|
UniProt ID: Q5XPT3 (30-690) Xylosyl- and glucuronyltransferase LARGE2
|
|
|
|
| Sample: |
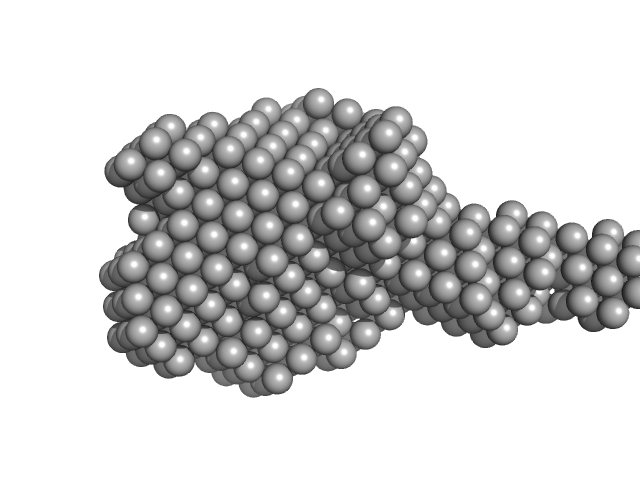
Xylosyl- and glucuronyltransferase LARGE2 dimer, 168 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Mar 18
|
LARGE1 processively polymerizes length-controlled matriglycan on prodystroglycan.
Nat Commun 16(1):9028 (2025)
Joseph S, Schnicker NJ, Spellmon N, Xu Z, Yan R, Yu Z, Davulcu O, Yang T, Hopkins J, Anderson ME, Venzke D, Campbell KP
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
231 |
nm3 |
|
|
UniProt ID: Q8KGS3 (2-83) DUF2285 domain-containing protein
|
|
|
|
| Sample: |
DUF2285 domain-containing protein monomer, 11 kDa Mesorhizobium loti protein
|
| Buffer: |
100 mM NaCl, 50 mM Tris-HCl, 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Dec 1
|
DUF2285 is a novel helix-turn-helix domain variant that orchestrates both activation and antiactivation of conjugative element transfer in proteobacteria.
Nucleic Acids Res (2023)
Jowsey WJ, Morris CRP, Hall DA, Sullivan JT, Fagerlund RD, Eto KY, Solomon PD, Mackay JP, Bond CS, Ramsay JP, Ronson CW
|
| RgGuinier |
1.5 |
nm |
| Dmax |
5.7 |
nm |
| VolumePorod |
15 |
nm3 |
|
|
UniProt ID: Q9VSK3 (757-991) Upstream of N-ras, isoform A
UniProt ID: None (None-None) polyA-15mer
|
|
|
|
| Sample: |
Upstream of N-ras, isoform A monomer, 26 kDa Drosophila melanogaster protein
PolyA-15mer monomer, 5 kDa RNA
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 8
|
Upstream of N-Ras C-terminal cold shock domains mediate poly(A) specificity in a novel RNA recognition mode and bind poly(A) binding protein.
Nucleic Acids Res (2023)
Hollmann NM, Jagtap PKA, Linse JB, Ullmann P, Payr M, Murciano B, Simon B, Hub JS, Hennig J
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
37 |
nm3 |
|
|