UniProt ID: P0DTC2 (319-566) Spike glycoprotein (ACE2 receptor binding domain)
|
|
|
|
| Sample: |
Spike glycoprotein (ACE2 receptor binding domain) monomer, 29 kDa Severe acute respiratory … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SANS
data collected at SPB/SFX, European XFEL on 2021 May 14
|
Form factor determination of biological molecules with X-ray free electron laser small-angle scattering (XFEL-SAS).
Commun Biol 6(1):1057 (2023)
Blanchet CE, Round A, Mertens HDT, Ayyer K, Graewert M, Awel S, Franke D, Dörner K, Bajt S, Bean R, Custódio TF, de Wijn R, Juncheng E, Henkel A, Gruzinov A, Jeffries CM, Kim Y, Kirkwood H, Kloos M, Knoška J, Koliyadu J, Letrun R, Löw C, Makroczyova J, Mall A, Meijers R, Pena Murillo GE, Oberthür D, Round E, Seuring C, Sikorski M, Vagovic P, Valerio J, Wollweber T, Zhuang Y, Schulz J, Haas H, Chapman HN, Mancuso AP, Svergun D
|
|
|
UniProt ID: P43351 (1-418) DNA repair protein RAD52 homolog
|
|
|
|
| Sample: |
DNA repair protein RAD52 homolog undecamer, 528 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 7.5, 250 mM NaCl, 1% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Jul 21
|
An integrative structural study of the human full-length RAD52 at 2.2 Å resolution
Communications Biology 7(1) (2024)
Balboni B, Marotta R, Rinaldi F, Milordini G, Varignani G, Girotto S, Cavalli A
|
| RgGuinier |
8.0 |
nm |
| Dmax |
40.2 |
nm |
| VolumePorod |
1218 |
nm3 |
|
|
UniProt ID: A0A6M7H0F8 (2-141) YdaT_toxin domain-containing protein
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDQB9_fit1_model1.png)
|
| Sample: |
YdaT_toxin domain-containing protein tetramer, 74 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
20 mM Tris-HCl, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 16
|
Structural basis of DNA binding by YdaT, a functional equivalent of the CII repressor in the cryptic prophage CP-933P from Escherichia coli
O157:H7
Acta Crystallographica Section D Structural Biology 79(3):245-258 (2023)
Prolič-Kalinšek M, Volkov A, Hadži S, Van Dyck J, Bervoets I, Charlier D, Loris R
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
130 |
nm3 |
|
|
UniProt ID: A0A6M7H0F8 (2-141) YdaT_toxin domain-containing protein (mutant: L111N, F118R)
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDQC9_fit1_model1.png)
|
| Sample: |
YdaT_toxin domain-containing protein (mutant: L111N, F118R) monomer, 18 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
20 mM Tris-HCl, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Apr 14
|
Structural basis of DNA binding by YdaT, a functional equivalent of the CII repressor in the cryptic prophage CP-933P from Escherichia coli
O157:H7
Acta Crystallographica Section D Structural Biology 79(3):245-258 (2023)
Prolič-Kalinšek M, Volkov A, Hadži S, Van Dyck J, Bervoets I, Charlier D, Loris R
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
UniProt ID: A0A6M7H0F8 (2-96) YdaT_toxin domain-containing protein
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDQD9_fit1_model1.png)
|
| Sample: |
YdaT_toxin domain-containing protein monomer, 13 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
20 mM Tris-HCl, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 16
|
Structural basis of DNA binding by YdaT, a functional equivalent of the CII repressor in the cryptic prophage CP-933P from Escherichia coli
O157:H7
Acta Crystallographica Section D Structural Biology 79(3):245-258 (2023)
Prolič-Kalinšek M, Volkov A, Hadži S, Van Dyck J, Bervoets I, Charlier D, Loris R
|
| RgGuinier |
1.7 |
nm |
| Dmax |
5.7 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
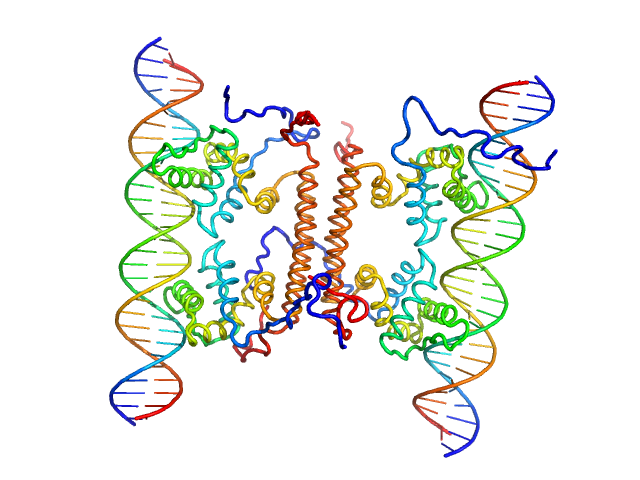
UniProt ID: A0A6M7H0F8 (2-141) YdaT_toxin domain-containing protein
UniProt ID: None (None-None) Om 30 base pair dsDNA
|
|
|
|
| Sample: |
YdaT_toxin domain-containing protein tetramer, 74 kDa Escherichia coli O157:H7 protein
Om 30 base pair dsDNA dimer, 37 kDa DNA
|
| Buffer: |
20 mM Tris-HCl, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 14
|
Structural basis of DNA binding by YdaT, a functional equivalent of the CII repressor in the cryptic prophage CP-933P from Escherichia coli
O157:H7
Acta Crystallographica Section D Structural Biology 79(3):245-258 (2023)
Prolič-Kalinšek M, Volkov A, Hadži S, Van Dyck J, Bervoets I, Charlier D, Loris R
|
| RgGuinier |
4.2 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
UniProt ID: P08113 (22-754) Endoplasmin
|
|
|
|
| Sample: |
Endoplasmin dimer, 169 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Jul 11
|
Visualization of conformational transition of GRP94 in solution.
Life Sci Alliance 7(2) (2024)
Sun S, Zhu R, Zhu M, Wang Q, Li N, Yang B
|
| RgGuinier |
5.6 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
UniProt ID: P08113 (22-754) Endoplasmin
|
|
|
|
| Sample: |
Endoplasmin dimer, 169 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Jul 11
|
Visualization of conformational transition of GRP94 in solution.
Life Sci Alliance 7(2) (2024)
Sun S, Zhu R, Zhu M, Wang Q, Li N, Yang B
|
| RgGuinier |
5.7 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
274 |
nm3 |
|
|
UniProt ID: P08113 (22-754) Endoplasmin
|
|
|
|
| Sample: |
Endoplasmin dimer, 169 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES, 200 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Jul 11
|
Visualization of conformational transition of GRP94 in solution.
Life Sci Alliance 7(2) (2024)
Sun S, Zhu R, Zhu M, Wang Q, Li N, Yang B
|
| RgGuinier |
5.5 |
nm |
| Dmax |
24.5 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
UniProt ID: P34710 (22-461) Netrin unc-6
UniProt ID: Q26261-1 (1-331) Isoform a of Netrin receptor unc-5
UniProt ID: None (None-None) Heparin, porcine intestinal mucosa
|
|
|
|
| Sample: |
Netrin unc-6 monomer, 52 kDa Caenorhabditis elegans protein
Isoform a of Netrin receptor unc-5 monomer, 38 kDa Caenorhabditis elegans protein
Heparin, porcine intestinal mucosa monomer, 15 kDa Sus scrofa domesticus
|
| Buffer: |
10 mM HEPES pH 7.2, 150 mM NaCl, 100 mM MgSO4, pH: 7.2 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Jun 18
|
Structural insights into the formation of repulsive netrin guidance complexes.
Sci Adv 10(7):eadj8083 (2024)
Priest JM, Nichols EL, Smock RG, Hopkins JB, Mendoza JL, Meijers R, Shen K, Özkan E
|
| RgGuinier |
10.4 |
nm |
| Dmax |
37.3 |
nm |
| VolumePorod |
2156 |
nm3 |
|
|