UniProt ID: Q14181 (155-598) DNA polymerase alpha subunit B
UniProt ID: P09884 (1265-1462) DNA polymerase alpha catalytic subunit
UniProt ID: P49643 (1-509) DNA primase large subunit
UniProt ID: P49642 (1-420) DNA primase small subunit
UniProt ID: None (None-None) 29 mer DNA template
UniProt ID: None (None-None) 8mer RNA primer
|
|
|
|
| Sample: |
DNA polymerase alpha subunit B monomer, 49 kDa Homo sapiens protein
DNA polymerase alpha catalytic subunit monomer, 23 kDa Homo sapiens protein
DNA primase large subunit monomer, 59 kDa Homo sapiens protein
DNA primase small subunit monomer, 50 kDa Homo sapiens protein
29 mer DNA template monomer, 9 kDa DNA
8mer RNA primer monomer, 3 kDa RNA
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM MgCl2, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Mar 28
|
Flexibility and distributive synthesis regulate RNA priming and handoff in human DNA polymerase α-primase
Journal of Molecular Biology :168330 (2023)
Cordoba J, Mullins E, Salay L, Eichman B, Chazin W
|
| RgGuinier |
4.9 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
344 |
nm3 |
|
|
UniProt ID: Q14181 (155-598) DNA polymerase alpha subunit B
UniProt ID: P09884 (1265-1462) DNA polymerase alpha catalytic subunit
UniProt ID: P49642 (1-420) DNA primase small subunit
UniProt ID: P49643 (1-265) DNA primase large subunit
|
|
|
|
| Sample: |
DNA polymerase alpha subunit B monomer, 49 kDa Homo sapiens protein
DNA polymerase alpha catalytic subunit monomer, 23 kDa Homo sapiens protein
DNA primase small subunit monomer, 50 kDa Homo sapiens protein
DNA primase large subunit monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM MgCl2, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Apr 15
|
Flexibility and distributive synthesis regulate RNA priming and handoff in human DNA polymerase α-primase
Journal of Molecular Biology :168330 (2023)
Cordoba J, Mullins E, Salay L, Eichman B, Chazin W
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
221 |
nm3 |
|
|
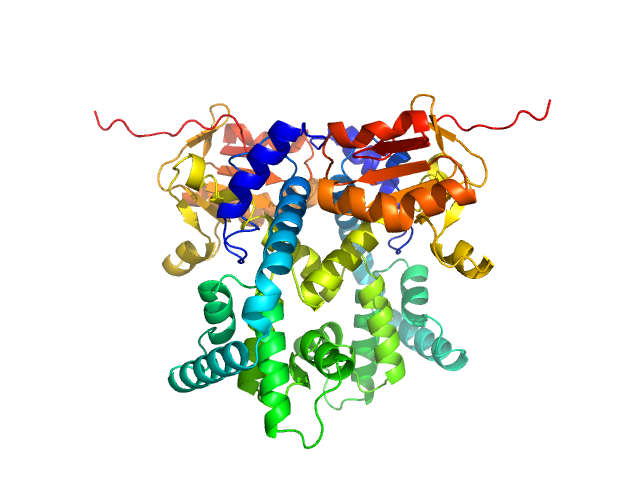
UniProt ID: Q9Y2V0 (1-281) CDAN1-interacting nuclease 1
|
|
|
|
| Sample: |
CDAN1-interacting nuclease 1 dimer, 67 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2021 Sep 2
|
Structural insights into inherited anemia CDA-I: disease-associated mutations disrupt Codanin1-CDIN1 complex
Tomas Brom
|
| RgGuinier |
3.0 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
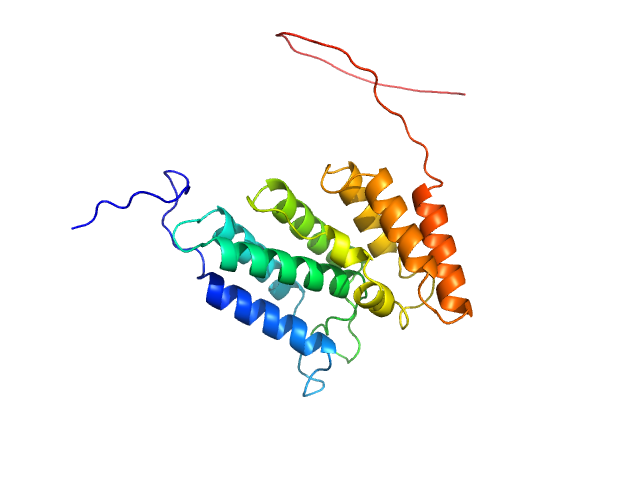
UniProt ID: Q8IWY9 (1005-1227) Codanin-1
|
|
|
|
| Sample: |
Codanin-1 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2022 Mar 5
|
Structural insights into inherited anemia CDA-I: disease-associated mutations disrupt Codanin1-CDIN1 complex
Tomas Brom
|
| RgGuinier |
2.7 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
54 |
nm3 |
|
|
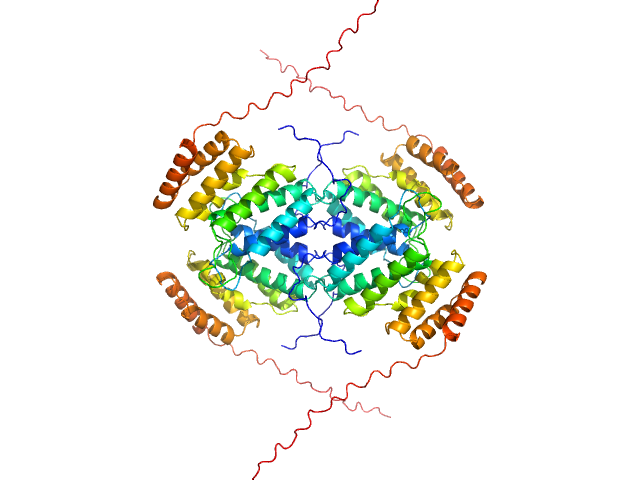
UniProt ID: Q8IWY9 (1005-1227) Codanin-1
|
|
|

|
| Sample: |
Codanin-1 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2022 Mar 5
|
Structural insights into inherited anemia CDA-I: disease-associated mutations disrupt Codanin1-CDIN1 complex
Tomas Brom
|
| RgGuinier |
4.1 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
220 |
nm3 |
|
|
UniProt ID: O75031 (2-334) Heat shock factor 2-binding protein
|
|
|
|
| Sample: |
Heat shock factor 2-binding protein tetramer, 150 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl pH 7.5, 250 mM NaCl, and 5 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2020 Oct 2
|
BRCA2-HSF2BP oligomeric ring disassembly by BRME1 promotes homologous recombination.
Sci Adv 9(43):eadi7352 (2023)
Ghouil R, Miron S, Sato K, Ristic D, van Rossum-Fikkert SE, Legrand P, Ouldali M, Winter JM, Ropars V, David G, Arteni AA, Wyman C, Knipscheer P, Kanaar R, Zelensky AN, Zinn-Justin S
|
| RgGuinier |
8.5 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
364 |
nm3 |
|
|
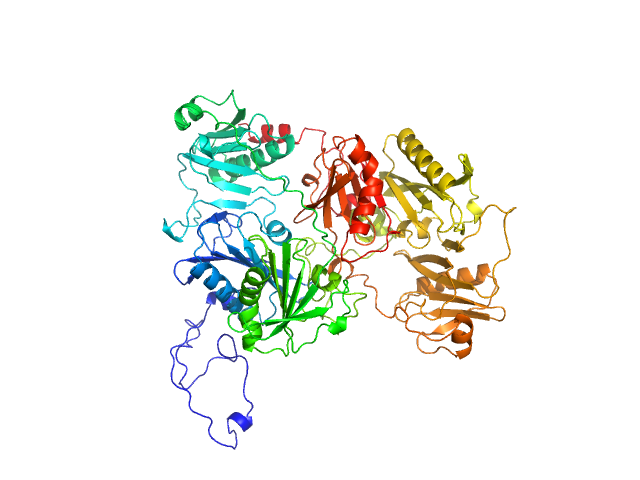
UniProt ID: P06396 (28-782) Gelsolin
|
|
|
|
| Sample: |
Gelsolin monomer, 85 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 1 mM EDTA, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Feb 24
|
Accurate and Efficient SAXS/SANS Implementation Including Solvation Layer Effects Suitable for Molecular Simulations.
J Chem Theory Comput (2023)
Ballabio F, Paissoni C, Bollati M, de Rosa M, Capelli R, Camilloni C
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
128 |
nm3 |
|
|
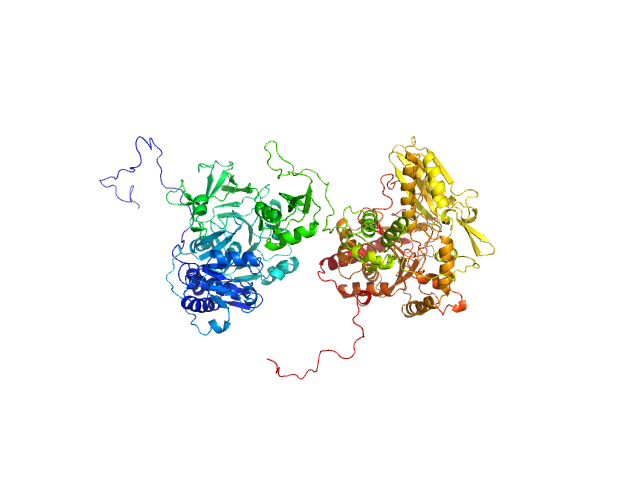
UniProt ID: P0C061 (1-1098) Gramicidin S synthase 1 (W239S)
|
|
|
|
| Sample: |
Gramicidin S synthase 1 (W239S) monomer, 127 kDa Aneurinibacillus migulanus protein
|
| Buffer: |
20 mM Tris pH 7.5, 100 mM NaCl, 10 mM MgCl2, 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Mar 14
|
Resting state of GrsA is a combination of two constitutively inactive isoforms that manifest long-range allosteric effects in presence of aminoacid and ATP
Raktim Roy
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
187 |
nm3 |
|
|
UniProt ID: P0C062 (1-1098) Gramicidin S synthase 1 (W239S)
|
|
|
|
| Sample: |
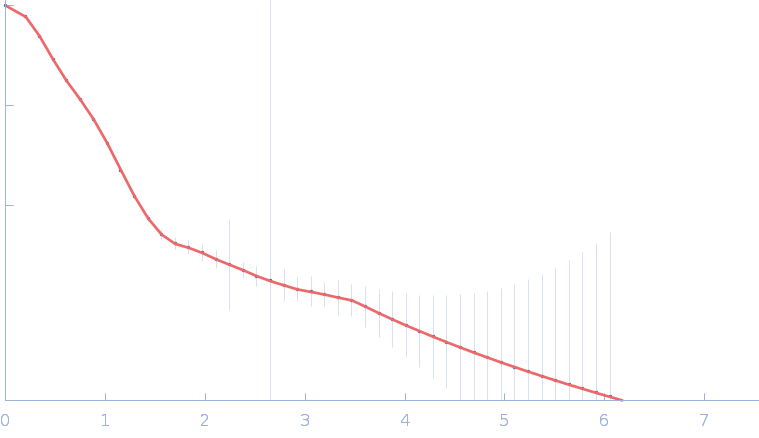
Gramicidin S synthase 1 (W239S) monomer, 126 kDa Brevibacillus brevis protein
|
| Buffer: |
20 mM Tris pH 7.5, 100 mM NaCl, 10 mM MgCl2, 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Mar 14
|
Resting state of GrsA is a combination of two constitutively inactive isoforms that manifest long-range allosteric effects in presence of aminoacid and ATP
Raktim Roy
|
| RgGuinier |
4.3 |
nm |
| Dmax |
15.4 |
nm |
| VolumePorod |
186 |
nm3 |
|
|
UniProt ID: P0C062 (1-1098) Gramicidin S synthase 1 (W239S)
|
|
|
|
| Sample: |
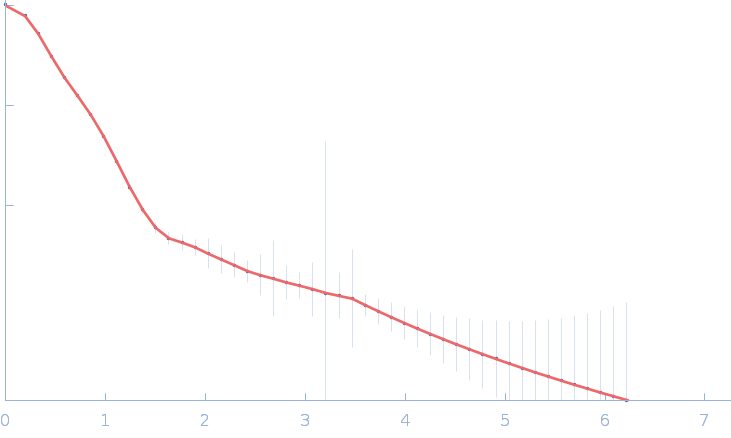
Gramicidin S synthase 1 (W239S) monomer, 126 kDa Brevibacillus brevis protein
|
| Buffer: |
20 mM Tris pH 7.5, 100 mM NaCl, 10 mM MgCl2, 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Mar 14
|
Resting state of GrsA is a combination of two constitutively inactive isoforms that manifest long-range allosteric effects in presence of aminoacid and ATP
Raktim Roy
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
186 |
nm3 |
|
|