UniProt ID: M1L6N7 (944-1066) Genome polyprotein
|
|
|
|
| Sample: |
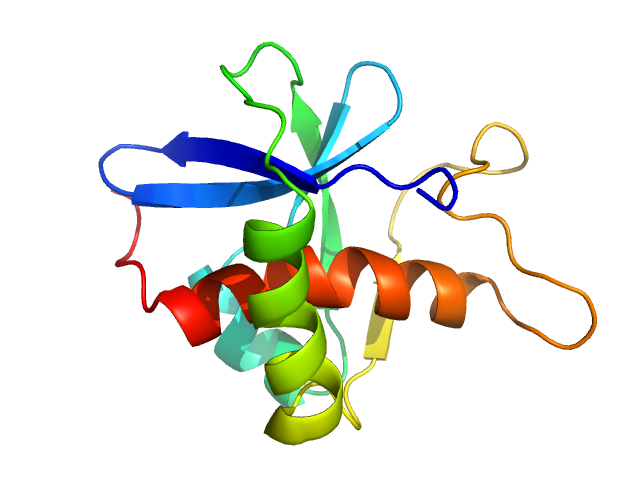
Genome polyprotein monomer, 13 kDa Turkey avisivirus protein
|
| Buffer: |
20mM Citrate buffer pH 6.5, 150mM NaCl, 2mM TCEP, 1% glycerol, pH: 6.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 May 15
|
Solution structure and functional characterization of an avian picornavirus 2AH/NC protein
Anna Wehlin
|
| RgGuinier |
1.6 |
nm |
| Dmax |
6.0 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
UniProt ID: M1L6N7 (944-1066) Genome polyprotein
|
|
|
|
| Sample: |
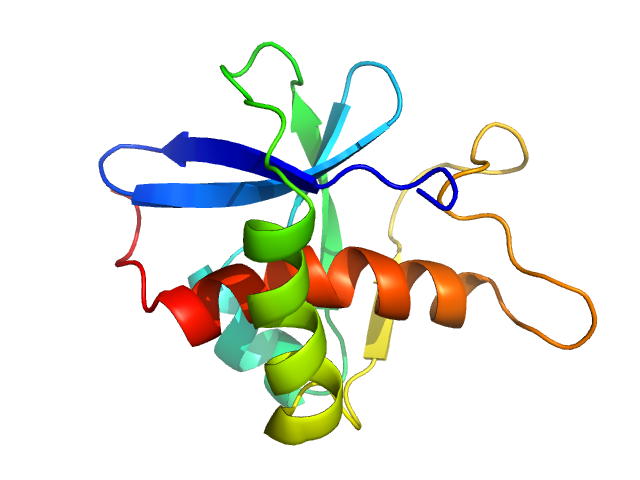
Genome polyprotein monomer, 13 kDa Turkey avisivirus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM TECP, 1% Glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 May 2
|
Solution structure and functional characterization of an avian picornavirus 2AH/NC protein
Anna Wehlin
|
| RgGuinier |
1.6 |
nm |
| Dmax |
4.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|
UniProt ID: Q9NRC8-1 (1-400) Isoform 1 of NAD-dependent protein deacetylase sirtuin-7
|
|
|
|
| Sample: |
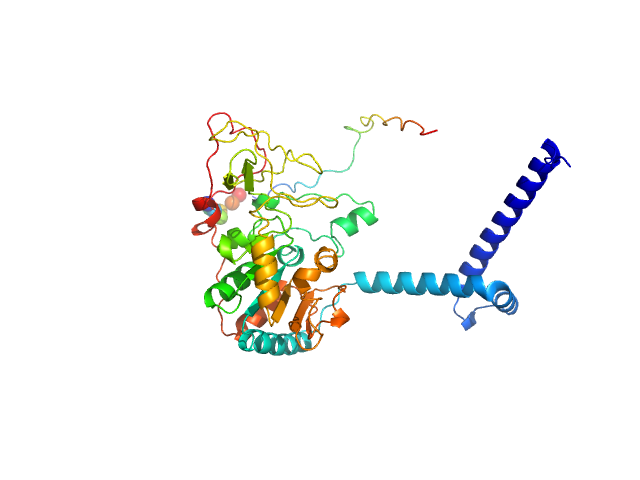
Isoform 1 of NAD-dependent protein deacetylase sirtuin-7 monomer, 45 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Dec 16
|
Substrates and Cyclic Peptide Inhibitors of the Oligonucleotide-Activated Sirtuin 7.
Angew Chem Int Ed Engl :e202314597 (2023)
Bolding JE, Nielsen AL, Jensen I, Hansen TN, Ryberg LA, Jameson ST, Harris P, Peters GHJ, Denu JM, Rogers JM, Olsen CA
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
89 |
nm3 |
|
|
UniProt ID: P30153 (1-589) Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform
UniProt ID: Q14738-1 (1-602) Isoform Delta-1 of Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (R160H)
UniProt ID: P67775-1 (1-309) Isoform 1 of Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform
|
|
|
|
| Sample: |
Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform monomer, 65 kDa Homo sapiens protein
Isoform Delta-1 of Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (R160H) monomer, 70 kDa Homo sapiens protein
Isoform 1 of Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform monomer, 36 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, 50 µM MnCl2, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Nov 20
|
B56δ long-disordered arms form a dynamic PP2A regulation interface coupled with global allostery and Jordan's syndrome mutations.
Proc Natl Acad Sci U S A 121(1):e2310727120 (2024)
Wu CG, Balakrishnan VK, Merrill RA, Parihar PS, Konovolov K, Chen YC, Xu Z, Wei H, Sundaresan R, Cui Q, Wadzinski BE, Swingle MR, Musiyenko A, Chung WK, Honkanen RE, Suzuki A, Huang X, Strack S, Xing Y
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
266 |
nm3 |
|
|
UniProt ID: P30153 (1-589) Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform
UniProt ID: P67775-1 (1-309) Isoform 1 of Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform
UniProt ID: Q14738-1 (1-602) Isoform Delta-1 of Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (R160H, E198K)
|
|
|
|
| Sample: |
Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform monomer, 65 kDa Homo sapiens protein
Isoform 1 of Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform monomer, 36 kDa Homo sapiens protein
Isoform Delta-1 of Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (R160H, E198K) monomer, 70 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, 50 µM MnCl2, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Nov 20
|
B56δ long-disordered arms form a dynamic PP2A regulation interface coupled with global allostery and Jordan's syndrome mutations.
Proc Natl Acad Sci U S A 121(1):e2310727120 (2024)
Wu CG, Balakrishnan VK, Merrill RA, Parihar PS, Konovolov K, Chen YC, Xu Z, Wei H, Sundaresan R, Cui Q, Wadzinski BE, Swingle MR, Musiyenko A, Chung WK, Honkanen RE, Suzuki A, Huang X, Strack S, Xing Y
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
305 |
nm3 |
|
|
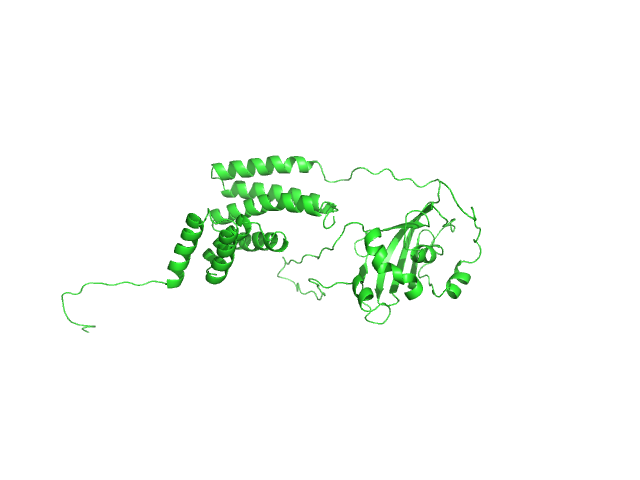
UniProt ID: O00170 (2-330) AH receptor-interacting protein (R9Q)
|
|
|

|
| Sample: |
AH receptor-interacting protein (R9Q) monomer, 39 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-Cl pH 7.5, 100 mM NaCl, 1 mM DTT, 5% (v/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 Jun 13
|
Human AIP HSP90 complex
Robert Rambo
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.9 |
nm |
| VolumePorod |
64 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
4.2 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
148 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
3.8 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
3.7 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
124 |
nm3 |
|
|
UniProt ID: P21980 (1-687) Protein-glutamine gamma-glutamyltransferase 2
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2022 Nov 17
|
Distinct conformational states enable transglutaminase 2 to promote cancer cell survival versus cell death.
Commun Biol 7(1):982 (2024)
Aplin C, Zielinski KA, Pabit S, Ogunribido D, Katt WP, Pollack L, Cerione RA, Milano SK
|
| RgGuinier |
3.6 |
nm |
| Dmax |
18.0 |
nm |
| VolumePorod |
121 |
nm3 |
|
|