UniProt ID: Q43349 (176-254) 29 kDa ribonucleoprotein, chloroplastic
|
|
|
|
| Sample: |
29 kDa ribonucleoprotein, chloroplastic monomer, 8 kDa Arabidopsis thaliana protein
|
| Buffer: |
20 mM sodium phosphate, 30 mM NaCl, pH: 6.2 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, SFB 1035, Technische Universität München on 2021 Oct 19
|
A prion-like domain is required for phase separation and chloroplast RNA processing during cold acclimation in Arabidopsis.
Plant Cell (2024)
Legen J, Lenzen B, Kachariya N, Feltgen S, Gao Y, Mergenthal S, Weber W, Klotzsch E, Zoschke R, Sattler M, Schmitz-Linneweber C
|
| RgGuinier |
2.4 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
UniProt ID: P02925 (25-296) Ribose import binding protein RbsB
|
|
|
|
| Sample: |
Ribose import binding protein RbsB monomer, 30 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 10% glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2019 Dec 9
|
SAXS/WAXS data of conformationally flexible ribose binding protein
Data in Brief :109932 (2023)
Choudhury J, Yonezawa K, Anu A, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.5 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
UniProt ID: P02925 (25-296) Ribose import binding protein RbsB
UniProt ID: None (None-None) Ribose
|
|
|
|
| Sample: |
Ribose import binding protein RbsB monomer, 30 kDa Escherichia coli (strain … protein
Ribose monomer, 0 kDa
|
| Buffer: |
50 mM Tris, 50 mM NaCl, 10% glycerol, 1 mM ribose, pH: 7 |
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2019 Dec 9
|
SAXS/WAXS data of conformationally flexible ribose binding protein
Data in Brief :109932 (2023)
Choudhury J, Yonezawa K, Anu A, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
UniProt ID: P02925 (25-296) Ribose import binding protein RbsB
UniProt ID: None (None-None) Ribose
|
|
|
|
| Sample: |
Ribose import binding protein RbsB monomer, 30 kDa Escherichia coli (strain … protein
Ribose monomer, 0 kDa
|
| Buffer: |
Tris 50 mM, 50 mM NaCl, 10% glycerol, 5 mM ribose, pH: 7 |
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2019 Dec 9
|
SAXS/WAXS data of conformationally flexible ribose binding protein
Data in Brief :109932 (2023)
Choudhury J, Yonezawa K, Anu A, Shimizu N, Chaudhuri B
|
| RgGuinier |
1.9 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
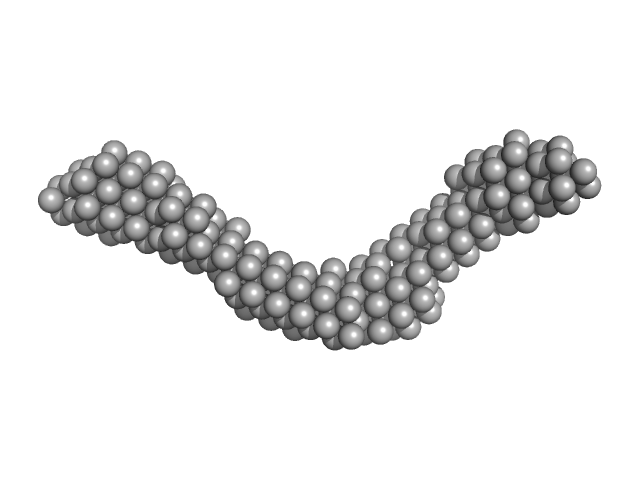
UniProt ID: Q9NYQ6 (557-995) Cadherin EGF LAG seven-pass G-type receptor 1
|
|
|
|
| Sample: |
Cadherin EGF LAG seven-pass G-type receptor 1 monomer, 49 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM KCl, 50 mM NaCl, 2 mM CaCl2, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Jul 1
|
CELSR1, a core planar cell polarity protein, features a weakly adhesive and flexible cadherin ectodomain.
Structure (2024)
Tamilselvan E, Sotomayor M
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.9 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
UniProt ID: A0A0A7LTX5 (48-342) Biofilm regulatory protein
|
|
|
|
| Sample: |
Biofilm regulatory protein monomer, 34 kDa Streptococcus dysgalactiae subsp. … protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 5 mM MgCl2, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Jun 1
|
Structural insights of an LCP protein-LytR-from Streptococcus dysgalactiae subs. dysgalactiae through biophysical and in silico methods.
Front Chem 12:1379914 (2024)
Paquete-Ferreira J, Freire F, Fernandes HS, Muthukumaran J, Ramos J, Bryton J, Panjkovich A, Svergun D, Santos MFA, Correia MAS, Fernandes AR, Romão MJ, Sousa SF, Santos-Silva T
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
83 |
nm3 |
|
|
UniProt ID: A0A0A7LTX5 (48-342) Biofilm regulatory protein
|
|
|
|
| Sample: |
Biofilm regulatory protein monomer, 34 kDa Streptococcus dysgalactiae subsp. … protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 5 mM MgCl2, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Jun 1
|
Structural insights of an LCP protein-LytR-from Streptococcus dysgalactiae subs. dysgalactiae through biophysical and in silico methods.
Front Chem 12:1379914 (2024)
Paquete-Ferreira J, Freire F, Fernandes HS, Muthukumaran J, Ramos J, Bryton J, Panjkovich A, Svergun D, Santos MFA, Correia MAS, Fernandes AR, Romão MJ, Sousa SF, Santos-Silva T
|
| RgGuinier |
2.4 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: P09936 (1-223) Ubiquitin carboxyl-terminal hydrolase isozyme L1
|
|
|
|
| Sample: |
Ubiquitin carboxyl-terminal hydrolase isozyme L1 monomer, 26 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.03% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 Sep 2
|
Altered protein dynamics and a more reactive catalytic cysteine in a neurodegeneration-associated UCHL1 mutant.
J Mol Biol :168438 (2024)
Kenny S, Lai CH, Chiang TS, Brown K, Hewitt CS, Krabill AD, Chang HT, Wang YS, Flaherty DP, Danny Hsu ST, Das C
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.9 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
UniProt ID: P09936 (1-223) Ubiquitin carboxyl-terminal hydrolase isozyme L1 (R178Q)
|
|
|
|
| Sample: |
Ubiquitin carboxyl-terminal hydrolase isozyme L1 (R178Q) monomer, 26 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.03% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 Sep 2
|
Altered protein dynamics and a more reactive catalytic cysteine in a neurodegeneration-associated UCHL1 mutant.
J Mol Biol :168438 (2024)
Kenny S, Lai CH, Chiang TS, Brown K, Hewitt CS, Krabill AD, Chang HT, Wang YS, Flaherty DP, Danny Hsu ST, Das C
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
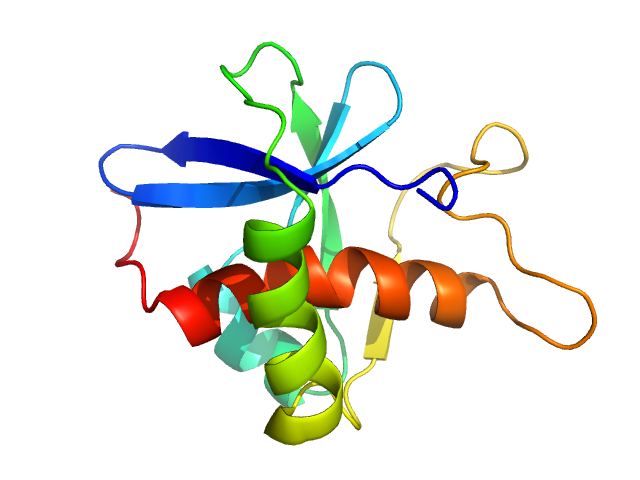
UniProt ID: M1L6N7 (944-1066) Genome polyprotein
|
|
|
|
| Sample: |
Genome polyprotein monomer, 13 kDa Turkey avisivirus protein
|
| Buffer: |
20 mM citrate pH 5.4, 150 mM NaCl, 2 mM TCEP, 1% glycerol, pH: 5.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 May 2
|
Solution structure and functional characterization of an avian picornavirus 2AH/NC protein
Anna Wehlin
|
| RgGuinier |
1.7 |
nm |
| Dmax |
7.2 |
nm |
| VolumePorod |
29 |
nm3 |
|
|