UniProt ID: P09651 (1-196) Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S )
|
|
|
|
| Sample: |
Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S ) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, 350 mM NaCl, 0.5 mM EDTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2023 Mar 1
|
Thermodynamic coupling of the tandem RRM domains of hnRNP A1 underlie its pleiotropic RNA binding functions.
Sci Adv 10(28):eadk6580 (2024)
Levengood JD, Potoyan D, Penumutchu S, Kumar A, Zhou Q, Wang Y, Hansen AL, Kutluay S, Roche J, Tolbert BS
|
|
|
UniProt ID: P09651 (1-196) Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S )
|
|
|
|
| Sample: |
Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S ) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, 350 mM NaCl, 0.5 mM EDTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2023 Mar 1
|
Thermodynamic coupling of the tandem RRM domains of hnRNP A1 underlie its pleiotropic RNA binding functions.
Sci Adv 10(28):eadk6580 (2024)
Levengood JD, Potoyan D, Penumutchu S, Kumar A, Zhou Q, Wang Y, Hansen AL, Kutluay S, Roche J, Tolbert BS
|
|
|
UniProt ID: P09651 (1-196) Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S )
|
|
|
|
| Sample: |
Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S ) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, 350 mM NaCl, 0.5 mM EDTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2023 Mar 1
|
Thermodynamic coupling of the tandem RRM domains of hnRNP A1 underlie its pleiotropic RNA binding functions.
Sci Adv 10(28):eadk6580 (2024)
Levengood JD, Potoyan D, Penumutchu S, Kumar A, Zhou Q, Wang Y, Hansen AL, Kutluay S, Roche J, Tolbert BS
|
|
|
UniProt ID: P09651 (1-196) Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S )
|
|
|
|
| Sample: |
Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S ) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, 350 mM NaCl, 0.5 mM EDTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2023 Mar 1
|
Thermodynamic coupling of the tandem RRM domains of hnRNP A1 underlie its pleiotropic RNA binding functions.
Sci Adv 10(28):eadk6580 (2024)
Levengood JD, Potoyan D, Penumutchu S, Kumar A, Zhou Q, Wang Y, Hansen AL, Kutluay S, Roche J, Tolbert BS
|
|
|
UniProt ID: P09651 (1-196) Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S )
|
|
|
|
| Sample: |
Heterogeneous nuclear ribonucleoprotein A1 (C43S/R75D/R88D/C175S ) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, 350 mM NaCl, 0.5 mM EDTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2023 Mar 1
|
Thermodynamic coupling of the tandem RRM domains of hnRNP A1 underlie its pleiotropic RNA binding functions.
Sci Adv 10(28):eadk6580 (2024)
Levengood JD, Potoyan D, Penumutchu S, Kumar A, Zhou Q, Wang Y, Hansen AL, Kutluay S, Roche J, Tolbert BS
|
|
|
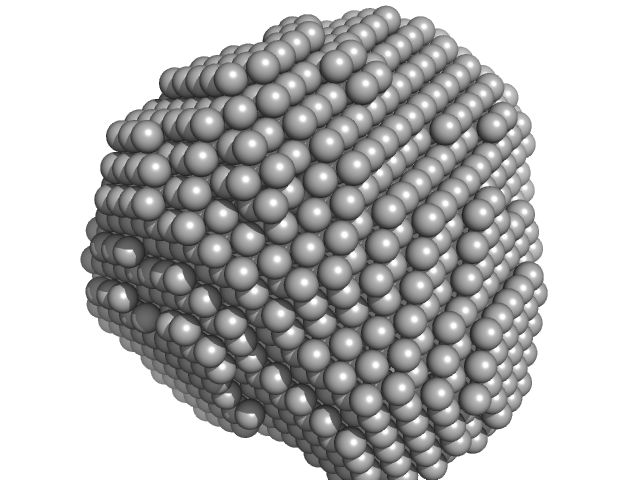
UniProt ID: O06310 (1-145) Conserved protein
|
|
|

|
| Sample: |
Conserved protein dimer, 31 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris pH 7.5, 200 mM NaCl, 2% glycerol, pH: |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2022 May 4
|
Molecular mechanisms underlying the functions of biofilm forming protein of mycobacterium tuberculosis
farheen jahan
|
| RgGuinier |
1.9 |
nm |
| Dmax |
4.4 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
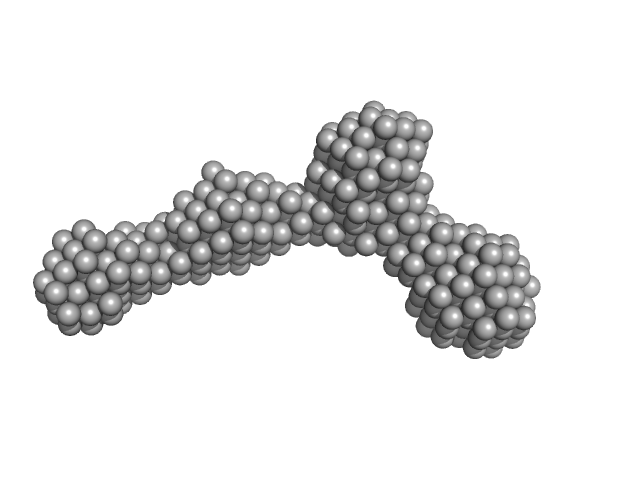
UniProt ID: P9WFC7 (22-719) UvrABC system protein B
|
|
|
|
| Sample: |
UvrABC system protein B dimer, 156 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris pH-8, 200 mM NaCl, 2% glycerol, pH: |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2023 Jun 6
|
Inhibition of Mycobacterium tuberculosis UvrB by small molecules: Potent NER disruption and structural insights into dimer conformation.
Int J Biol Macromol :147338 (2025)
Jahan F, Anand S, Kumar S, Pandey G, Siddiqi MI, Krishnan MY, Ramachandran R
|
| RgGuinier |
6.7 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
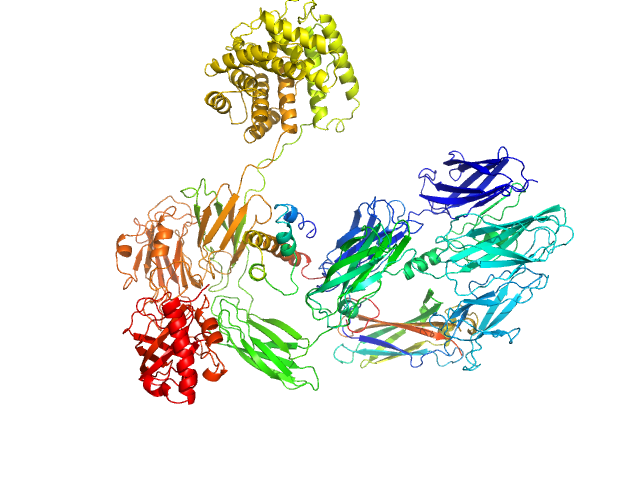
UniProt ID: P01024 (1-1663) Complement C3 (Δ668-671)
|
|
|
|
| Sample: |
Complement C3 (Δ668-671) monomer, 187 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES pH 6.0, 200 mM NaCl, pH: 6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Nov 7
|
Cryo-EM analysis of complement C3 reveals a reversible major opening of the macroglobulin ring.
Nat Struct Mol Biol (2025)
Gadeberg TAF, Jørgensen MH, Olesen HG, Lorentzen J, Harwood SL, Almeida AV, Fruergaard MU, Jensen RK, Kanis P, Pedersen H, Tranchant E, Petersen SV, Thøgersen IB, Kragelund BB, Lyons JA, Enghild JJ, Andersen GR
|
| RgGuinier |
5.4 |
nm |
| Dmax |
21.8 |
nm |
| VolumePorod |
357 |
nm3 |
|
|
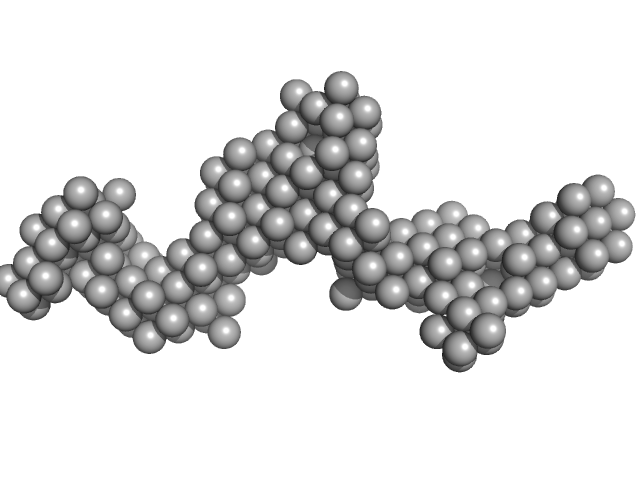
UniProt ID: E9Q9C6 (27-2583) Fc fragment of IgG binding protein
|
|
|

|
| Sample: |
Fc fragment of IgG binding protein dimer, 552 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES, 100 mM NaCl, 10 mM CaCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Nov 20
|
The structure of FCGBP is formed as a disulfide-mediated homodimer between its C-terminal domains.
FEBS J (2025)
Ehrencrona E, Gallego P, Trillo-Muyo S, Garcia-Bonete MJ, Recktenwald CV, Hansson GC, Johansson MEV
|
| RgGuinier |
10.0 |
nm |
| Dmax |
42.0 |
nm |
| VolumePorod |
1462 |
nm3 |
|
|
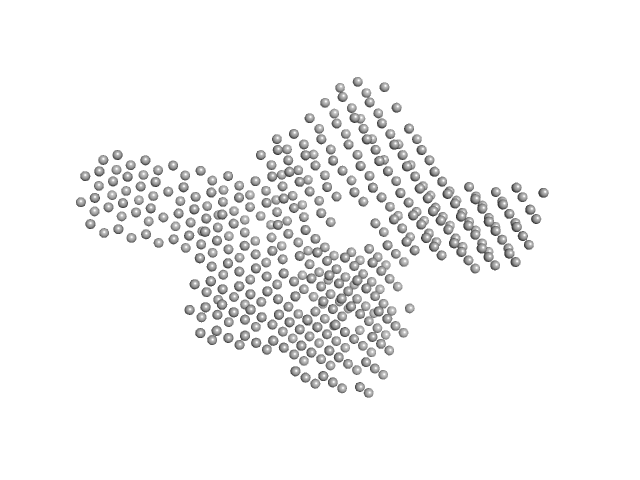
UniProt ID: Q2FHL2 (1-548) Uncharacterized protein SAUSA300_1119
|
|
|

|
| Sample: |
Uncharacterized protein SAUSA300_1119 dimer, 125 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM KCl, 1 mM TCEP, 5% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Oct 5
|
Molecular insights into the structure and function of the Staphylococcus aureus fatty acid kinase.
J Biol Chem 300(12):107920 (2024)
Myers MJ, Xu Z, Ryan BJ, DeMars ZR, Ridder MJ, Johnson DK, Krute CN, Flynn TS, Kashipathy MM, Battaile KP, Schnicker N, Lovell S, Freudenthal BD, Bose JL
|
| RgGuinier |
4.3 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
219 |
nm3 |
|
|