UniProt ID: None (1-113) phosphorylated Heterochromatin protein HP1α, delta CSD b4 mutant
|
|
|
|
| Sample: |
Phosphorylated Heterochromatin protein HP1α, delta CSD b4 mutant monomer, 13 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 50 mM NaCl, 1 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Nov 27
|
A dynamic structural unit of phase-separated heterochromatin protein 1α as revealed by integrative structural analyses
Nucleic Acids Research 53(6) (2025)
Furukawa A, Yonezawa K, Negami T, Yoshimura Y, Hayashi A, Nakayama J, Adachi N, Senda T, Shimizu K, Terada T, Shimizu N, Nishimura Y
|
| RgGuinier |
2.7 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
34 |
nm3 |
|
|
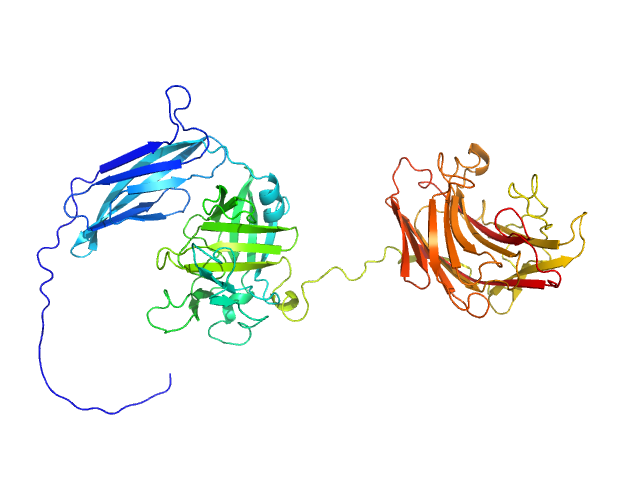
UniProt ID: A0M710-1 (29-538) Glycosyl hydrolase, family 16
|
|
|
|
| Sample: |
Glycosyl hydrolase, family 16 monomer, 58 kDa Christiangramia forsetii (strain … protein
|
| Buffer: |
10 mM MOPS, 100 mM NaCl, pH: 7.8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 1
|
Unveiling the role of novel carbohydrate-binding modules in laminarin interaction of multimodular proteins from marine Bacteroidota during phytoplankton blooms.
Environ Microbiol 26(5):e16624 (2024)
Zühlke MK, Ficko-Blean E, Bartosik D, Terrapon N, Jeudy A, Jam M, Wang F, Welsch N, Dürwald A, Martin LT, Larocque R, Jouanneau D, Eisenack T, Thomas F, Trautwein-Schult A, Teeling H, Becher D, Schweder T, Czjzek M
|
| RgGuinier |
3.8 |
nm |
| Dmax |
12.9 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
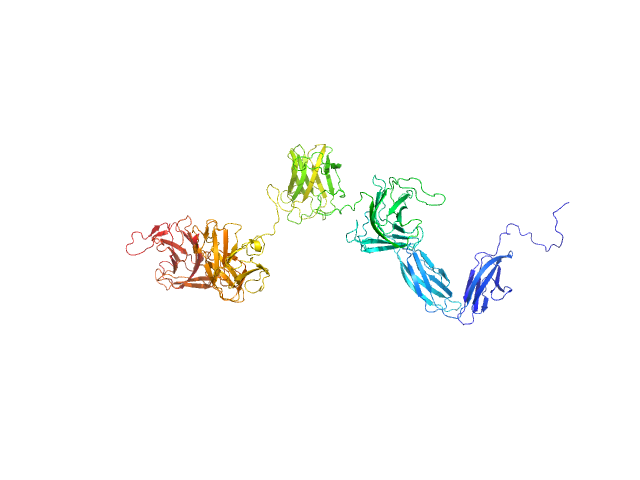
UniProt ID: A0M709 (21-896) PKD domain-containing protein
|
|
|
|
| Sample: |
PKD domain-containing protein monomer, 96 kDa Christiangramia forsetii (strain … protein
|
| Buffer: |
10 mM MOPS, 100 mM NaCl, pH: 7.8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 1
|
Unveiling the role of novel carbohydrate-binding modules in laminarin interaction of multimodular proteins from marine Bacteroidota during phytoplankton blooms.
Environ Microbiol 26(5):e16624 (2024)
Zühlke MK, Ficko-Blean E, Bartosik D, Terrapon N, Jeudy A, Jam M, Wang F, Welsch N, Dürwald A, Martin LT, Larocque R, Jouanneau D, Eisenack T, Thomas F, Trautwein-Schult A, Teeling H, Becher D, Schweder T, Czjzek M
|
| RgGuinier |
5.0 |
nm |
| Dmax |
17.8 |
nm |
| VolumePorod |
138 |
nm3 |
|
|
UniProt ID: Q04917 (1-246) 14-3-3 protein eta
|
|
|
|
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM EDTA, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 4
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
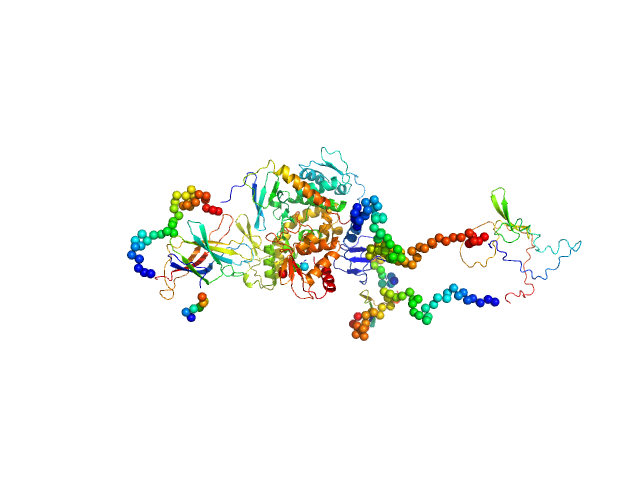
UniProt ID: Q96PU5-5 (1-955) Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like
|
|
|
|
| Sample: |
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM EDTA, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
Structural basis of ubiquitin ligase Nedd4-2 autoinhibition and regulation by calcium and 14-3-3 proteins
Nature Communications 16(1) (2025)
Janosev M, Kosek D, Tekel A, Joshi R, Honzejkova K, Pohl P, Obsil T, Obsilova V
|
| RgGuinier |
4.9 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
206 |
nm3 |
|
|
UniProt ID: Q96PU5-5 (1-955) Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like
|
|
|
|
| Sample: |
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM CaCl2, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
Structural basis of ubiquitin ligase Nedd4-2 autoinhibition and regulation by calcium and 14-3-3 proteins
Nature Communications 16(1) (2025)
Janosev M, Kosek D, Tekel A, Joshi R, Honzejkova K, Pohl P, Obsil T, Obsilova V
|
| RgGuinier |
4.9 |
nm |
| Dmax |
22.5 |
nm |
| VolumePorod |
201 |
nm3 |
|
|
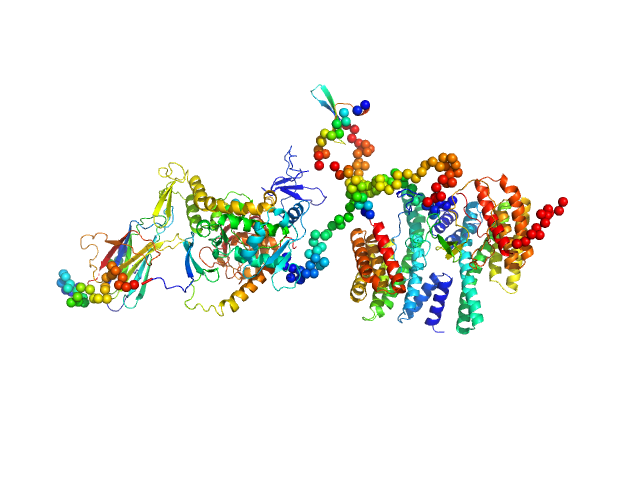
UniProt ID: Q04917 (1-246) 14-3-3 protein eta
UniProt ID: Q96PU5-5 (1-955) Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like
|
|
|
|
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM EDTA, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
5.4 |
nm |
| Dmax |
21.4 |
nm |
| VolumePorod |
281 |
nm3 |
|
|
UniProt ID: Q04917 (1-246) 14-3-3 protein eta
UniProt ID: Q96PU5-5 (1-955) Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like
|
|
|
|
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM CaCl2, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 4
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
5.5 |
nm |
| Dmax |
23.5 |
nm |
| VolumePorod |
290 |
nm3 |
|
|
UniProt ID: N4UN01 (21-303) Endoglucanase-7
|
|
|
|
| Sample: |
Endoglucanase-7 monomer, 30 kDa Colletotrichum orbiculare (strain … protein
|
| Buffer: |
50 mM MES, 150 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 1
|
The disordered C-terminal tail of fungal LPMOs from phytopathogens mediates protein dimerization and impacts plant penetration.
Proc Natl Acad Sci U S A 121(13):e2319998121 (2024)
Tamburrini KC, Kodama S, Grisel S, Haon M, Nishiuchi T, Bissaro B, Kubo Y, Longhi S, Berrin JG
|
| RgGuinier |
3.0 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
UniProt ID: N4UN01 (21-303) Endoglucanase-7
|
|
|
|
| Sample: |
Endoglucanase-7 dimer, 60 kDa Colletotrichum orbiculare (strain … protein
|
| Buffer: |
50 mM MES, 150 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 1
|
The disordered C-terminal tail of fungal LPMOs from phytopathogens mediates protein dimerization and impacts plant penetration.
Proc Natl Acad Sci U S A 121(13):e2319998121 (2024)
Tamburrini KC, Kodama S, Grisel S, Haon M, Nishiuchi T, Bissaro B, Kubo Y, Longhi S, Berrin JG
|
| RgGuinier |
4.8 |
nm |
| Dmax |
21.7 |
nm |
| VolumePorod |
83 |
nm3 |
|
|