UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-mannose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris/HCl, 1 mM DTT, 10 mM D-glucose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
|
|
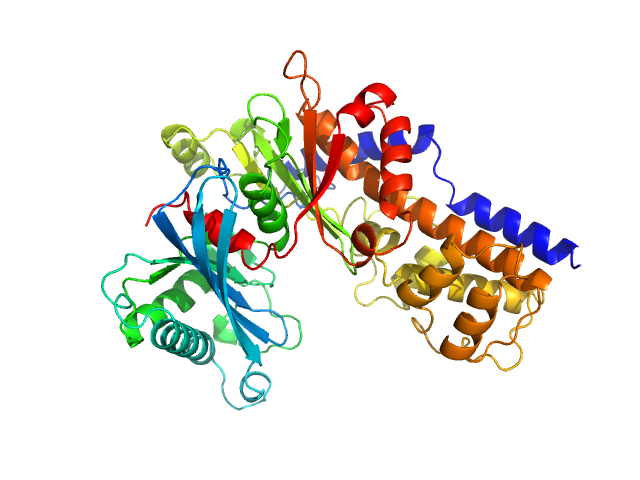
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
|
|
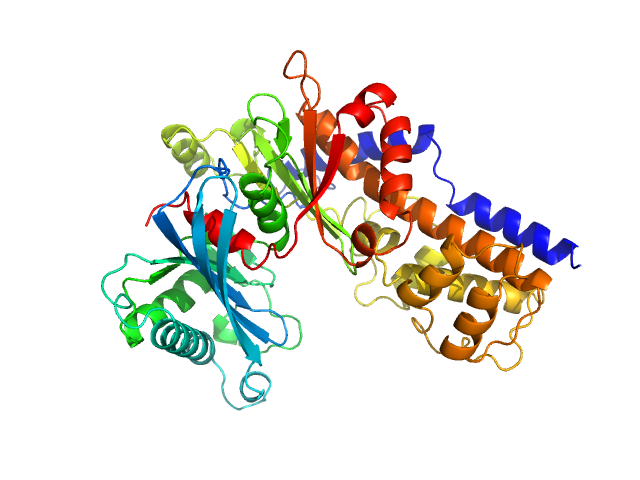
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-fructose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
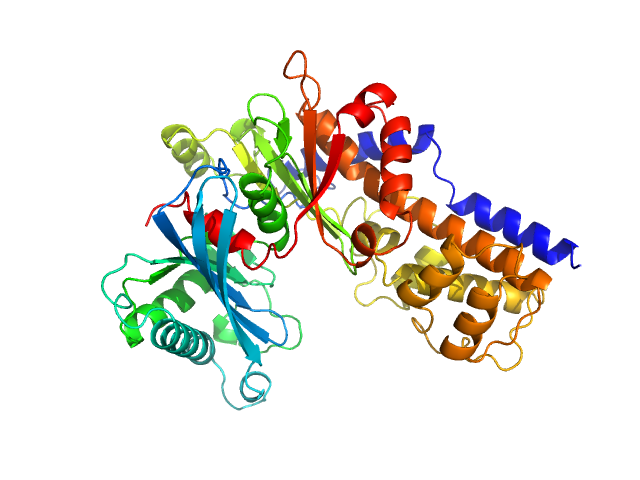
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-mannose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
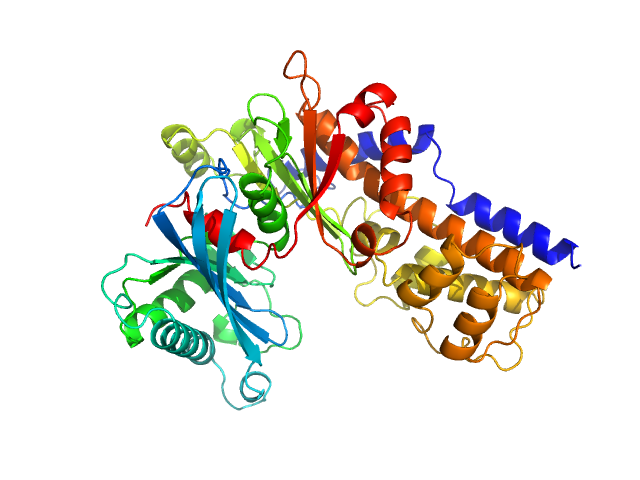
UniProt ID: Q6CUZ3 (1-481) Glucokinase-1 (H304Q, I356D, Y419D, H420D)
|
|
|
|
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris/HCl, 1 mM DTT, 10 mM D-glucose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: P61825 (756-836) Structural polyprotein
|
|
|
|
| Sample: |
Structural polyprotein dimer, 26 kDa Infectious bursal disease … protein
|
| Buffer: |
50 mM Tris, 500 mM NaCl, 2 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Sep 27
|
Infectious Bursal Disease Virus VP3
Diego Ferrero
|
| RgGuinier |
2.4 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
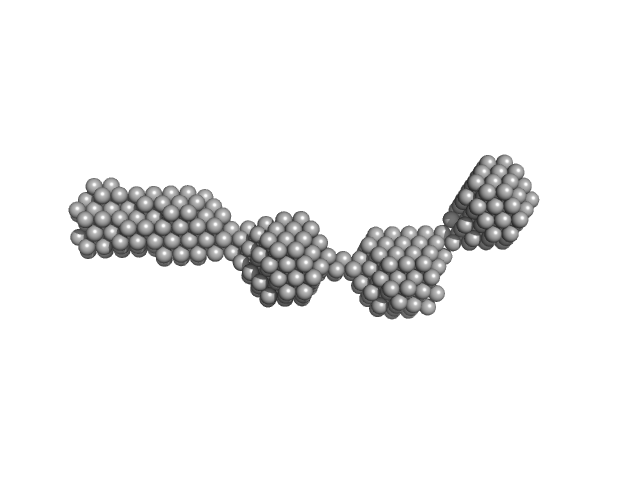
UniProt ID: P11709 (2-440) Nuclear fusion protein BIK1
|
|
|
|
| Sample: |
Nuclear fusion protein BIK1 dimer, 106 kDa Saccharomyces cerevisiae (strain … protein
|
| Buffer: |
20 mM Tris-HCl pH 7.5, 500 mM NaCl, 2% glycerol, 1 mM DTT, pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Jul 13
|
Phase separation of a microtubule plus-end tracking protein into a fluid fractal network
(2024)
Czub M, Uliana F, Grubić T, Padeste C, Rosowski K, Dufresne E, Menzel A, Vakonakis I, Gasser U, Steinmetz M
|
| RgGuinier |
8.8 |
nm |
| Dmax |
28.0 |
nm |
| VolumePorod |
467 |
nm3 |
|
|