UniProt ID: A0A5N5URA7 (4-661) Peptidase S9
|
|
|

|
| Sample: |
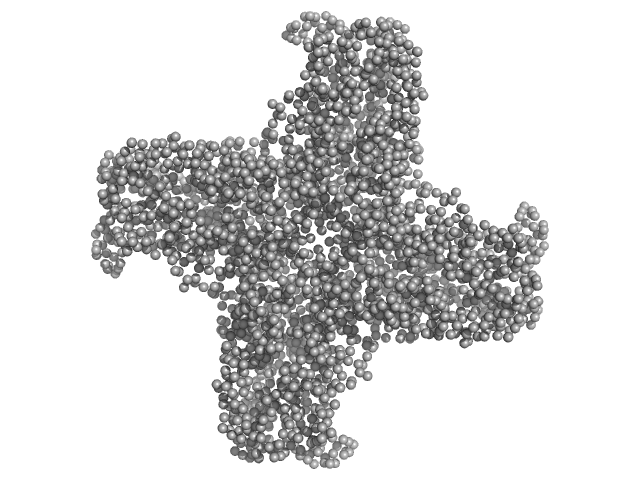
Peptidase S9 tetramer, 304 kDa Mycolicibacterium phlei DSM … protein
|
| Buffer: |
10 mM Tris-HCl, 135 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL-18, INDUS-2 on 2024 Mar 21
|
Structural and Functional Investigation of Putative Peptidase from Mycolicibacterium phlei
: An Exclusive Endopeptidase among S9C Subfamily
Biochemistry (2025)
Chandravanshi K, Jamdar S, Singh R, Kumar A, Mahato A, Agrawal R, Kumar S, Kumar A, Makde R
|
| RgGuinier |
5.1 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
563 |
nm3 |
|
|
UniProt ID: P23246 (214-707) Splicing factor, proline- and glutamine-rich
|
|
|
|
| Sample: |
Splicing factor, proline- and glutamine-rich dimer, 153 kDa Homo sapiens protein
|
| Buffer: |
150 mM KCl, 0.075% glycerol, 20 mM HEPES, 0.5 mM DTT,, pH: 7.4 |
| Experiment: |
SANS
data collected at Quokka - Small Angle Neutron Scattering, Australian Centre for Neutron Scattering (ANSTO) on 2023 Jan 20
|
Structural dynamics of IDR interactions in human SFPQ and implications for liquid–liquid phase separation
Acta Crystallographica Section D Structural Biology 81(7):357-379 (2025)
Koning H, Lai V, Sethi A, Chakraborty S, Ang C, Fox A, Duff A, Whitten A, Marshall A, Bond C
|
| RgGuinier |
6.1 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
181 |
nm3 |
|
|
UniProt ID: P29268 (26-348) Cellular communication network factor 2
|
|
|
|
| Sample: |
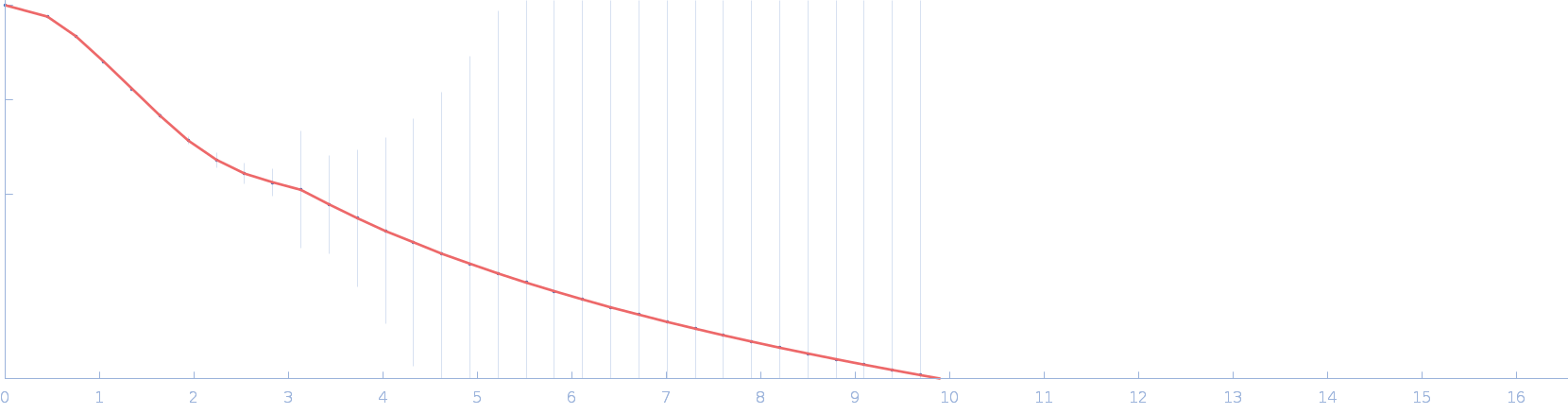
Cellular communication network factor 2 monomer, 36 kDa protein
|
| Buffer: |
50 mM Tris pH 7.5, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 6
|
Structural basis for therapeutic antibody binding to CCN2 and its mechanisms on TGF-β family signaling
Haben Gabir
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
UniProt ID: P29268 (26-348) Cellular communication network factor 2
UniProt ID: None (None-None) anti-CCN2 37-45-MH1 F(ab) antibody
|
|
|
|
| Sample: |
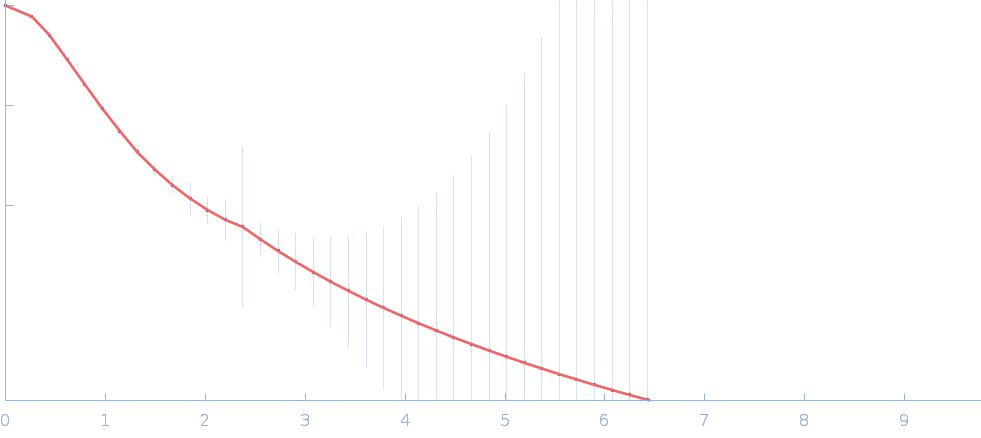
Cellular communication network factor 2 monomer, 36 kDa protein
Anti-CCN2 37-45-MH1 F(ab) antibody monomer, 49 kDa protein
|
| Buffer: |
50 mM Tris pH 7.5, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 6
|
Structural basis for therapeutic antibody binding to CCN2 and its mechanisms on TGF-β family signaling
Haben Gabir
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.9 |
nm |
| VolumePorod |
95 |
nm3 |
|
|
UniProt ID: Q9SQQ6 (172-338) NAC domain-containing protein 46
|
|
|
|
| Sample: |
NAC domain-containing protein 46 monomer, 18 kDa Arabidopsis thaliana protein
|
| Buffer: |
20mM Na2HPO4/NaH2PO4, 100mM NaCl, 5mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 1
|
Hierarchy in regulator interactions with distant transcriptional activation domains empowers rheostatic regulation
Protein Science 34(6) (2025)
Due A, Davey N, Thomasen F, Morffy N, Prestel A, Brakti I, O'Shea C, Strader L, Lindorff‐Larsen K, Skriver K, Kragelund B
|
| RgGuinier |
3.5 |
nm |
| Dmax |
16.5 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
UniProt ID: Q9SQQ6 (172-338) NAC domain-containing protein 46
UniProt ID: Q8RY59 (499-572) Inactive poly [ADP-ribose] polymerase RCD1
|
|
|
|
| Sample: |
NAC domain-containing protein 46 monomer, 18 kDa Arabidopsis thaliana protein
Inactive poly [ADP-ribose] polymerase RCD1 monomer, 9 kDa Arabidopsis thaliana protein
|
| Buffer: |
20mM Na2HPO4/NaH2PO4, 100mM NaCl, 5mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 1
|
Hierarchy in regulator interactions with distant transcriptional activation domains empowers rheostatic regulation
Protein Science 34(6) (2025)
Due A, Davey N, Thomasen F, Morffy N, Prestel A, Brakti I, O'Shea C, Strader L, Lindorff‐Larsen K, Skriver K, Kragelund B
|
| RgGuinier |
3.8 |
nm |
| Dmax |
20.6 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
UniProt ID: P48047 (None-None) ATP synthase subunit O, mitochondrial
|
|
|
|
| Sample: |
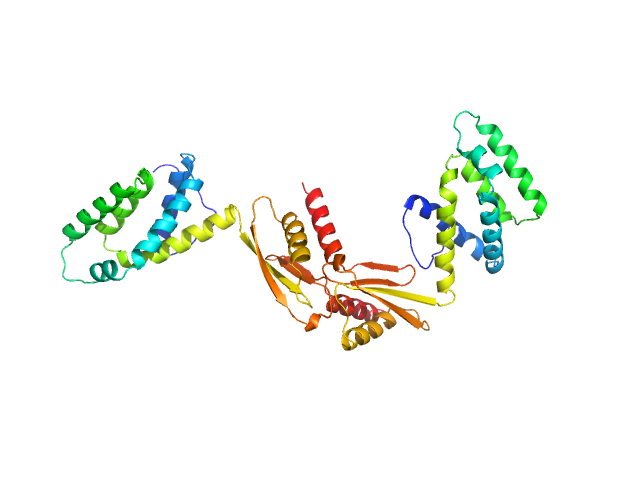
ATP synthase subunit O, mitochondrial dimer, 42 kDa Homo sapiens protein
|
| Buffer: |
50 mM Na2HPO4, 300 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Mar 15
|
The OSCP Subunit of ATP Synthase is a Dimer in Solution: Strategy to Induce the Monomeric Protein as a New Tool for Drug Discovery.
J Mol Biol 437(17):169267 (2025)
Fabbian S, Gabbatore L, Morbiato L, Zotti M, Giorgio V, Battisutta R, Giachin G, Sosic A, Bellanda M
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
78 |
nm3 |
|
|
UniProt ID: P48047 (137-213) ATP synthase subunit O, mitochondrial C-terminal section
|
|
|
|
| Sample: |
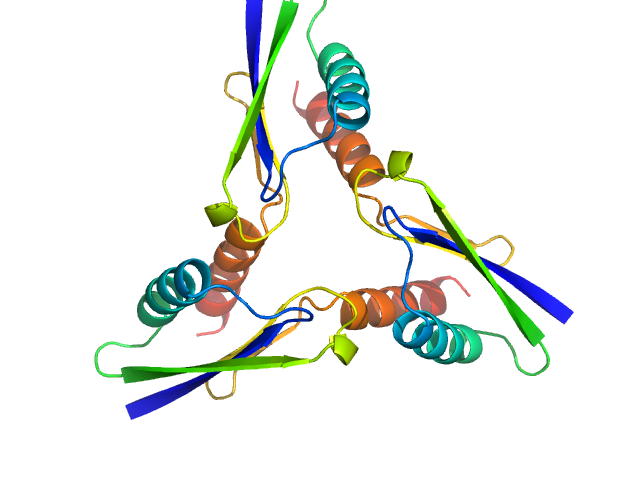
ATP synthase subunit O, mitochondrial C-terminal section trimer, 25 kDa Homo sapiens protein
|
| Buffer: |
50 mM Na2HPO4, 300 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Mar 15
|
The OSCP Subunit of ATP Synthase is a Dimer in Solution: Strategy to Induce the Monomeric Protein as a New Tool for Drug Discovery.
J Mol Biol 437(17):169267 (2025)
Fabbian S, Gabbatore L, Morbiato L, Zotti M, Giorgio V, Battisutta R, Giachin G, Sosic A, Bellanda M
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
UniProt ID: Q8A9M2 (1-438) Xylose isomerase
|
|
|

|
| Sample: |
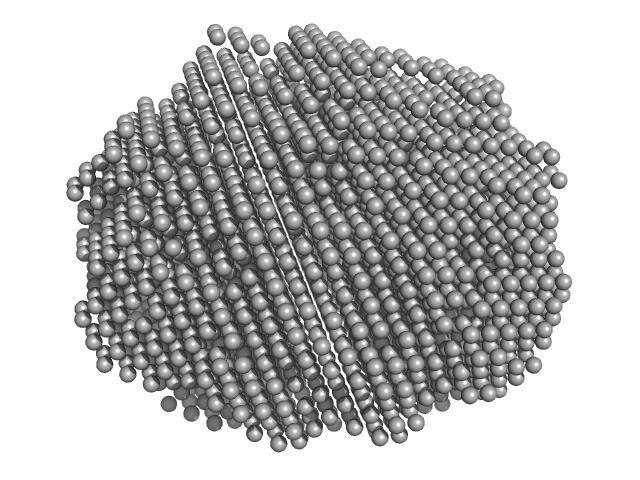
Xylose isomerase tetramer, 196 kDa Bacteroides thetaiotaomicron (strain … protein
|
| Buffer: |
PBS, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2024 Oct 22
|
BioSAXS Australian Synchrotron Standard Protein
Annmaree Warrender
|
| RgGuinier |
3.5 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
222 |
nm3 |
|
|
UniProt ID: P12109 (566-1028) Collagen alpha-1(VI) chain
UniProt ID: P12110 (564-1019) Collagen alpha-2(VI) chain (R680H, K966N, Q967E)
UniProt ID: P12111 (2347-2820) Collagen alpha-3(VI) chain (D2357R, K2367R, D2431V, R2441T, R2609A, R2610A)
|
|
|
|
| Sample: |
Collagen alpha-1(VI) chain, 52 kDa Homo sapiens protein
Collagen alpha-2(VI) chain (R680H, K966N, Q967E), 53 kDa Homo sapiens protein
Collagen alpha-3(VI) chain (D2357R, K2367R, D2431V, R2441T, R2609A, R2610A), 54 kDa Homo sapiens protein
|
| Buffer: |
Tris-buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Jul 27
|
Collagen VI microfibril structure reveals mechanism for molecular assembly and clustering of inherited pathogenic mutations.
Nat Commun 16(1):7549 (2025)
Godwin ARF, Becker MH, Dajani R, Snee M, Roseman AM, Baldock C
|
| RgGuinier |
4.8 |
nm |
| Dmax |
18.7 |
nm |
| VolumePorod |
478 |
nm3 |
|
|