UniProt ID: Q8LBL5 (1-410) E3 ubiquitin-protein ligase PRT1
|
|
|
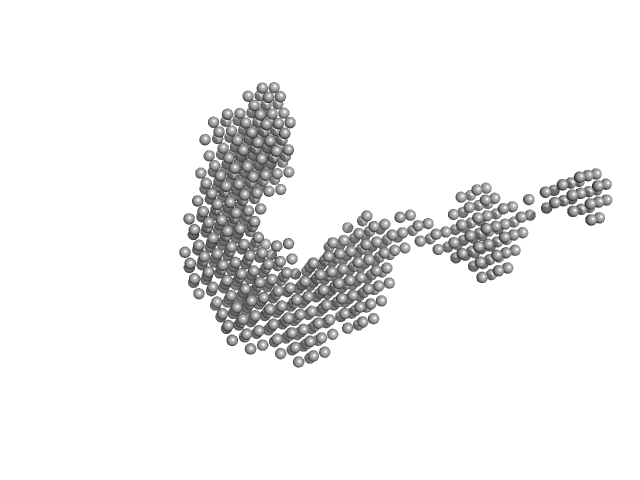
|
| Sample: |
E3 ubiquitin-protein ligase PRT1 monomer, 46 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM TCEP, 2% w/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2018 May 22
|
Structural basis for the recognition and ubiquitylation of type-2 N-degron substrate by PRT1 plant N-recognin
Nature Communications 16(1) (2025)
Yang W, Kim S, Kim M, Shin H, Lee J, Sandmann A, Park O, Dissmeyer N, Song H
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
UniProt ID: Q71MD6 (1-697) Prolyl endopeptidase
|
|
|
|
| Sample: |
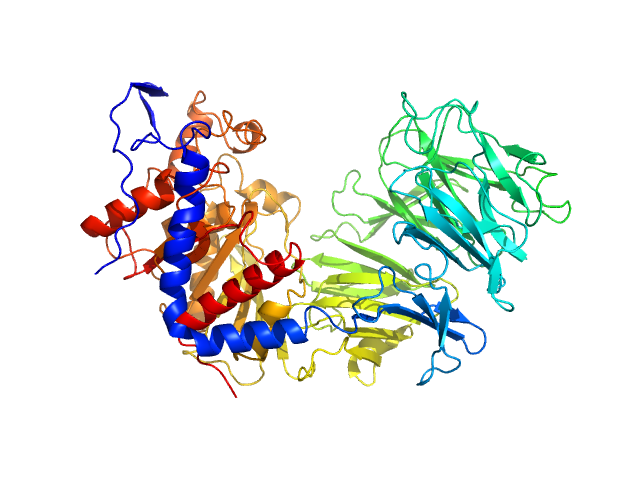
Prolyl endopeptidase monomer, 78 kDa Trypanosoma cruzi protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Apr 24
|
Cryo-EM led analysis of open and closed conformations of Chagas vaccine candidate TcPOP.
Nat Commun 16(1):7164 (2025)
Batra S, Olmo F, Ragan TJ, Kaplan M, Calvaresi V, Frank AM, Lancey C, Assadipapari M, Ying C, Struwe WB, Hesketh EL, Kelly JM, Barfod L, Campeotto I
|
| RgGuinier |
3.0 |
nm |
| Dmax |
0.0 |
nm |
| VolumePorod |
142 |
nm3 |
|
|
UniProt ID: Q71MD6 (None-None) Prolyl endopeptidase
|
|
|
|
| Sample: |
Prolyl endopeptidase monomer, 78 kDa Trypanosoma cruzi protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Apr 24
|
Cryo-EM led analysis of open and closed conformations of Chagas vaccine candidate TcPOP.
Nat Commun 16(1):7164 (2025)
Batra S, Olmo F, Ragan TJ, Kaplan M, Calvaresi V, Frank AM, Lancey C, Assadipapari M, Ying C, Struwe WB, Hesketh EL, Kelly JM, Barfod L, Campeotto I
|
| RgGuinier |
3.8 |
nm |
| Dmax |
0.0 |
nm |
| VolumePorod |
170 |
nm3 |
|
|
UniProt ID: P01012 (2-386) Ovalbumin
|
|
|
|
| Sample: |
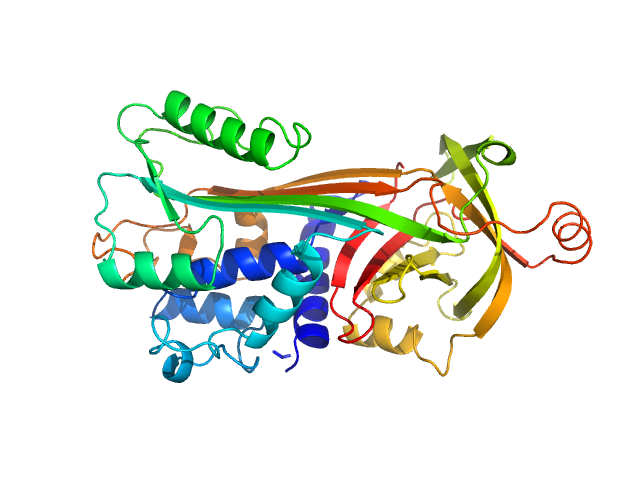
Ovalbumin monomer, 43 kDa Gallus gallus protein
|
| Buffer: |
100 mM Tris-HCl, 100 mM NaCl in 100% v/v D₂O, pH: 7.5 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Apr 22
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.5 |
nm |
|
|
UniProt ID: P02769 (None-None) Albumin (natural variant A214T)
|
|
|
|
| Sample: |
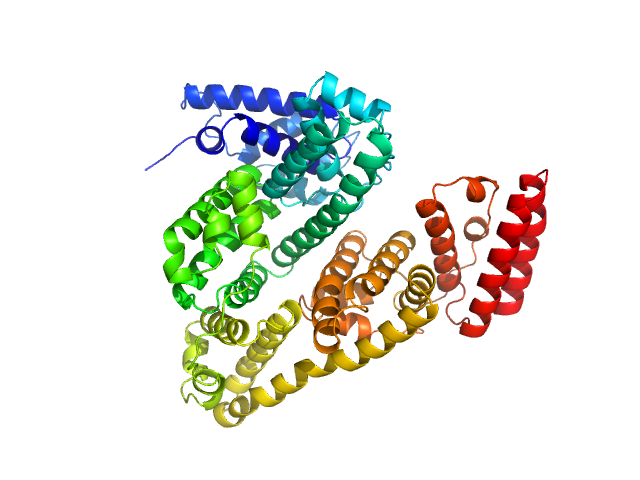
Albumin (natural variant A214T) monomer, 66 kDa Bos taurus protein
|
| Buffer: |
100 mM Tris-HCl, 100 mM NaCl in 100% v/v D₂O, pH: 7.5 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Apr 22
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.7 |
nm |
|
|
UniProt ID: P02791 (2-175) Ferritin light chain
|
|
|
|
| Sample: |
Ferritin light chain 24-mer, 476 kDa Equus caballus protein
|
| Buffer: |
100 mM Tris-HCl, 100 mM NaCl in 100% v/v D₂O, pH: 7.5 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Apr 22
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
5.4 |
nm |
| Dmax |
23.1 |
nm |
|
|
UniProt ID: Q5N595 (1-102) Circadian clock oscillator protein KaiB
UniProt ID: Q79PF4 (2-519) Circadian clock oscillator protein KaiC
|
|
|
|
| Sample: |
Circadian clock oscillator protein KaiB hexamer, 71 kDa Synechococcus sp. (strain … protein
Circadian clock oscillator protein KaiC hexamer, 356 kDa Synechococcus elongatus (strain … protein
|
| Buffer: |
50 mM sodium phosphate buffer, 150 mM NaCl, 5 mM MgCl₂, 0.5 mM EDTA, 1 mM ATP, 1 mM DTT, 50 mM arginine, 50 mM glutamine, in 100% D₂O, pH: 7.8 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Jul 2
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
4.9 |
nm |
| Dmax |
14.4 |
nm |
|
|
UniProt ID: Q5N595 (1-102) Circadian clock oscillator protein KaiB
UniProt ID: Q79PF4 (2-519) Circadian clock oscillator protein KaiC
|
|
|
|
| Sample: |
Circadian clock oscillator protein KaiB hexamer, 71 kDa Synechococcus sp. (strain … protein
Circadian clock oscillator protein KaiC hexamer, 356 kDa Synechococcus elongatus (strain … protein
|
| Buffer: |
50 mM sodium phosphate buffer, 150 mM NaCl, 5 mM MgCl₂, 0.5 mM EDTA, 1 mM ATP, 1 mM DTT, 50 mM arginine, 50 mM glutamine, in 100% D₂O, pH: 7.8 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Jul 2
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
4.3 |
nm |
| Dmax |
11.7 |
nm |
|
|
UniProt ID: Q79PF4 (2-519) Circadian clock oscillator protein KaiC
|
|
|
|
| Sample: |
Circadian clock oscillator protein KaiC hexamer, 356 kDa Synechococcus elongatus (strain … protein
|
| Buffer: |
50 mM sodium phosphate buffer, 150 mM NaCl, 5 mM MgCl₂, 0.5 mM EDTA, 1 mM ATP, 1 mM DTT, 50 mM arginine, 50 mM glutamine, in 100% D₂O, pH: 7.8 |
| Experiment: |
SANS
data collected at SANS-U, JRR-3 (Japan Atomic Energy Agency) on 2024 Jul 2
|
Size-exclusion chromatography–small-angle neutron scattering system optimized for an instrument with medium neutron flux
Journal of Applied Crystallography 58(2) (2025)
Morishima K, Inoue R, Nakagawa T, Shimizu M, Sakamoto R, Oda T, Mayumi K, Sugiyama M
|
| RgGuinier |
4.4 |
nm |
| Dmax |
11.9 |
nm |
|
|
UniProt ID: A0A0H3MBY4 (89-215) Candidate inclusion membrane protein (MBP-tagged)
|
|
|
|
| Sample: |
Candidate inclusion membrane protein (MBP-tagged) tetramer, 235 kDa Chlamydia trachomatis serovar … protein
|
| Buffer: |
20 mM Tris-HCl, 50 mM NaCl, 1 mM EDTA, 2% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2023 Feb 15
|
Tetramer formation of CpoS facilitates Inc-Inc interactions during Chlamydia trachomatis infection
Zhen Xu
|
| RgGuinier |
7.1 |
nm |
| Dmax |
30.3 |
nm |
| VolumePorod |
625 |
nm3 |
|
|