UniProt ID: P68104 (1-462) Mammalian translation elongation factor eEF1A1
|
|
|
|
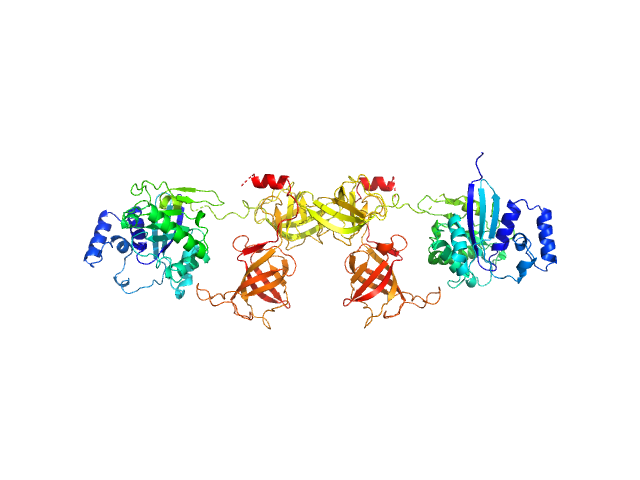
| Sample: |
Mammalian translation elongation factor eEF1A1 dimer, 100 kDa protein
|
| Buffer: |
25 mM Tris HCl, 150 mM NaCl, 6 mM βME, 20% glycerol, 0.01mM GDP,, pH: 7.5 |
| Experiment: |
SAXS
data collected at Bruker Nanostar w Excillum source, Department of Chemistry, iNANO building, Aarhus Uinversity on 2023 Mar 20
|
Dynamics and structural features of the eEF1A1 and eEF1A2 paralogs
Nucleic Acids Research 53(21) (2025)
Novosylna O, Shalak V, Dąbrowska K, Patmanidis I, Lozhko D, Bondarchuk T, Schiøtt B, Pedersen J, Knudsen C, Nissen P, Dadlez M, Negrutskii B
|
| RgGuinier |
4.6 |
nm |
| Dmax |
143.1 |
nm |
| VolumePorod |
144 |
nm3 |
|
|
UniProt ID: Q9Z2Y3 (1-118) EVH1 domain of Homer protein homolog 1 from mouse
|
|
|
|
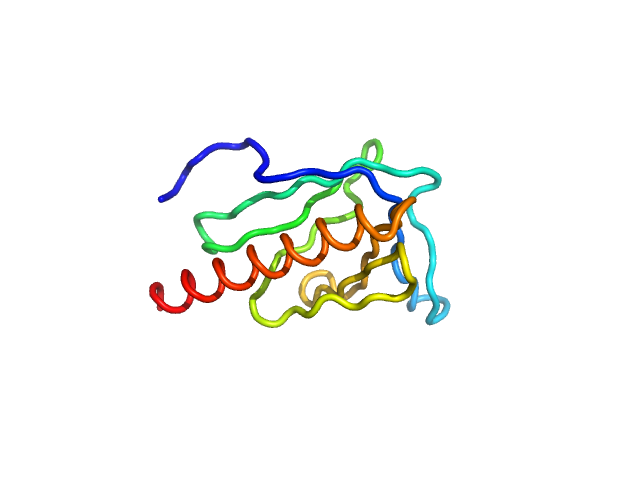
| Sample: |
EVH1 domain of Homer protein homolog 1 from mouse monomer, 14 kDa Mus musculus protein
|
| Buffer: |
20mM NaCl, 50 mM NaPi, 0.02% of sodium-azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, CEITEC on 2023 Nov 9
|
Structural Modeling and Dynamics of the Full-Length Homer1 Multimer.
Proteins (2025)
Kálmán ZE, Czajlik A, Maruzs B, Farkas F, Pap I, Homonnay C, Klumpler T, Batta G, Gáspári Z, Péterfia B
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) Direct Repeat binding site with zero-base-pair spacing
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
Direct Repeat binding site with zero-base-pair spacing monomer, 10 kDa DNA
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2024 Apr 18
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
3.6 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
121 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: None (None-None) Direct Repeat binding site with zero-base-pair spacing (long sequence)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Direct Repeat binding site with zero-base-pair spacing (long sequence) monomer, 19 kDa DNA
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 12
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
4.1 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
152 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Direct Repeat binding site with one-base-pair spacing
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Direct Repeat binding site with one-base-pair spacing monomer, 11 kDa DNA
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 20
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Direct Repeat binding site with five-base-pair spacing
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Direct Repeat binding site with five-base-pair spacing monomer, 13 kDa DNA
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Sep 12
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.3 |
nm |
| VolumePorod |
130 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Inverted Repeat binding site with zero-base-pair spacing
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Inverted Repeat binding site with zero-base-pair spacing monomer, 11 kDa DNA
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 20
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.9 |
nm |
| VolumePorod |
121 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: None (None-None) Direct Repeat binding site with zero-base-pair spacing (long sequence)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: Q60974 (2059-2325) Nuclear receptor corepressor 1
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Direct Repeat binding site with zero-base-pair spacing (long sequence) monomer, 19 kDa DNA
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Nuclear receptor corepressor 1 monomer, 29 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 12
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
5.4 |
nm |
| Dmax |
20.1 |
nm |
| VolumePorod |
363 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Direct Repeat binding site with one-base-pair spacing
UniProt ID: Q60974 (2059-2325) Nuclear receptor corepressor 1
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Direct Repeat binding site with one-base-pair spacing monomer, 11 kDa DNA
Nuclear receptor corepressor 1 monomer, 29 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 20
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
5.6 |
nm |
| Dmax |
21.8 |
nm |
| VolumePorod |
330 |
nm3 |
|
|
UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Direct Repeat binding site with five-base-pair spacing
UniProt ID: Q60974 (2059-2325) Nuclear receptor corepressor 1
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Direct Repeat binding site with five-base-pair spacing monomer, 13 kDa DNA
Nuclear receptor corepressor 1 monomer, 29 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Dec 12
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
5.7 |
nm |
| Dmax |
21.8 |
nm |
| VolumePorod |
363 |
nm3 |
|
|