UniProt ID: P11416 (82-421) Retinoic acid receptor alpha
UniProt ID: None (None-None) BMS493 (RAR inverse agonist ligand)
UniProt ID: P19793 (126-462) Retinoid X receptor alpha
UniProt ID: None (None-None) Inverted Repeat binding site with zero-base-pair spacing
UniProt ID: Q60974 (2059-2325) Nuclear receptor corepressor 1
|
|
|
|
| Sample: |
Retinoic acid receptor alpha monomer, 39 kDa Mus musculus protein
BMS493 (RAR inverse agonist ligand) monomer, 0 kDa
Retinoid X receptor alpha monomer, 38 kDa Homo sapiens protein
Inverted Repeat binding site with zero-base-pair spacing monomer, 11 kDa DNA
Nuclear receptor corepressor 1 monomer, 29 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris HCl, 75 mM NaCl, 75 mM KCl, 2 mM CHAPS, 4 mM MgCl₂, 12 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Apr 20
|
New structural insights into the control of the retinoic acid receptors RAR/RXR by DNA, ligands, and transcriptional coregulators.
Nucleic Acids Res 53(18) (2025)
Tambones I, Sagar A, Vankova P, Loginov D, Carivenc C, Rochel N, Bourguet W, Man P, Bernadó P, le Maire A
|
| RgGuinier |
5.5 |
nm |
| Dmax |
22.9 |
nm |
| VolumePorod |
257 |
nm3 |
|
|
UniProt ID: Q57YA3 (2-134) Histone H2A
UniProt ID: Q389T1 (2-112) Histone H2B
UniProt ID: Q4GYX7 (2-133) Histone H3, putative
UniProt ID: Q57Z31 (2-100) Histone H4
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 1
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 2
|
|
|
|
| Sample: |
Histone H2A, 28 kDa Trypanosoma brucei brucei … protein
Histone H2B, 25 kDa Trypanosoma brucei brucei … protein
Histone H3, putative, 29 kDa Trypanosoma brucei brucei … protein
Histone H4, 22 kDa Trypanosoma brucei brucei … protein
Widom 601 145 bp DNA - strand 1, 45 kDa DNA
Widom 601 145 bp DNA - strand 2, 45 kDa DNA
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Oct 10
|
Trypanosome histone variants H3.V and H4.V promote nucleosome plasticity in repressed chromatin
|
| RgGuinier |
4.3 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
435 |
nm3 |
|
|
UniProt ID: Q57YA3 (2-134) Histone H2A
UniProt ID: Q389T1 (2-112) Histone H2B
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 1
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 2
UniProt ID: Q387X7 (2-139) Histone H3 variant
UniProt ID: Q587H6 (2-100) Histone H4
|
|
|
|
| Sample: |
Histone H2A, 28 kDa Trypanosoma brucei brucei … protein
Histone H2B, 25 kDa Trypanosoma brucei brucei … protein
Widom 601 145 bp DNA - strand 1, 45 kDa DNA
Widom 601 145 bp DNA - strand 2, 45 kDa DNA
Histone H3 variant, 32 kDa Trypanosoma brucei brucei … protein
Histone H4, 22 kDa Trypanosoma brucei brucei … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Oct 10
|
Trypanosome histone variants H3.V and H4.V promote nucleosome plasticity in repressed chromatin
|
| RgGuinier |
4.5 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
483 |
nm3 |
|
|
UniProt ID: Q57YA3 (2-134) Histone H2A
UniProt ID: Q389T1 (2-112) Histone H2B
UniProt ID: Q57Z31 (2-100) Histone H4
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 1
UniProt ID: None (None-None) Widom 601 145 bp DNA - strand 2
UniProt ID: Q387X7 (2-139) Histone H3 variant
|
|
|
|
| Sample: |
Histone H2A, 28 kDa Trypanosoma brucei brucei … protein
Histone H2B, 25 kDa Trypanosoma brucei brucei … protein
Histone H4, 22 kDa Trypanosoma brucei brucei … protein
Widom 601 145 bp DNA - strand 1, 45 kDa DNA
Widom 601 145 bp DNA - strand 2, 45 kDa DNA
Histone H3 variant, 32 kDa Trypanosoma brucei brucei … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Oct 10
|
Trypanosome histone variants H3.V and H4.V promote nucleosome plasticity in repressed chromatin
Gauri Deak
|
| RgGuinier |
4.4 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
481 |
nm3 |
|
|
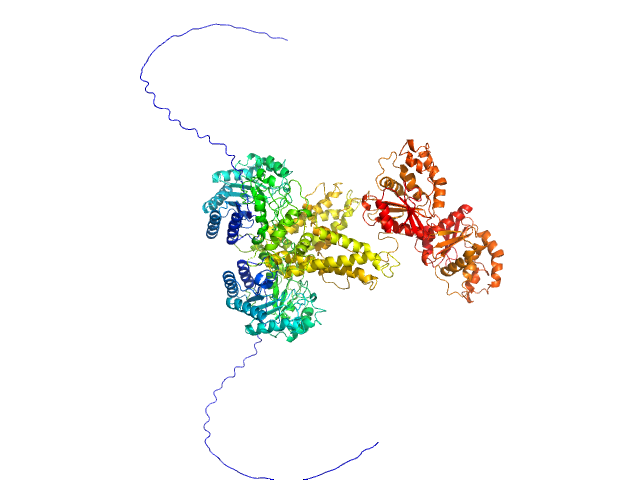
UniProt ID: Q93GI2 (None-None) RTX cytotoxin pro-MbxA without acylation
|
|
|
|
| Sample: |
RTX cytotoxin pro-MbxA without acylation monomer, 100 kDa Moraxella bovis protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 10mM CaCl2, pH: 7.8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 May 9
|
Acylation of the RTX toxin MbxA stimulates host membrane disruption through a specific interaction with cholesterol
Biochimica et Biophysica Acta (BBA) - Biomembranes :184487 (2025)
Chacko F, Ganz S, Pfitzer-Bilsing A, Hänsch S, Westhoff P, Weidtkamp-Peters S, Smits S, Exterkate M, Schmitt L
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
154 |
nm3 |
|
|
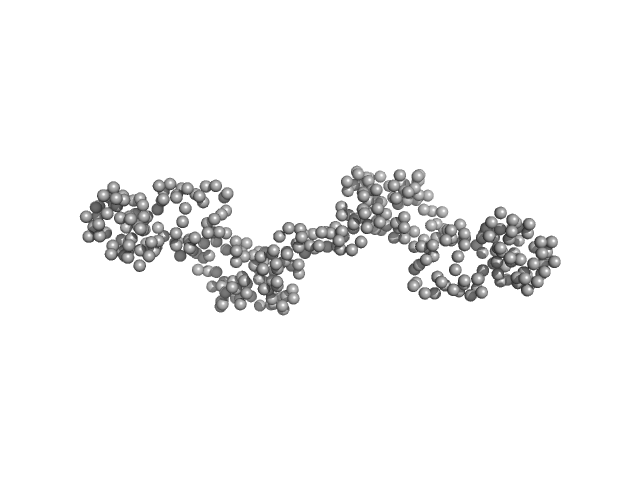
UniProt ID: B8BQS1 (None-None) CP12 domain-containing protein
|
|
|

|
| Sample: |
CP12 domain-containing protein dimer, 34 kDa Thalassiosira pseudonana protein
|
| Buffer: |
30 mM Tris, 50 mM NaCl, 2 mM EDTA, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Feb 23
|
A new type of flexible CP12 protein in the marine diatom Thalassiosira pseudonana
Cell Communication and Signaling 19(1) (2021)
Shao H, Huang W, Avilan L, Receveur-Bréchot V, Puppo C, Puppo R, Lebrun R, Gontero B, Launay H
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.5 |
nm |
|
|
UniProt ID: Q14764 (1-893) Major vault protein
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDXJ3_fit1_model1.png)
|
| Sample: |
Major vault protein, 7747 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Jun 8
|
Structural flexibility of the human vault particle revealed by high-resolution cryo-EM and molecular dynamics simulations
Jana Aupič
|
| RgGuinier |
16.3 |
nm |
| Dmax |
65.0 |
nm |
| VolumePorod |
16650 |
nm3 |
|
|
UniProt ID: Q93IQ4 (151-509) Alkaline serine protease
|
|
|
|
| Sample: |
Alkaline serine protease monomer, 36 kDa Stenotrophomonas maltophilia protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Jun 10
|
Unveiling the structure, function and dynamics of StmPr1 in Stenotrophomonas maltophilia virulence.
Sci Rep 15(1):20193 (2025)
Sommer M, Negm A, Outzen L, Windhorst S, Gabdulkhakov A, Weber W, Betzel C
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.5 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
UniProt ID: Q93IQ4 (151-617) Alkaline serine protease (Δ1-150)
|
|
|
|
| Sample: |
Alkaline serine protease (Δ1-150) monomer, 47 kDa Stenotrophomonas maltophilia protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Nov 25
|
Unveiling the structure, function and dynamics of StmPr1 in Stenotrophomonas maltophilia virulence.
Sci Rep 15(1):20193 (2025)
Sommer M, Negm A, Outzen L, Windhorst S, Gabdulkhakov A, Weber W, Betzel C
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
112 |
nm3 |
|
|
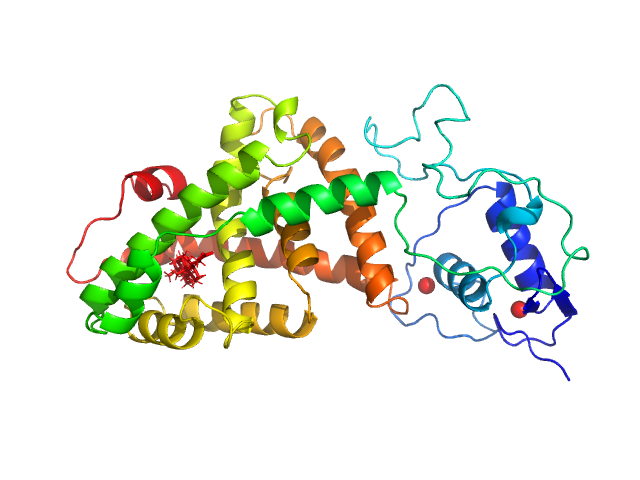
UniProt ID: Q96RI1-1 (110-466) Isoform 3 of Bile acid receptor
|
|
|
|
| Sample: |
Isoform 3 of Bile acid receptor monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 5 % glycerol, 1 mM DTT, 20 µM CDCA, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Pennsylvania State University on 2024 May 9
|
DNA induces non-canonical dimerization of the farnesoid X receptor.
Nucleic Acids Res 54(4) (2026)
Khan SH, Yennawar N, Okafor CD
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
75 |
nm3 |
|
|