UniProt ID: O07411 (1-554) Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase)
|
|
|
|
| Sample: |
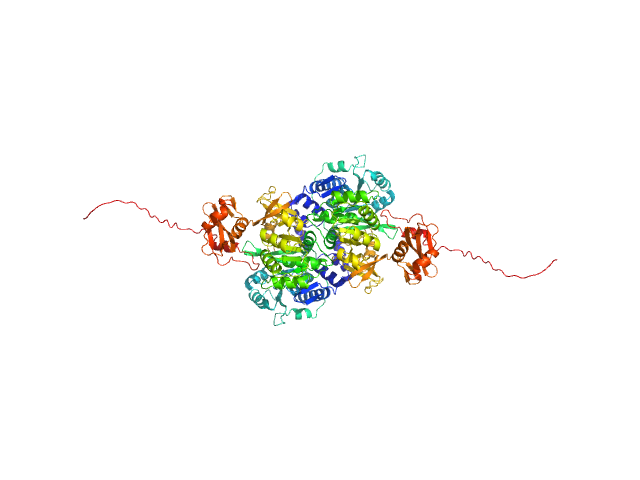
Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase) dimer, 123 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM HEPES, 500 mM NaCl, 5 mM MgCl2, 1 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Oct 20
|
Structural enzymological studies of the long chain fatty acyl-CoA synthetase FadD5 from the mce1 operon of Mycobacterium tuberculosis.
Biochem Biophys Res Commun 769:151960 (2025)
Rahman MA, Dalwani S, Venkatesan R
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.3 |
nm |
| VolumePorod |
221 |
nm3 |
|
|
UniProt ID: O07411 (1-554) Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase)
|
|
|
|
| Sample: |
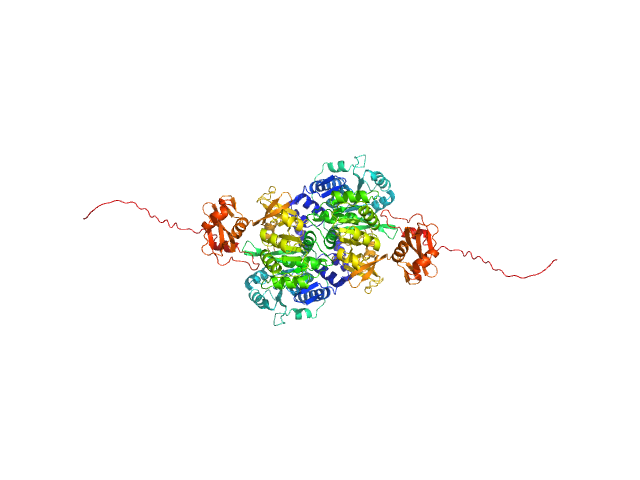
Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase) dimer, 123 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM HEPES, 500 mM NaCl, 5 mM MgCl2, 1 mM β-mercaptoethanol, 2mM ATP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Oct 20
|
Structural enzymological studies of the long chain fatty acyl-CoA synthetase FadD5 from the mce1 operon of Mycobacterium tuberculosis.
Biochem Biophys Res Commun 769:151960 (2025)
Rahman MA, Dalwani S, Venkatesan R
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.1 |
nm |
| VolumePorod |
213 |
nm3 |
|
|
UniProt ID: O07411 (1-554) Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase)
|
|
|

|
| Sample: |
Probable fatty-acid-CoA ligase FadD5 (Fatty-acid-CoA synthetase) (Fatty-acid-CoA synthase) dimer, 123 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM HEPES, 500 mM NaCl, 5 mM MgCl2, 1 mM β-mercaptoethanol, 2 mM coenzyme A (CoA), pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Oct 20
|
Structural enzymological studies of the long chain fatty acyl-CoA synthetase FadD5 from the mce1 operon of Mycobacterium tuberculosis.
Biochem Biophys Res Commun 769:151960 (2025)
Rahman MA, Dalwani S, Venkatesan R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
210 |
nm3 |
|
|
UniProt ID: E9RJ23 (1-232) Relaxase
|
|
|
|
| Sample: |
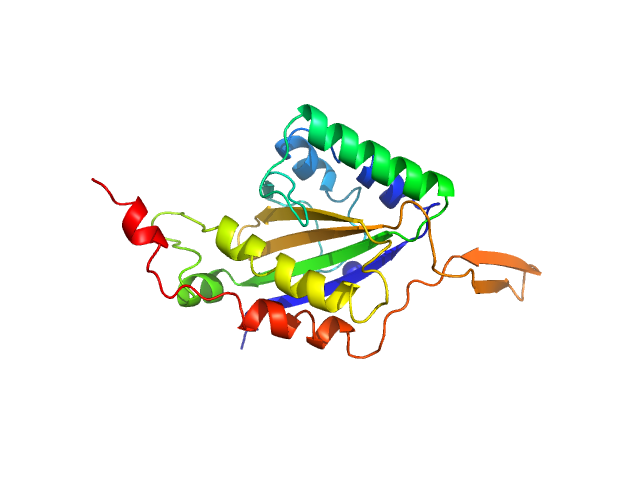
Relaxase monomer, 29 kDa Bacillus subtilis subsp. … protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BL11 - NCD, ALBA on 2019 Dec 3
|
Activation and dimerization of the relaxase from the Bacillus subtilis conjugative plasmid pLS20 by DNA-induced cross-over of the catalytic residue
Isidro Crespo
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
UniProt ID: E9RJ23 (1-232) Relaxase
UniProt ID: None (None-None) core region, single stranded
|
|
|
|
| Sample: |
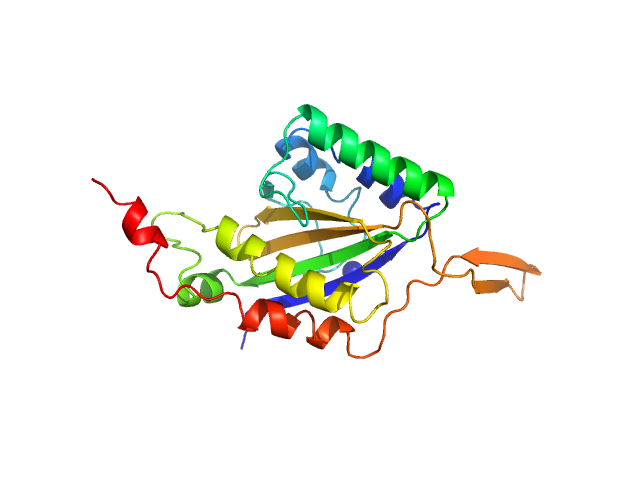
Relaxase dimer, 57 kDa Bacillus subtilis subsp. … protein
Core region, single stranded dimer, 12 kDa Bacillus subtilis DNA
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BL11 - NCD, ALBA on 2019 Dec 3
|
Activation and dimerization of the relaxase from the Bacillus subtilis conjugative plasmid pLS20 by DNA-induced cross-over of the catalytic residue
Isidro Crespo
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
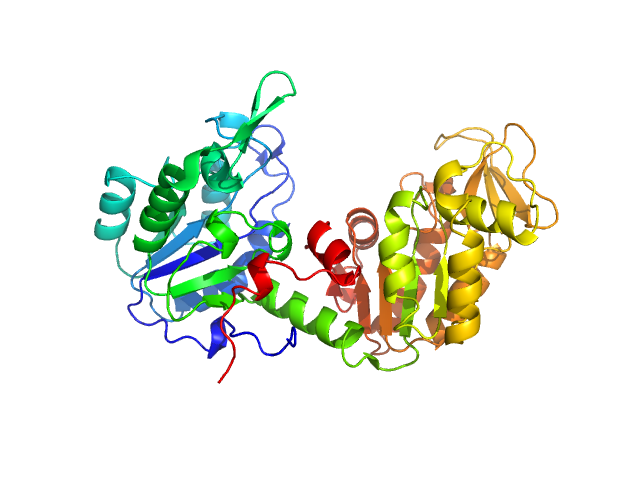
UniProt ID: A0A0W0EGC5 (1-416) Phosphoglycerate kinase
|
|
|
|
| Sample: |
Phosphoglycerate kinase monomer, 45 kDa Nakaseomyces glabratus protein
|
| Buffer: |
50 mM sodium phosphate, 500 mM NaCl, 500 mM imidazole, 1 mM phenylmethylsulfonyl fluoride (PMSF), pH: 8.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2025 Feb 13
|
Integrative Structural Characterization of Candida glabrata
Phosphoglycerate Kinase by Small-Angle X-ray Scattering and AlphaFold: Implications for Therapeutic Targeting in Candidiasis
ACS Omega (2026)
Cuéllar-Cruz M, Maqueda Cabrera E, Siliqi D, Moreno A
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
79 |
nm3 |
|
|
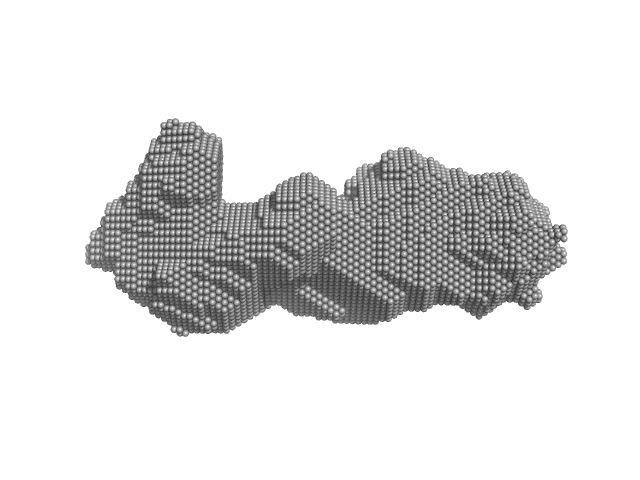
UniProt ID: Q504U8 (651-993) Receptor protein-tyrosine kinase (duplication mutant)
|
|
|

|
| Sample: |
Receptor protein-tyrosine kinase (duplication mutant) monomer, 80 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 250 mM NaCl, 250 mM KCl, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax-007HF, Yale University on 2022 Mar 22
|
The role of kinase domain dimerization in EGFR activation
Zaritza Petrova
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
135 |
nm3 |
|
|
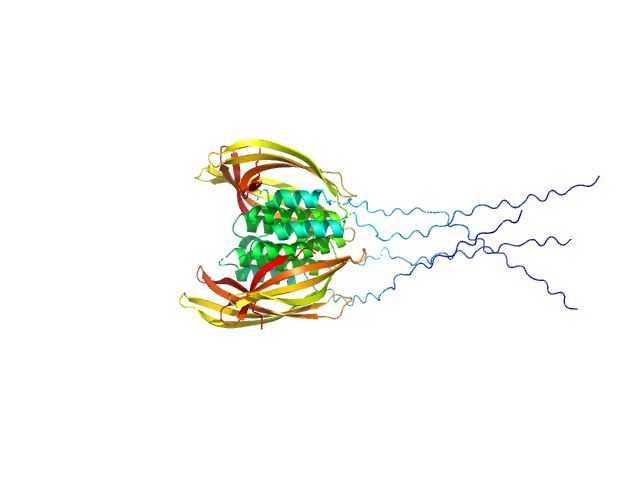
UniProt ID: O07419 (107-213) Probable conserved Mce associated membrane protein
|
|
|
|
| Sample: |
Probable conserved Mce associated membrane protein tetramer, 59 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50mM Tris, 500mM NaCl, pH: 8.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 10
|
Insight into the structure and interactions of the M. tuberculosis
Mce-associated membrane proteins Mam1A-1D
(2025)
Hynönen M, Perumal P, Hynönen N, Doutch J, Ma K, Venkatesan R
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
177 |
nm3 |
|
|
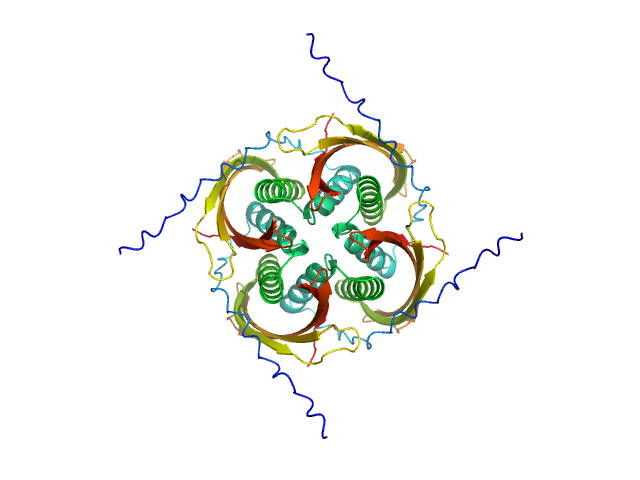
UniProt ID: O07419 (107-213) Probable conserved Mce associated membrane protein
|
|
|
|
| Sample: |
Probable conserved Mce associated membrane protein tetramer, 59 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris, 500 mM NaCl, D-C12E9, pH: 8.5 |
| Experiment: |
SANS
data collected at SANS2D, ISIS Neutron and Muon Source on 2019 Dec 10
|
Insight into the structure and interactions of the M. tuberculosis
Mce-associated membrane proteins Mam1A-1D
(2025)
Hynönen M, Perumal P, Hynönen N, Doutch J, Ma K, Venkatesan R
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
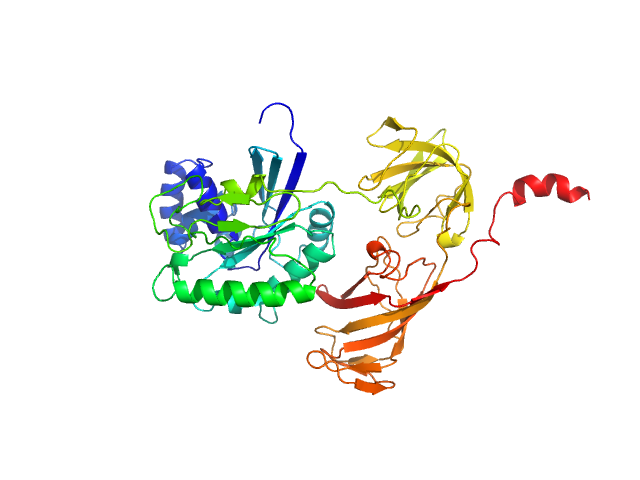
UniProt ID: Q05639 (1-463) Mammalian translation elongation factor eEF1A2
|
|
|
|
| Sample: |
Mammalian translation elongation factor eEF1A2 monomer, 50 kDa protein
|
| Buffer: |
25 mM Tris HCl, 150 mM NaCl, 6 mM βME, 20% glycerol, 0.01mM GDP,, pH: 7.5 |
| Experiment: |
SAXS
data collected at Bruker Nanostar w Excillum source, Department of Chemistry, iNANO building, Aarhus Uinversity on 2023 Mar 21
|
Dynamics and structural features of the eEF1A1 and eEF1A2 paralogs
Nucleic Acids Research 53(21) (2025)
Novosylna O, Shalak V, Dąbrowska K, Patmanidis I, Lozhko D, Bondarchuk T, Schiøtt B, Pedersen J, Knudsen C, Nissen P, Dadlez M, Negrutskii B
|
| RgGuinier |
2.6 |
nm |
| Dmax |
97.1 |
nm |
| VolumePorod |
60 |
nm3 |
|
|