UniProt ID: P04198 (1-100) N-myc proto-oncogene protein, residues 1-100
UniProt ID: O14965 (122-403) Aurora kinase A mutant C290A:C393A
|
|
|
|
| Sample: |
N-myc proto-oncogene protein, residues 1-100 monomer, 11 kDa Homo sapiens protein
Aurora kinase A mutant C290A:C393A monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Dec 9
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.4 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
UniProt ID: B2FN79 (None-None) Stenotrophomonas maltophilia threonyl tRNA synthetase
|
|
|
|
| Sample: |
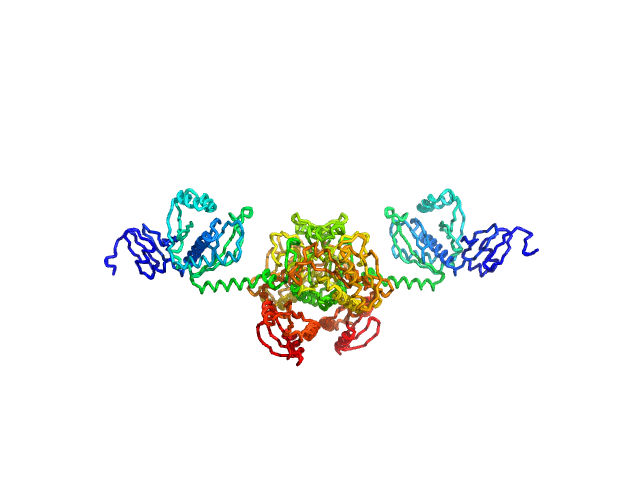
Stenotrophomonas maltophilia threonyl tRNA synthetase dimer, 146 kDa Stenotrophomonas maltophilia (strain … protein
|
| Buffer: |
20 mM HEPES, 0.2 M NaCl, 1.5% v/v glycerol, and 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2024 Nov 7
|
Crystal structure of a complete threonyl tRNA synthetase: toggle clamp model?
Biochemical and Biophysical Research Communications :152677 (2025)
Tran T, Fenwick M, Edwards T, Dranow D, Abendroth J, Shek R, Tillery L, Craig J, Van Voorhis W, Myler P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.2 |
nm |
|
|
UniProt ID: B2FN79 (None-None) Stenotrophomonas maltophilia threonyl tRNA synthetase
|
|
|
|
| Sample: |
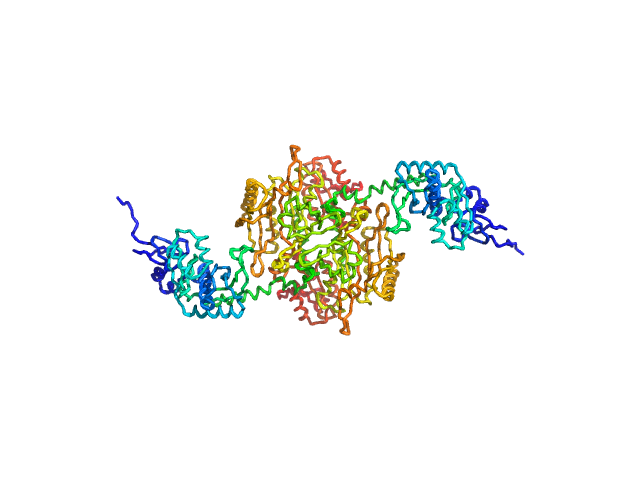
Stenotrophomonas maltophilia threonyl tRNA synthetase dimer, 146 kDa Stenotrophomonas maltophilia (strain … protein
|
| Buffer: |
20 mM HEPES, 0.2 M NaCl, 1.5% v/v glycerol, and 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2024 Nov 7
|
Crystal structure of a complete threonyl tRNA synthetase: toggle clamp model?
Biochemical and Biophysical Research Communications :152677 (2025)
Tran T, Fenwick M, Edwards T, Dranow D, Abendroth J, Shek R, Tillery L, Craig J, Van Voorhis W, Myler P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.2 |
nm |
|
|
UniProt ID: B2FN79 (None-None) Stenotrophomonas maltophilia threonyl tRNA synthetase
|
|
|
|
| Sample: |
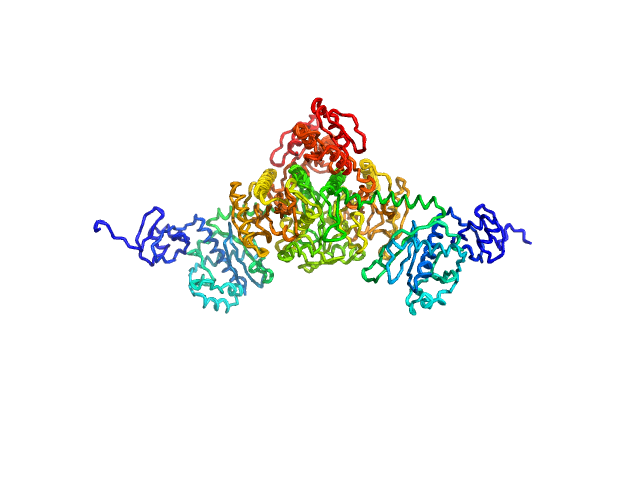
Stenotrophomonas maltophilia threonyl tRNA synthetase dimer, 146 kDa Stenotrophomonas maltophilia (strain … protein
|
| Buffer: |
20 mM HEPES, 0.2 M NaCl, 1.5% v/v glycerol, and 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2024 Nov 7
|
Crystal structure of a complete threonyl tRNA synthetase: toggle clamp model?
Biochemical and Biophysical Research Communications :152677 (2025)
Tran T, Fenwick M, Edwards T, Dranow D, Abendroth J, Shek R, Tillery L, Craig J, Van Voorhis W, Myler P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.2 |
nm |
|
|
UniProt ID: B2FN79 (None-None) Stenotrophomonas maltophilia threonyl tRNA synthetase
|
|
|
|
| Sample: |
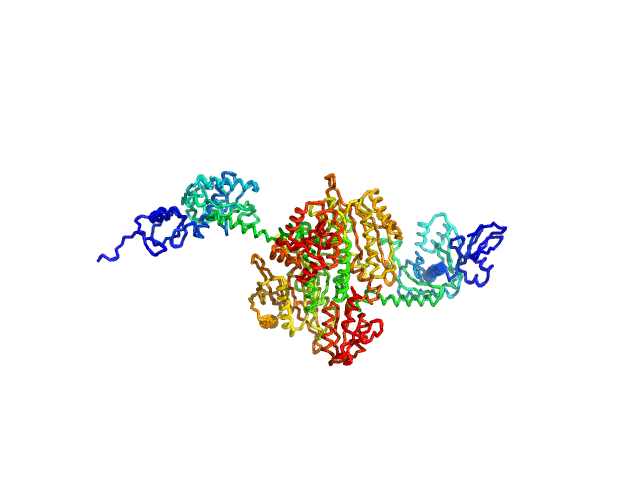
Stenotrophomonas maltophilia threonyl tRNA synthetase dimer, 146 kDa protein
|
| Buffer: |
20 mM HEPES, 0.2 M NaCl, 1.5% v/v glycerol, and 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2024 Nov 7
|
Crystal structure of a complete threonyl tRNA synthetase: toggle clamp model?
Biochemical and Biophysical Research Communications :152677 (2025)
Tran T, Fenwick M, Edwards T, Dranow D, Abendroth J, Shek R, Tillery L, Craig J, Van Voorhis W, Myler P
|
|
|
UniProt ID: Q13148 (1-277) TAR DNA-binding protein 43
|
|
|
|
| Sample: |
TAR DNA-binding protein 43 monomer, 63 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM KCl, 2 mM TCEP, pH: 7.6 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Nov 30
|
From TDP-43/RNA complex formation to disease-linked TDP-43 aggregation through a structural and cellular approach.
Nat Commun (2026)
Feng Y, Joshi V, Pankivskyi S, Clément MJ, Rengifo-Gonzalez JC, Thureau A, Pastré D, Bouhss A
|
| RgGuinier |
4.0 |
nm |
| Dmax |
16.5 |
nm |
| VolumePorod |
135 |
nm3 |
|
|
UniProt ID: P0DKX7 (1529-1680) Repeats-in-toxin domain Block V of adenylate cyclase toxin
|
|
|
|
| Sample: |
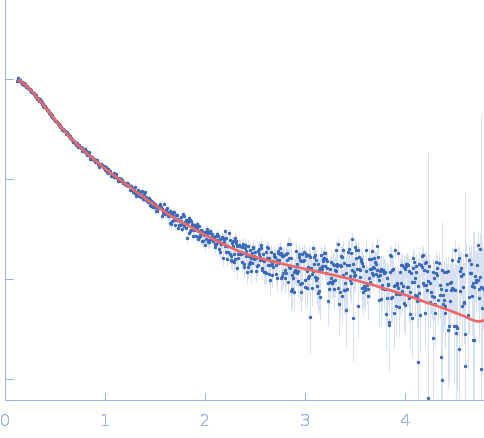
Repeats-in-toxin domain Block V of adenylate cyclase toxin monomer, 19 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM Tris, 5 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2024 Apr 1
|
Ion-selective conformational stabilization of a disordered repeats-in-toxin protein domain.
Biophys J (2025)
Gudinas AP, Shambharkar GM, Chang MP, Fernández D, Matsui T, Mai DJ
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.4 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
UniProt ID: P0DKX7 (1529-1680) Repeats-in-toxin domain Block V of adenylate cyclase toxin
|
|
|
|
| Sample: |
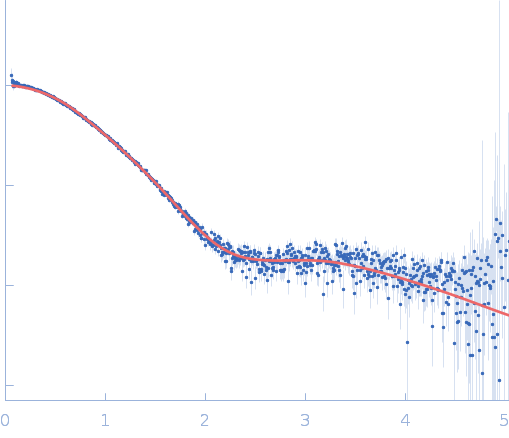
Repeats-in-toxin domain Block V of adenylate cyclase toxin monomer, 19 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM Tris, 5 mM DTT, 1 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2024 Jun 28
|
Ion-selective conformational stabilization of a disordered repeats-in-toxin protein domain.
Biophys J (2025)
Gudinas AP, Shambharkar GM, Chang MP, Fernández D, Matsui T, Mai DJ
|
| RgGuinier |
2.0 |
nm |
| Dmax |
8.1 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
UniProt ID: P0DKX7 (1529-1680) Repeats-in-toxin domain Block V of adenylate cyclase toxin
|
|
|
|
| Sample: |
Repeats-in-toxin domain Block V of adenylate cyclase toxin monomer, 19 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM Tris, 5 mM DTT, 10 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2024 Dec 12
|
Ion-selective conformational stabilization of a disordered repeats-in-toxin protein domain.
Biophys J (2025)
Gudinas AP, Shambharkar GM, Chang MP, Fernández D, Matsui T, Mai DJ
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
UniProt ID: P0DKX7 (1529-1680) Repeats-in-toxin domain Block V of adenylate cyclase toxin
|
|
|
|
| Sample: |
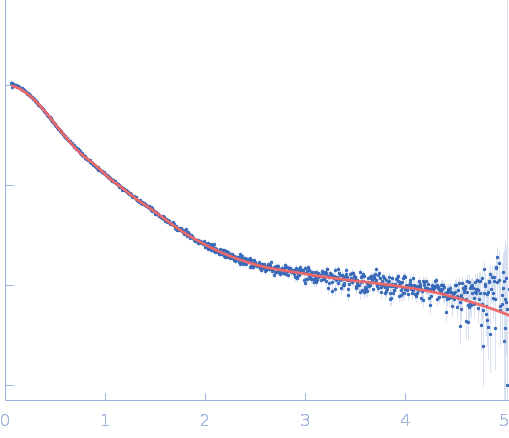
Repeats-in-toxin domain Block V of adenylate cyclase toxin monomer, 19 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM Tris, 5 mM DTT, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2024 Jun 28
|
Ion-selective conformational stabilization of a disordered repeats-in-toxin protein domain.
Biophys J (2025)
Gudinas AP, Shambharkar GM, Chang MP, Fernández D, Matsui T, Mai DJ
|
| RgGuinier |
3.5 |
nm |
| Dmax |
14.9 |
nm |
| VolumePorod |
49 |
nm3 |
|
|