UniProt ID: Q92560 (1-238) Ubiquitin C-terminal hydrolase (UCH) domain of BAP1 wild type
|
|
|
|
| Sample: |
Ubiquitin C-terminal hydrolase (UCH) domain of BAP1 wild type monomer, 27 kDa protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3 mM CaCl2, 0.02% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Jun 23
|
Allosteric network of dynamic coupling within BAP1-UCH revealed by methyl NMR
Yong-Sheng Wang
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
UniProt ID: Q92560 (1-238) Ubiquitin C-terminal hydrolase (UCH) domain of BAP1 L49V
|
|
|
|
| Sample: |
Ubiquitin C-terminal hydrolase (UCH) domain of BAP1 L49V monomer, 27 kDa protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.02% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Jun 23
|
Allosteric network of dynamic coupling within BAP1-UCH revealed by methyl NMR
Yong-Sheng Wang
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.7 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
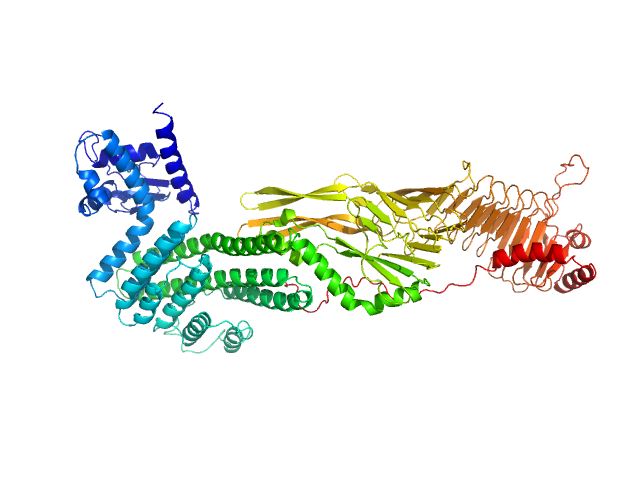
UniProt ID: O60502 (11-766) Protein O-GlcNAcase
|
|
|
|
| Sample: |
Protein O-GlcNAcase dimer, 163 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris buffer, 150 mM NaCl, and 0.5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at Bruker Nanostar w Excillum source, Department of Chemistry, iNANO building, Aarhus Uinversity on 2023 Jun 22
|
Multi-domain O-GlcNAcase structures reveal allosteric regulatory mechanisms
Nature Communications 16(1) (2025)
Hansen S, Bartual S, Yuan H, Raimi O, Gorelik A, Ferenbach A, Lytje K, Pedersen J, Drace T, Boesen T, van Aalten D
|
| RgGuinier |
5.4 |
nm |
| Dmax |
18.9 |
nm |
| VolumePorod |
443 |
nm3 |
|
|
UniProt ID: Q9UQL6 (655-1122) Histone deacetylase 5
|
|
|
|
| Sample: |
Histone deacetylase 5 monomer, 51 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM KCl, 1 mM DTT,, pH: 7.5 |
| Experiment: |
SAXS
data collected at Xenocs Xeuss 2.0 Q-Xoom, Center for Structural Studies, Heinrich-Heine-University on 2024 Feb 29
|
In vitro characterization of the catalytic domain of human histone deacetylase 5
Scientific Reports (2026)
Mammen C, Hornung F, Anzenhofer C, Schumacher J, Reiners J, Li J, Mazzone F, Bilsing F, Kassack M, Kurz T, Smits S
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
UniProt ID: P0DTD1 (6465-6798) Non structural Protein 15
|
|
|
|
| Sample: |
Non structural Protein 15 monomer, 251 kDa Severe acute respiratory … protein
|
| Buffer: |
20mM HEPES, 150mM Nacl, pH: 7.5 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2024 Jun 14
|
Non Structural Protein 15 in Complex with RNA 9mer Oligo of SARS CoV 2
Hira Singh Gariya
|
| RgGuinier |
4.7 |
nm |
| Dmax |
13.8 |
nm |
| VolumePorod |
356 |
nm3 |
|
|
UniProt ID: Q9ET54 (1022-1257) Palladin Ig3-45Linker_Ig4
|
|
|
|
| Sample: |
Palladin Ig3-45Linker_Ig4 monomer, 22 kDa Mus musculus protein
|
| Buffer: |
HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2023 Aug 29
|
The role of linker length and composition in actin binding and bundling by palladin.
Biochem J 483(3):301-318 (2026)
Sargent RA, Bradford CW, Hughes LM, Ta NH, Limpiado MJ, Vattepu R, Beck MR
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
UniProt ID: P04198 (1-69) N-myc proto-oncogene protein, residues 1-69
|
|
|
|
| Sample: |
N-myc proto-oncogene protein, residues 1-69 monomer, 8 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Nov 28
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
2.4 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
12 |
nm3 |
|
|
UniProt ID: P04198 (1-100) N-myc proto-oncogene protein, residues 1-100
|
|
|
|
| Sample: |
N-myc proto-oncogene protein, residues 1-100 monomer, 11 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Nov 28
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
3.0 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
UniProt ID: O14965 (122-403) Aurora kinase A mutant C290A:C393A
|
|
|
|
| Sample: |
Aurora kinase A mutant C290A:C393A monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Nov 28
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
2.2 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
59 |
nm3 |
|
|
UniProt ID: P04198 (1-69) N-myc proto-oncogene protein, residues 1-69
UniProt ID: O14965 (122-403) Aurora kinase A mutant C290A:C393A
|
|
|
|
| Sample: |
N-myc proto-oncogene protein, residues 1-69 monomer, 8 kDa Homo sapiens protein
Aurora kinase A mutant C290A:C393A monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Nov 28
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
62 |
nm3 |
|
|