UniProt ID: Q6PDM4 (109-267) Interactor of HORMAD1 protein 1 (M198V, V230I)
|
|
|
|
| Sample: |
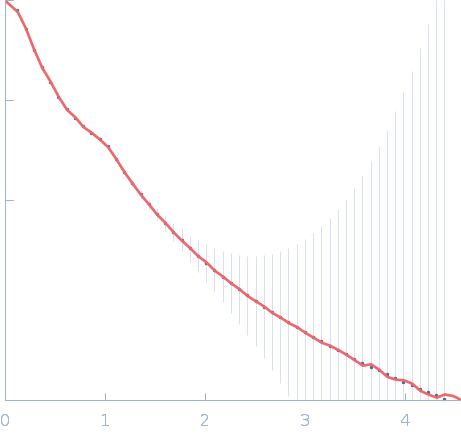
Interactor of HORMAD1 protein 1 (M198V, V230I) tetramer, 127 kDa Mus musculus protein
|
| Buffer: |
25 mM HEPES-NaOH, 500 mM NaCl, 5 mM EDTA, 5% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2023 Apr 14
|
Evolutionary conservation of the structure and function of meiotic Rec114-Mei4 and Mer2 complexes.
Genes Dev 37(11-12):535-553 (2023)
Daccache D, De Jonge E, Liloku P, Mechleb K, Haddad M, Corthaut S, Sterckx YG, Volkov AN, Claeys Bouuaert C
|
| RgGuinier |
7.2 |
nm |
| Dmax |
25.5 |
nm |
|
|
UniProt ID: Q86YC2 (1-200) Partner and localizer of BRCA2
|
|
|
|
| Sample: |
Partner and localizer of BRCA2 dimer, 46 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 160 mM NaCl, 0.5 mM TCEP, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 11
|
The strand exchange domain of tumor suppressor PALB2 is intrinsically disordered and promotes oligomerization-dependent DNA compaction.
iScience 27(12):111259 (2024)
Kyriukha Y, Watkins MB, Redington JM, Chintalapati N, Ganti A, Dastvan R, Uversky VN, Hopkins JB, Pozzi N, Korolev S
|
| RgGuinier |
4.7 |
nm |
| Dmax |
21.3 |
nm |
| VolumePorod |
197 |
nm3 |
|
|
UniProt ID: O75400 (141-236) Pre-mRNA-processing factor 40 homolog A
UniProt ID: Q15637 (575-590) Splicing factor 1
|
|
|
|
| Sample: |
Pre-mRNA-processing factor 40 homolog A monomer, 11 kDa Homo sapiens protein
Splicing factor 1 monomer, 2 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 100 mM NaCl, 0.5 mM DTT, pH: 6.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Sep 19
|
Intramolecular autoinhibition regulates the selectivity of PRPF40A tandem WW domains for proline-rich motifs.
Nat Commun 15(1):3888 (2024)
Martínez-Lumbreras S, Träger LK, Mulorz MM, Payr M, Dikaya V, Hipp C, König J, Sattler M
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.9 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
UniProt ID: Q8N5S9 (67-481) Calcium/calmodulin-dependent protein kinase kinase 1 (M474L)
|
|
|
|
| Sample: |
Calcium/calmodulin-dependent protein kinase kinase 1 (M474L) monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM TCEP, 3% (w/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
14-3-3 protein inhibits CaMKK1 by blocking the kinase active site with its last two C-terminal helices.
Protein Sci :e4805 (2023)
Petrvalska O, Honzejkova K, Koupilova N, Herman P, Obsilova V, Obsil T
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
114 |
nm3 |
|
|
UniProt ID: Q9WTS6-1 (343-2715) Isoform A1B1 of Teneurin-3
|
|
|
|
| Sample: |
Isoform A1B1 of Teneurin-3 dimer, 545 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM EDTA, pH: 7.8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Sep 10
|
Alternative splicing controls teneurin-3 compact dimer formation for neuronal recognition
Nature Communications 15(1) (2024)
Gogou C, Beugelink J, Frias C, Kresik L, Jaroszynska N, Drescher U, Janssen B, Hindges R, Meijer D
|
| RgGuinier |
8.8 |
nm |
| Dmax |
35.0 |
nm |
| VolumePorod |
1035 |
nm3 |
|
|
UniProt ID: M4N7P1 (1-482) Variant surface glycoprotein 3.1
|
|
|
|
| Sample: |
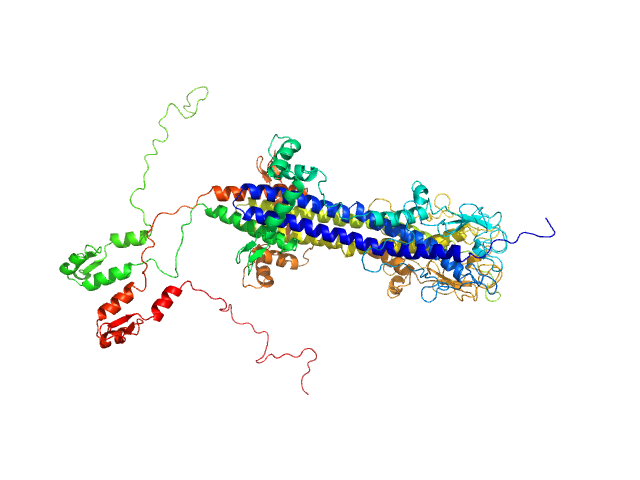
Variant surface glycoprotein 3.1 dimer, 103 kDa Trypanosoma brucei gambiense protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Sep 8
|
Beyond the VSG Layer: Exploring the Role of Intrinsic Disorder in the Invariant Surface Glycoproteins of African Trypanosomes.
PLoS Pathog 20(4):e1012186 (2024)
Sülzen H, Volkov AN, Geens R, Zahedifard F, Stijlemans B, Zoltner M, Magez S, Sterckx YG, Zoll S
|
| RgGuinier |
4.7 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
149 |
nm3 |
|
|
UniProt ID: P0DTC9 (1-365) Nucleoprotein
|
|
|
|
| Sample: |
Nucleoprotein tetramer, 163 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 13
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
5.8 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
181 |
nm3 |
|
|
UniProt ID: P0DN75 (2-132) Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Jun 8
|
A hidden property of the iron-sulfur protein in the mononuclear iron-bound state: species-dependent structural ordering induced by magnetic fields.
FEBS J (2025)
Arai S, Soga S, Hirai M, Kobayashi R, Masai H, Kimura K, Maeda K, Nagashima H
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
UniProt ID: J3QLL0 (2-73) importin-beta-binding domain of importin subunit alpha-1 labelled with unnatural amino acid diBrK
|
|
|
|
| Sample: |
Importin-beta-binding domain of importin subunit alpha-1 labelled with unnatural amino acid diBrK monomer, 8 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline containing 0.5 M urea, 0.3 M KCl, 5 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Dec 19
|
A genetically encoded anomalous SAXS ruler to probe the dimensions of intrinsically disordered proteins.
Proc Natl Acad Sci U S A 121(50):e2415220121 (2024)
Yu M, Gruzinov AY, Ruan H, Scheidt T, Chowdhury A, Giofrè S, Mohammed ASA, Caria J, Sauter PF, Svergun DI, Lemke EA
|
| RgGuinier |
2.6 |
nm |
| Dmax |
13.8 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
UniProt ID: P08311 (22-245) Cathepsin G
UniProt ID: A0A0H3JUK5 (32-146) MAP domain-containing protein
UniProt ID: P08246 (31-249) Neutrophil elastase
|
|
|
|
| Sample: |
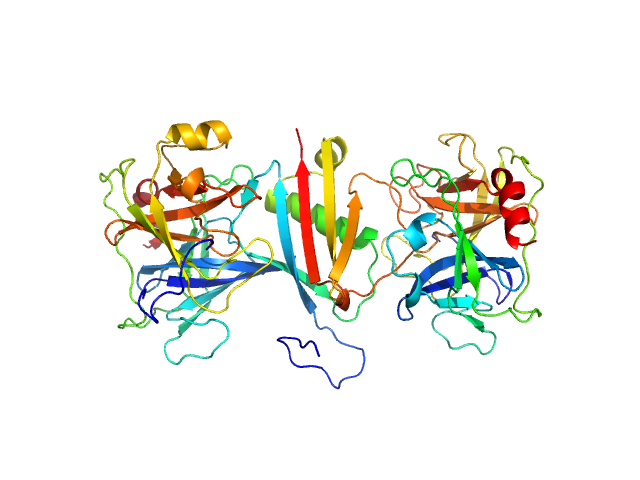
Cathepsin G monomer, 25 kDa Homo sapiens protein
MAP domain-containing protein monomer, 13 kDa Staphylococcus aureus (strain … protein
Neutrophil elastase monomer, 23 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Jun 3
|
S. aureus Eap is a polyvalent inhibitor of neutrophil serine proteases.
J Biol Chem 300(9):107627 (2024)
Mishra N, Gido CD, Herdendorf TJ, Hammel M, Hura GL, Fu ZQ, Geisbrecht BV
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
77 |
nm3 |
|
|