UniProt ID: Q92974-1 (28-582) Isoform 1 of Rho guanine nucleotide exchange factor 2
|
|
|
|
| Sample: |
Isoform 1 of Rho guanine nucleotide exchange factor 2 monomer, 64 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris-Cl, 150mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Feb 19
|
Structural basis of microtubule-mediated signal transduction.
Cell (2025)
Choi SR, Blum TB, Giono M, Roy B, Vakonakis I, Schmid D, Oelgarth N, Ranganathan A, Gossert AD, Shivashankar GV, Zippelius A, Steinmetz MO
|
|
|
UniProt ID: Q8N4Q1 (1-142) Mitochondrial intermembrane space import and assembly protein 40
|
|
|
|
| Sample: |
Mitochondrial intermembrane space import and assembly protein 40 monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Oct 26
|
NADH-bound AIF activates the mitochondrial CHCHD4/MIA40 chaperone by a substrate-mimicry mechanism.
EMBO J (2025)
Brosey CA, Shen R, Tainer JA
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
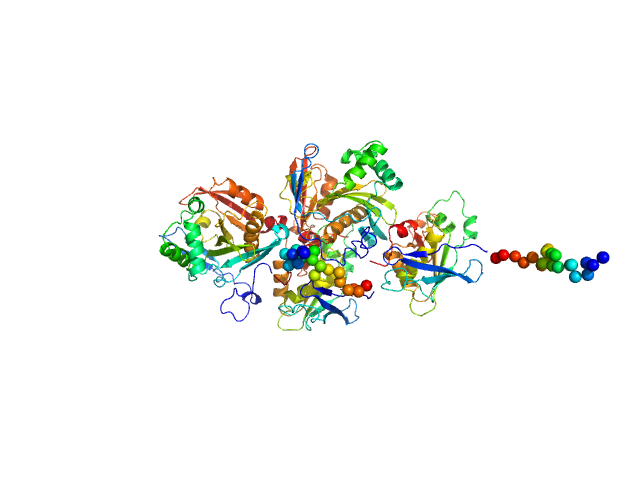
UniProt ID: Q4WF55 (1-443) N(5)-hydroxyornithine:cis-anhydromevalonyl coenzyme A-N(5)-transacylase sidF (Δ444-462)
|
|
|
|
| Sample: |
N(5)-hydroxyornithine:cis-anhydromevalonyl coenzyme A-N(5)-transacylase sidF (Δ444-462) dimer, 107 kDa Aspergillus fumigatus (strain … protein
|
| Buffer: |
50 mM Tris, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2023 Jan 18
|
SidF, a dual substrate N5-acetyl-N5-hydroxy-L-ornithine transacetylase involved in Aspergillus fumigatus siderophore biosynthesis
Journal of Structural Biology: X 11:100119 (2025)
Poonsiri T, Stransky J, Demitri N, Haas H, Cianci M, Benini S
|
| RgGuinier |
3.5 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
189 |
nm3 |
|
|
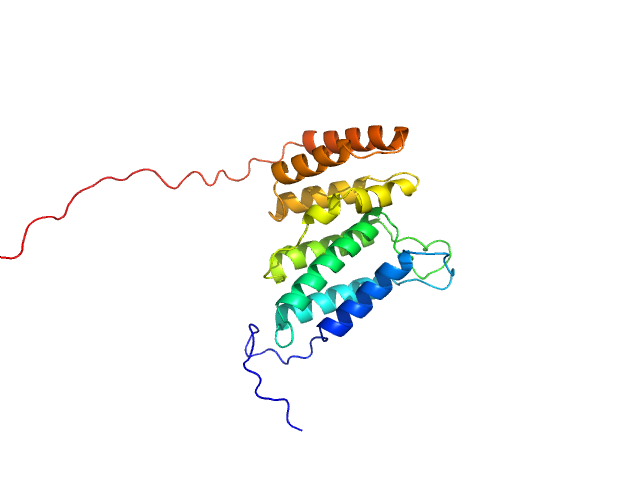
UniProt ID: Q8IWY9 (1005-1227) Codanin-1
|
|
|
|
| Sample: |
Codanin-1 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Sep 29
|
Anemia-associated mutations disrupt the CDIN1-Codanin1 complex in inherited congenital dyserythropoietic anemia I (CDA-I) disease.
FEBS J (2026)
Stojaspal M, Brom T, Nečasová I, Janovič T, Veverka P, Verma N, Uhrík L, Hernychova L, Hofr C
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
UniProt ID: P30681 (1-210) High mobility group protein B2
UniProt ID: None (None-None) Double stranded DNA 20mer
|
|
|
|
| Sample: |
High mobility group protein B2 monomer, 26 kDa Mus musculus protein
Double stranded DNA 20mer monomer, 12 kDa DNA
|
| Buffer: |
Tris 50mM, NaCl 150mM, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Apr 27
|
High mobility group protein B2 (HMGB2)
Chun I Tu
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.2 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
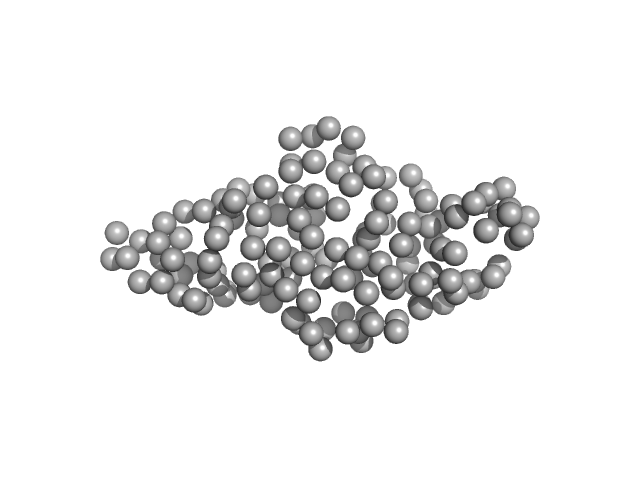
UniProt ID: P84243 (1-40) Histone H3.3
UniProt ID: Q9UBC3 (206-355) DNA (cytosine-5)-methyltransferase 3B Pro-Trp-Trp-Pro (PWWP) domain
|
|
|

|
| Sample: |
Histone H3.3 monomer, 4 kDa Homo sa protein
DNA (cytosine-5)-methyltransferase 3B Pro-Trp-Trp-Pro (PWWP) domain monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM TCEP, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Sep 17
|
Histone modification-driven structural remodeling unleashes DNMT3B in DNA methylation.
Sci Adv 11(13):eadu8116 (2025)
Cho CC, Huang HH, Jiang BC, Yang WZ, Chen YN, Yuan HS
|
| RgGuinier |
1.8 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
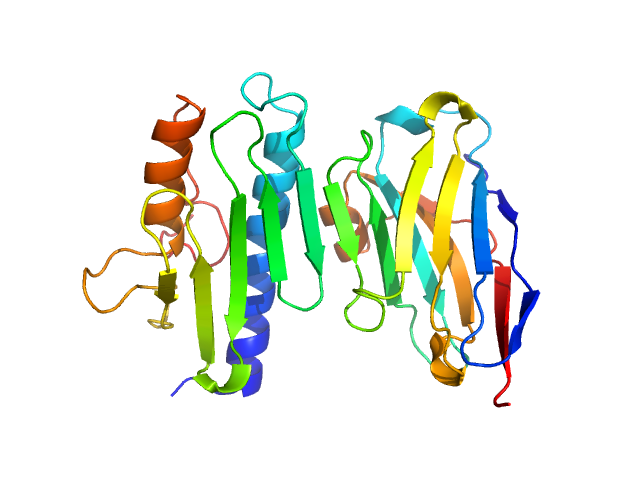
UniProt ID: Q16595 (90-210) Frataxin, mitochondrial
UniProt ID: None (1-142) Nanobody 28F6
|
|
|
|
| Sample: |
Frataxin, mitochondrial monomer, 14 kDa Homo sapiens protein
Nanobody 28F6 monomer, 15 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Sep 18
|
Nanobodies as tools for studying human frataxin biology.
Commun Biol (2026)
Pignataro MF, Fernández NB, Garay-Alvarez A, Pavan MF, Molina R, Muñoz IG, Grossi J, Noguera M, Vila A, García AE, Gentili HG, Rodríguez NA, Aran M, Parreño V, Bok M, Hermoso JA, Ibañez LI, Santos J
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
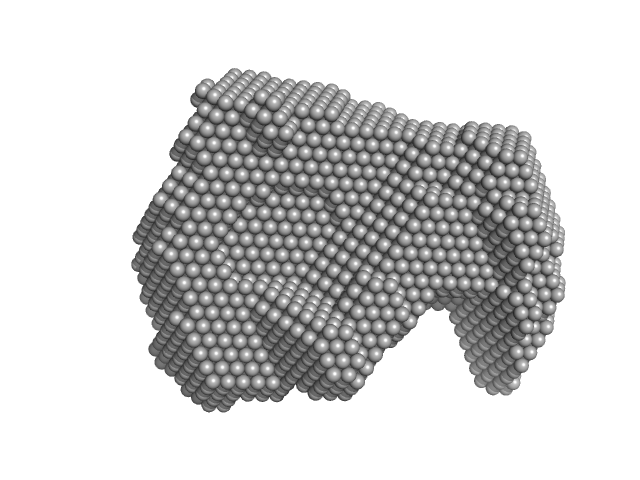
UniProt ID: C4LX08 (1-234) Anti-Silencing Function 1
UniProt ID: Q06196 (1-135) Histone H3
UniProt ID: P40287 (1-118) Histone H4
|
|
|

|
| Sample: |
Anti-Silencing Function 1 monomer, 28 kDa Entamoeba histolytica strain … protein
Histone H3 monomer, 15 kDa Entamoeba histolytica strain … protein
Histone H4 monomer, 13 kDa Entamoeba histolytica strain … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, and 2 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Oct 1
|
Characterisation of Entamoeba histolytica Anti-Silencing Function 1 as a histone chaperone.
Biochimie (2025)
Gandhi S, Vasudevan D
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
UniProt ID: P18472 (68-210) TraW (Δ1-67)
|
|
|

|
| Sample: |
TraW (Δ1-67) monomer, 20 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 5% glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 17
|
Solution characterization of TraW, a regulatory protein of the F plasmid type 4 secretion system
Structural Dynamics 13(2) (2026)
Rodriguez C, Audette G
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
|
|